Team:British Columbia/Notebook/Week 4
From 2011.igem.org
| Line 1: | Line 1: | ||
{{Template:Notebook}} | {{Template:Notebook}} | ||
| - | + | ==Lab Meeting== | |
| - | [[File:ubcnotebook.jpg]] | + | [[File:ubcnotebook.jpg | frame]] |
We created a set of lab rules due to common mistakes made in the lab over the past few weeks. | We created a set of lab rules due to common mistakes made in the lab over the past few weeks. | ||
| Line 13: | Line 13: | ||
For our modeling projects, Jacob obtained the ArcGIS program for modeling the pine beetle infestation and Gurpal created a wet lab flow chart to match the modeling. | For our modeling projects, Jacob obtained the ArcGIS program for modeling the pine beetle infestation and Gurpal created a wet lab flow chart to match the modeling. | ||
| - | + | ==Alpha-pinene== | |
| - | Joe's SDM-PCR worked (using the QuikChange SDM Kit protocol)! | + | Joe's SDM-PCR worked (using the QuikChange SDM Kit protocol)! Hurrah! |
| - | + | ==3-carene== | |
Daisy made primers for sequencing because the mutagen site is in the middle of the gene and her flanking primers will only sequence 700-800bp into the sequence. | Daisy made primers for sequencing because the mutagen site is in the middle of the gene and her flanking primers will only sequence 700-800bp into the sequence. | ||
| Line 27: | Line 27: | ||
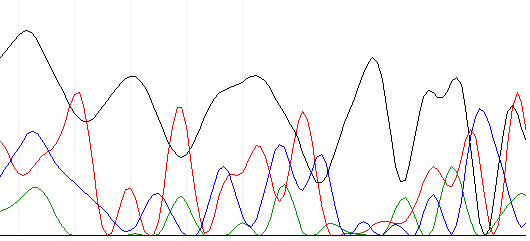
Below is an example of the chromatogram that came. | Below is an example of the chromatogram that came. | ||
| + | [[File:Sequencingseq.png | frame | left]] | ||
| - | + | <br><br><br><br><br><br><br><br><br><br><br><br><br><br> | |
| + | ==1,8-Cineole== | ||
| - | + | [[File:ubcwk5pcrtubes.jpg| thumb | left | 200px]] | |
| - | + | ||
| - | + | ||
| - | [[File:ubcwk5pcrtubes.jpg]] | + | |
Jacob ran successful a SDM-PCR to remove a restriction site as verified by restriction digest and gel electrophoresis. | Jacob ran successful a SDM-PCR to remove a restriction site as verified by restriction digest and gel electrophoresis. | ||
He transformed competent cells with his SDM-PCR product and also ran SDM-PCR on a variant of the original synthase. He ran Raf's mini-prep protocol on transformed cells to isolate a plasmid containing the SDMed pg-TPS-cin gene. | He transformed competent cells with his SDM-PCR product and also ran SDM-PCR on a variant of the original synthase. He ran Raf's mini-prep protocol on transformed cells to isolate a plasmid containing the SDMed pg-TPS-cin gene. | ||
| - | + | <br><br><br><br><br><br> | |
| + | ==Beta-pinene & (-)-limonene== | ||
| - | Marianne and Vicki ran SDM-PCR on their synthase genes and verified their PCR products through gel electrophoresis. No bands were present in any of the lanes. Troubleshooting: The dNTP could have been too old/contaminated (we had trouble with this particular tube of dNTP from last year), the concentrations for the reagents were not optimized so... Marianne looked into some protocols and optimized the SDM-PCR protocol. | + | Marianne and Vicki ran SDM-PCR on their synthase genes and verified their PCR products through gel electrophoresis. No bands were present in any of the lanes. |
| + | *Troubleshooting: The dNTP could have been too old/contaminated (we had trouble with this particular tube of dNTP from last year), the concentrations for the reagents were not optimized so... Marianne looked into some protocols and optimized the SDM-PCR protocol. Let the failures begin. | ||
| - | [[File:ubccookglove.jpg]] | + | [[File:ubccookglove.jpg | thumb | right | 300px]] |
| - | In a group, Vicki, Marianne, Gurpal and Jacob attempted to perform SDM to remove an internal cut-site. ThermoPol was used as a buffer and PFU | + | In a group, Vicki, Marianne, Gurpal and Jacob again attempted to perform SDM to remove an internal cut-site. ThermoPol was used as a buffer and PFU was used. After adding the primers and other constituents, we each ended up with ~ 25uL samples. We set our samples to run in a thermocycler for about 4 hours with specified conditions. When the cycling was completed, DpnI enzyme was added to each PCR tube and incubated overnight at 37oC. |
| - | + | To check if our SDM worked, we performed gel electrophoresis. Unfortunately, our gel showed that our site directed mutagenesis did not work as our product was still cut in two by the restriction enzyme. But, each of us learnt what not to do for next time which is always good. | |
Revision as of 05:50, 8 August 2011

 |
 |
 |
 |
 |
Contents |
Lab Meeting
We created a set of lab rules due to common mistakes made in the lab over the past few weeks.
Vicki and Marianne will have training sessions throughout the week to show team members without wet lab experience how to PCR, cast and run a gel, digest, ligate, transform, design primers and send mini-prepped products to be sequenced.
We also received and fulfilled a request from Grinnell College (a new iGEM team this year!) for the DspB part created by the 2010 UBC iGEM team.
For our modeling projects, Jacob obtained the ArcGIS program for modeling the pine beetle infestation and Gurpal created a wet lab flow chart to match the modeling.
Alpha-pinene
Joe's SDM-PCR worked (using the QuikChange SDM Kit protocol)! Hurrah!
3-carene
Daisy made primers for sequencing because the mutagen site is in the middle of the gene and her flanking primers will only sequence 700-800bp into the sequence.
She did a single digest with EcoRI-HF to test if the EcoRI site is in the synthase or not. Chris Keeling emailed Daisy the sequence he used for the truncation of the 3-Carene synthase.
A couple days later, the sequencing results came back. The NAPS unit (the place we send our samples to be sequenced) said there was no signal from the results.
Below is an example of the chromatogram that came.
1,8-Cineole
Jacob ran successful a SDM-PCR to remove a restriction site as verified by restriction digest and gel electrophoresis. He transformed competent cells with his SDM-PCR product and also ran SDM-PCR on a variant of the original synthase. He ran Raf's mini-prep protocol on transformed cells to isolate a plasmid containing the SDMed pg-TPS-cin gene.
Beta-pinene & (-)-limonene
Marianne and Vicki ran SDM-PCR on their synthase genes and verified their PCR products through gel electrophoresis. No bands were present in any of the lanes.
- Troubleshooting: The dNTP could have been too old/contaminