Team:Cambridge/Experiments/Synthetic Reflectin PCR and Construction of GA1 to 6
From 2011.igem.org
(→Synopsis) |
(→Construct Design) |
||
| Line 82: | Line 82: | ||
We managed to join pBAD/araC promoter and B0015 ribosome binding site with the low copy number plasmid backbone using a pair of primers complementary to the region around the standard annealing site, included in every standard biobrick backbone. | We managed to join pBAD/araC promoter and B0015 ribosome binding site with the low copy number plasmid backbone using a pair of primers complementary to the region around the standard annealing site, included in every standard biobrick backbone. | ||
| - | For GA13 and GA14 constructs, the template DNA we used were GA1 and GA2 plasmids, respectively. We inserted GFP gene [http://partsregistry.org/Part:BBa_E0040 E0040], contributed by J69511:pSB3K3 plasmid, between the reflectin coding region and the B0015 terminator. | + | For '''GA13''' and '''GA14''' constructs, the template DNA we used were GA1 and GA2 plasmids, respectively. We inserted GFP gene [http://partsregistry.org/Part:BBa_E0040 E0040], contributed by J69511:pSB3K3 plasmid, between the reflectin coding region and the B0015 terminator. |
Sequences of the primers used in the experiment and their predicted melting temperatures are the following: | Sequences of the primers used in the experiment and their predicted melting temperatures are the following: | ||
| - | 1 | + | {| |
| + | |Name | ||
| + | |Sequence | ||
| + | |Tm | ||
| + | |- | ||
| + | |1 | ||
| + | |5' AGAAAGAGGAGAAATACTAG ATGGGATCCATGAACCGTTACC 3' | ||
| + | |67.89°C | ||
| + | |||
| + | |||
| + | |||
2 | 2 | ||
3 | 3 | ||
Revision as of 17:21, 19 September 2011
Contents |
Synthetic Gene Amplification & Plasmid Construction
Synopsis
The synthetic genes were extracted from the filter paper on which they were sent, and amplified by PCR.
Eight plasmids were then constructed, named GA1-6, GA13 and GA14 as shown below. Each puts reflectin expression under the control of pBAD promoter, an arabinose inducible promoter. HIS tags were included on some of the constructs in order to purify the protein for the in vitro work. We also put both reflectin A2 and 1B in a low expression plasmid. In addition, we created Reflectin A1 translational fusions with GFP at the C-terminus, which allowed for tracing the localization of reflectin in a bacterial cell at high and low level of expression.
| Name | Description | ||
| Gene | Plasmid | Promoter | |
| GA1 | Reflectin A1 | on PSB1A2 (high copy) | pBAD promoter |
| GA2 | Reflectin A1 | on PSB3K3 (low copy) | pBAD promoter |
| GA3 | Reflectin A1 + HIS tag | on PSB1A2 (high copy) | pBAD promoter |
| GA4 | Reflectin A1 + HIS tag | on PSB3K3 (low copy) | pBAD promoter |
| GA5 | Reflectin A2 | on PSB3K3 (low copy) | pBAD promoter |
| GA6 | Reflectin 1B | on PSB3K3 (low copy) | pBAD promoter |
| GA13 | Reflectin A1 : GFP | on PSB3K3 (low copy) | pBAD promoter |
| GA14 | Reflectin A1 : GFP | on PSB3K3 (low copy) | pBAD promoter |
These constructs were grown up in E.coli and verified by colony PCR. The plasmids were then miniprepped and stored at -20°C.
Extraction & Amplification
Wendy Crookes-Goodson kindly supported us with three [http://www.bio-protech.com.tw/databank/DataSheet/CustomSC/pUC57map.pdf pUC57] plasmids containg reflectin genes: Euprymna scolopes [http://www.ncbi.nlm.nih.gov/nuccore/33521697 Reflectin 1B], and Loligo pealei [http://www.ncbi.nlm.nih.gov/nuccore/269996957 Reflectin A1] and [http://www.ncbi.nlm.nih.gov/nuccore/FJ824805.1 Reflectin A2]. These are synthetic genes, codon optimized for expression in E.coli, and with all restriction sites removed from the coding region.
The plasmids with the reflectin gene were first extracted from filter paper, and then used to transform competent E. coli cells. A standard miniprep was used to recover the plasmid DNA from the overnight culture of the transformed bacteria in LB. The purpose of this procedure was to amplify and store the plasmid DNA.
Construct Design
For the assembly of the required plasmids, we amplified desired components by PCR and then joined them using Gibson Assembly procedure.
We used three different DNA templates:
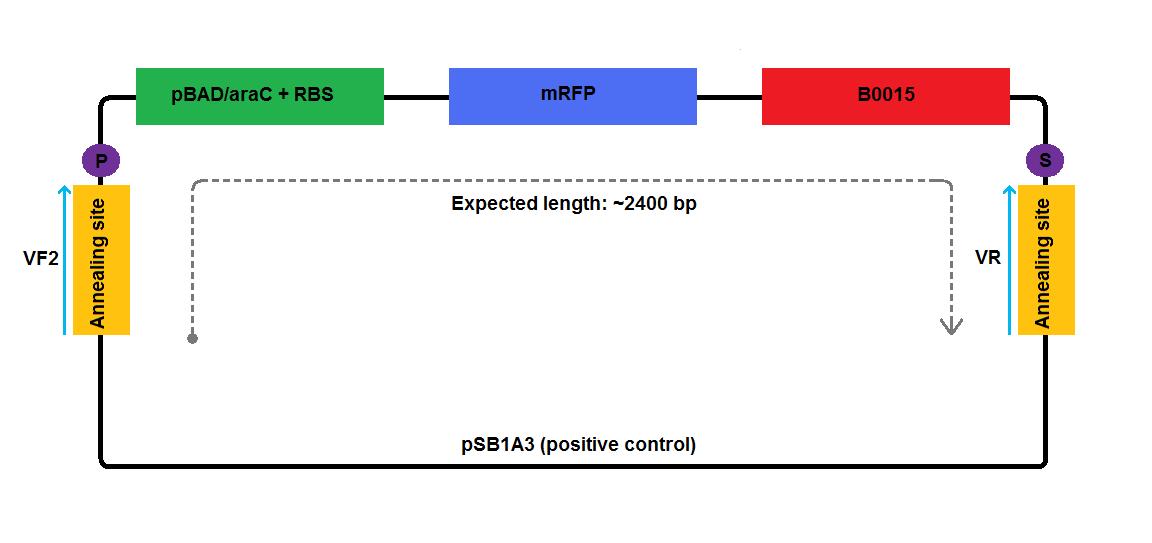
- [http://partsregistry.org/Part:BBa_I13520 I13520]:[http://partsregistry.org/Part:pSB1A3 pSB1A3], which contains mRFP gene under the control of pBAD promoter. It provided high copy number plasmid backbone and elements required for the expression of the gene of interest, such as pBAD/araC promoter [http://partsregistry.org/Part:BBa_I0500 I0500], ribosome binding site [http://partsregistry.org/Part:BBa_B0034 B0034] and terminator [http://partsregistry.org/Part:BBa_B0015 B0015].
- [http://partsregistry.org/Part:BBa_J69511 J69511]:[http://partsregistry.org/Part:pSB3K3 pSB3K3], which contains GFP under the control of [http://partsregistry.org/wiki/index.php?title=Part:BBa_R0011 R0011] promoter that allows for strong constitutive expression unless it is repressed by LacI. It provided low copy number plasmid backbone and terminator [http://partsregistry.org/Part:BBa_B0015 B0015].
- pUC57 plasmid with reflectin A1/A2/1B coding regions.
The his tag present in GA3 and GA4 constructs was included in primers.
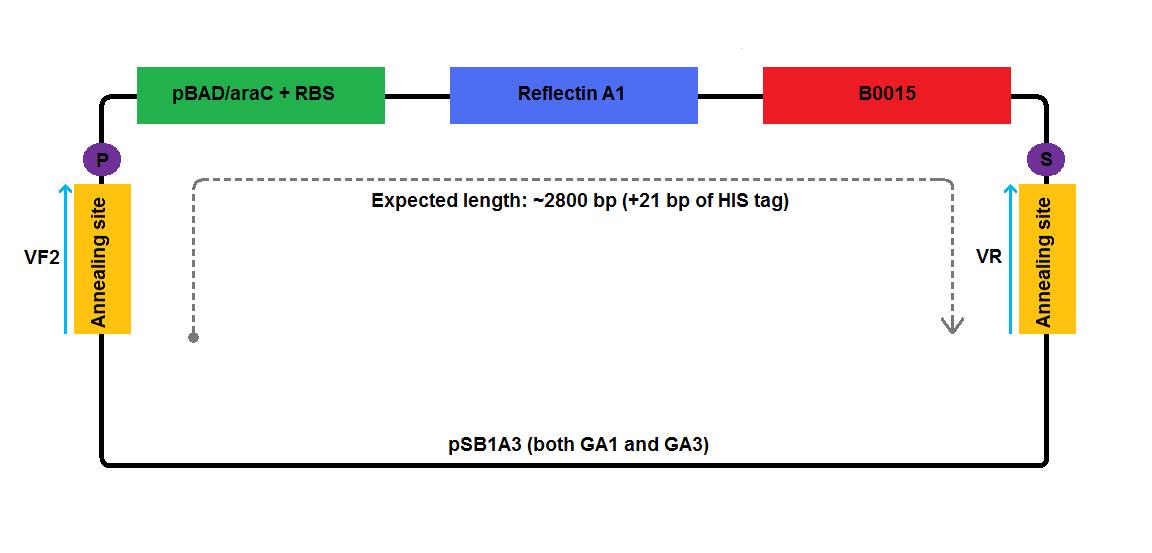
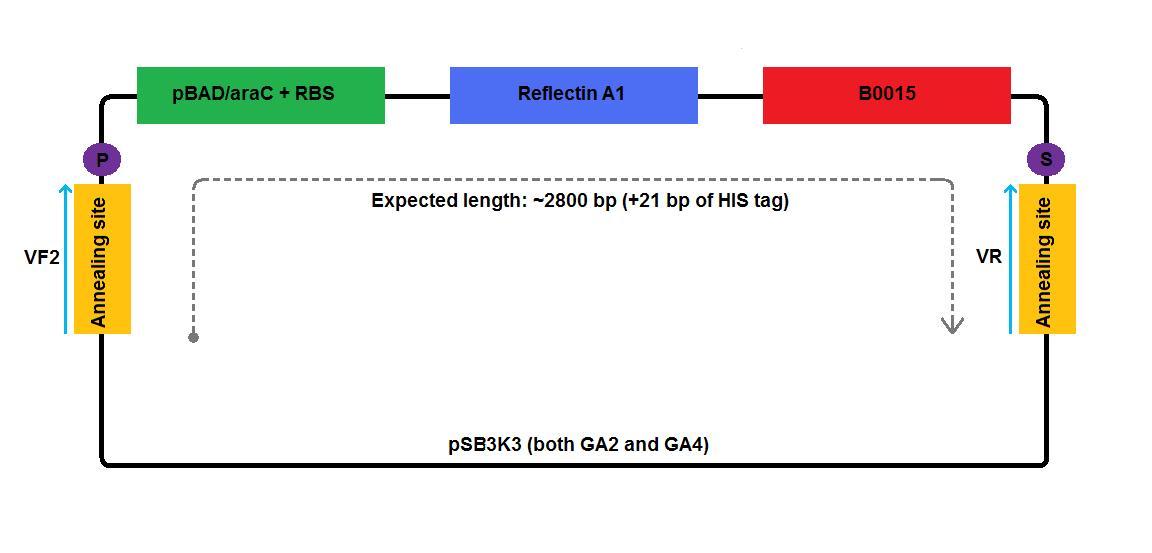
We created high copy number plasmids GA1 and GA3 by replacing mRFP coding region of pSB1A2:I13520 with reflectin A1 coding region.
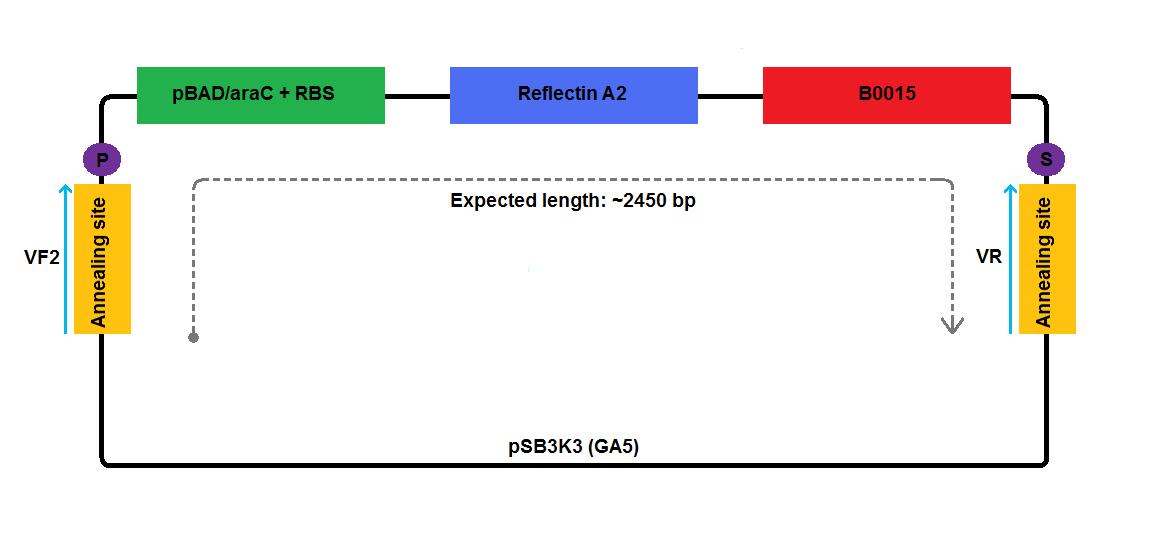
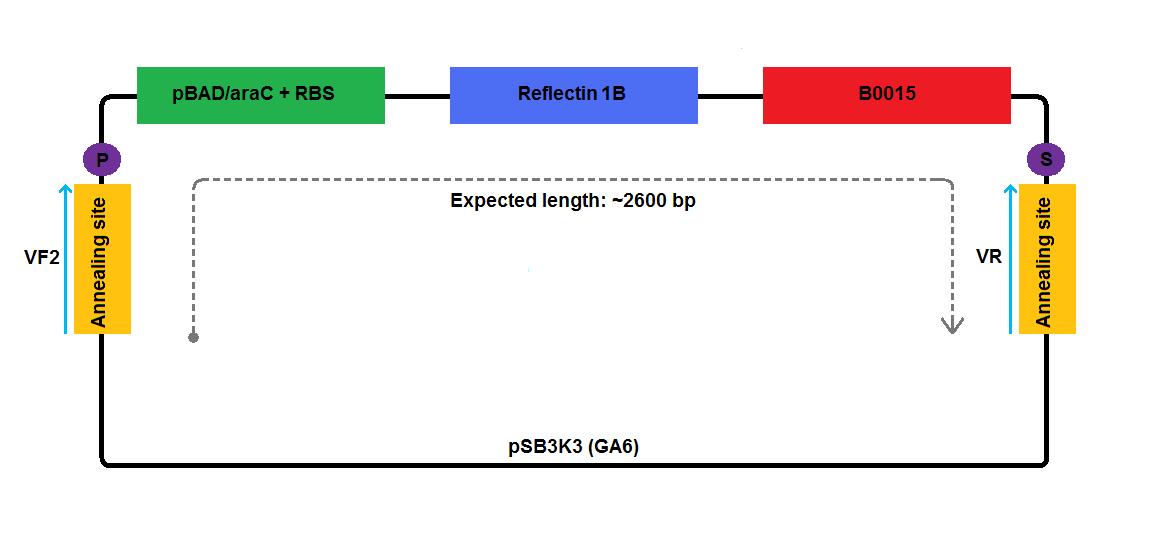
In the case of low copy number plasmids, GA2 and GA4-6, the respective reflectin coding region was joined to a part of pSB1A2 upstream (carrying pBAD/araC and RBS) and was attached to a part of the pSB3K3 downstream (contributing a terminator and a backbone). We managed to join pBAD/araC promoter and B0015 ribosome binding site with the low copy number plasmid backbone using a pair of primers complementary to the region around the standard annealing site, included in every standard biobrick backbone.
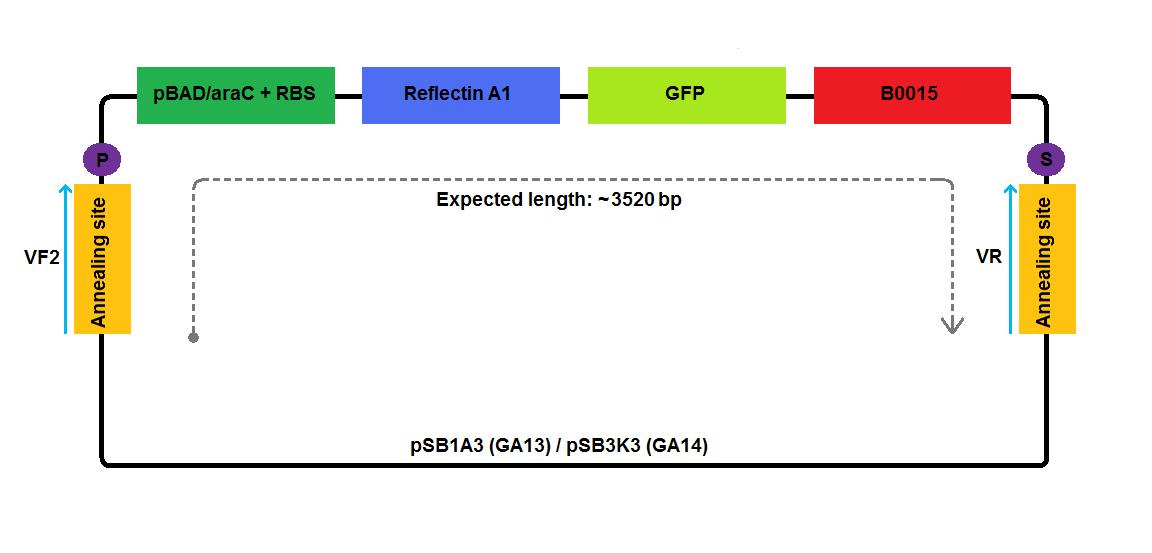
For GA13 and GA14 constructs, the template DNA we used were GA1 and GA2 plasmids, respectively. We inserted GFP gene [http://partsregistry.org/Part:BBa_E0040 E0040], contributed by J69511:pSB3K3 plasmid, between the reflectin coding region and the B0015 terminator.
Sequences of the primers used in the experiment and their predicted melting temperatures are the following:
| Name | Sequence | Tm | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 1 | 5' AGAAAGAGGAGAAATACTAG ATGGGATCCATGAACCGTTACC 3' | 67.89°C
2 3 4 5 6 1-his 4-his 1-a2 2-a2 3-a2 4-a2 1-1b 2-1b 3-1b 4-1b The pictures below give an overview of the assembly of the plasmids:
Assembly of GA1, GA2, GA3 and GA4 - First AttemptPCRIn the first round of PCR, we amplified fragments required for the assembly of GA1, GA2, GA3 and GA4 constructs.
We decided to use the 55°C annealing temperature, although the predicted temperature for most primers is 5-10°C higher, because of a low annealing temperature of the VF2 primer.
For the gel extraction of DNA we followed the protocol, assuming that one slice of gel is 100μl.
Gibson AssemblyWe conducted Gibson Assembly of GA1, GA2, GA3 and GA4 constructs according to the protocol. The volumes of Master Mix and solutions of amplified DNA were the following:
TransformationWe transformed competent E.coli cells according to the following protocol. We cultured each class of transformants on four different plates.
After overnight incubation at 37°C of transformed E.coli cells, we could see colonies on only some of our plates. We examined successful plates under a fluorescence microscope to check if the cells had been transformed with a carry-through of the template DNA used in the initial PCR reactions. The risk of contaminating Gibson Assembly reactions with template DNA was fairly high because circular DNA of around 4kb was likely to co-localize with the linear PCR products.
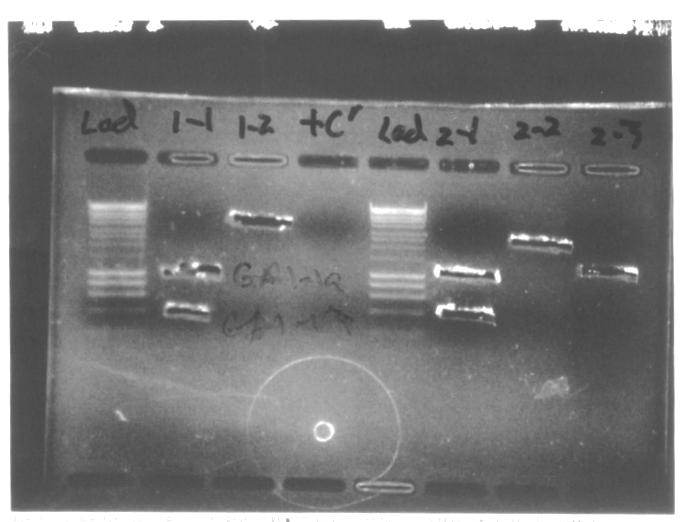
Colony PCRWe decided to conduct colony PCR of E.coli transformed with GA1 and GA3 constructs in order to confirm successful assembly of the plasmids.
pSB1AK3 or pSB1A3 backbone?As a side experiment, we decided to determine whether constructs with pSB1A3 backbone also carry a gene for kanamycin resistance. When preparing glycerol stocks of successful transformants, we cultured GA1x and GA3c bacteria in LB + kanamycin liquid medium in addition to LB + ampicillin medium. After 4 hour incubation at 37° the turbidity of both cultures was the same.
DiagnosticsIn addition, we performed a series of tests to identify the cause of the low efficiency of transformation. The proposed sources of error included:
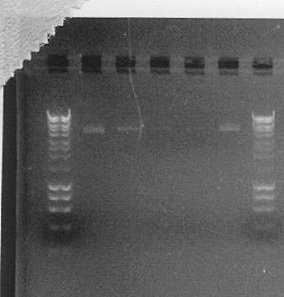
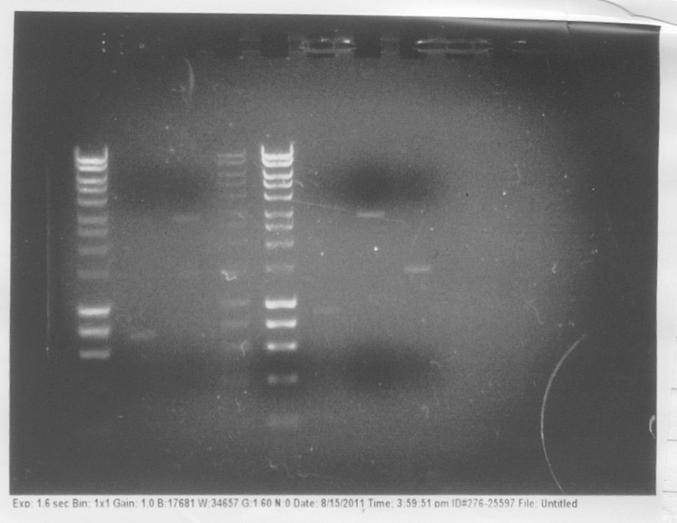
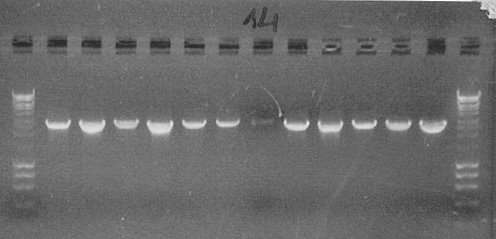
In order to check if gel extraction of DNA was successful, we ran a 1% agarose gel with samples of purified products of the initial PCR reactions. Although we could see distinct, fairly thick bands on the first gels, from which PCR products were purified, bands on the diagnostic gel were very faint, even missing in some lanes. This indicates that the yield of our extraction procedure was very low and probably this was the main reason why the performed transformation was fairly unsuccessful. Therefore, we decided to repeat PCR reactions, this time at higher 50μl volume, and try different conditions of the gel extraction procedure.
Assembly of GA1, GA2, GA3, GA4, GA5 and GA6 - Second AttemptPCRIn the second round of PCR, we amplified fragments required for the assembly of GA1, GA2, GA3, GA4, as well as GA5 and GA6 constructs.
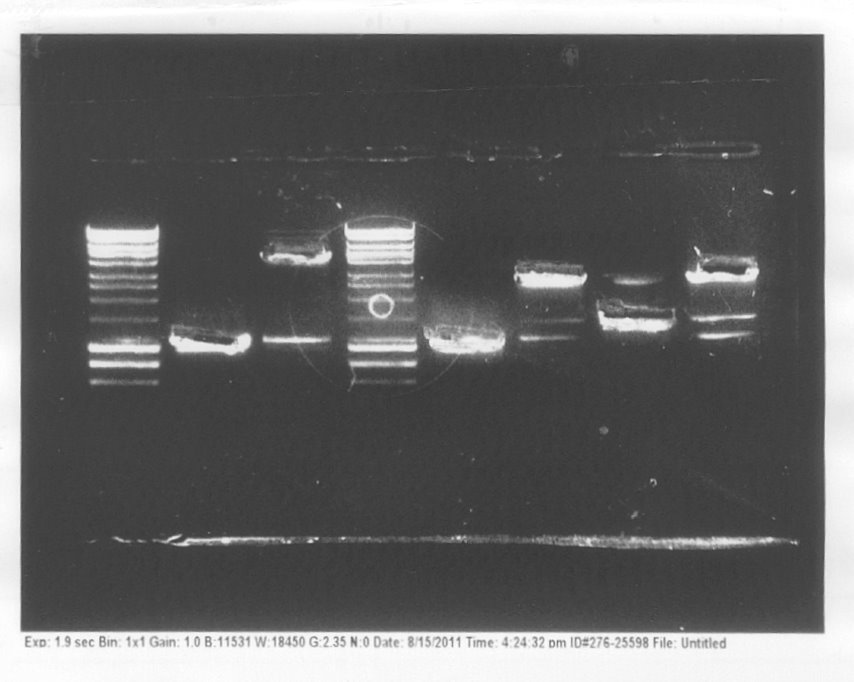
We also ran a gel with products of extraction to check how efficient the procedure was.
Gibson AssemblyWe conducted Gibson Assembly of GA1, GA2, GA3, GA4, GA5 and GA6 constructs according to the protocol. The total reaction volume was 20μl.
TransformationWe transformed competent E.coli cells according to the following protocol. We cultured each class of transformants on four different plates.
As a positive control, we transformed cells with I13520:pSB1A3 plasmid, expecting to detect RFP fluorescence on induced plates. As a negative control, we plated non-transformed heat-shocked competent cells on LB + kanamycin and LB + ampicillin plates. After overnight incubation at 37°C we examined plates under a fluorescence microscope to check if the cells had been also transformed with the template DNA used in the initial PCR reactions. The table presents expected phenotype of cells transformed with the correct Gibson Assembly construct or two DNA templates.
Colony PCRWe conducted colony PCR of E.coli transformed with GA1 - GA6 constructs in order to confirm successful assembly of these plasmids.
Assembly of GA13 and GA14PCRIn the third round of PCR, we amplified fragments required for the assembly of GA13 and GA14 constructs.
Gibson AssemblyWe performed Gibson Assembly of GA13 and GA14 construct in 20μ reactions.
TransformationWe transformed competent E.coli cells according to the following protocol. We cultured each class of transformants on four different plates, and we transformed cells with I13520:pSB1A3 as a positive control. Non-transformed heat-shocked competent E.coli cells plated on LB + ampicillin and LB + kanamycin served as a negative control.
After overnight incubation at 37° we screened colonies under a fluorescence microscope in order to check if they had been transformed with template DNA.
For both classes of transformants, colonies on non-induced plates showed no fluorescence, whereas they showed green fluorescence on induced plates, which served as a confirmation of correct expression of reflectin:GFP fusion. In order to prepare liquid cultures for colony PCR, we picked 4 colonies for each construct - two colonies from LB + antibiotic 100μ plate, one colony from LB + antibiotic 10μ plate and one colony from LB + antibiotic + arabinose 100μ plate.
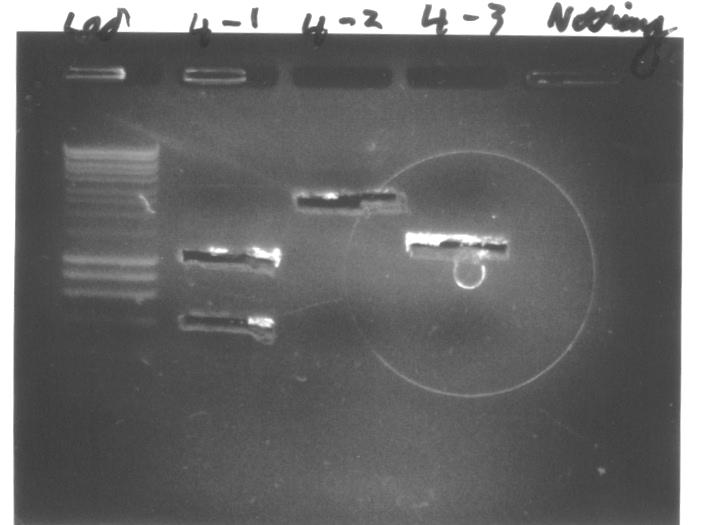
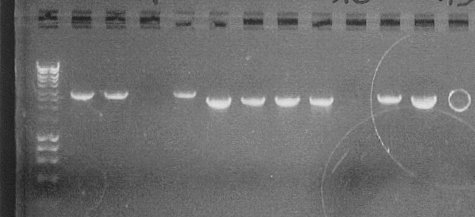
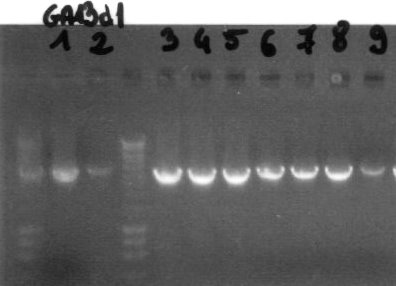
Colony PCRWe conducted colony PCR of E.coli transformed with GA13 and GA14 constructs in order to confirm successful assembly of these plasmids.
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
 "
"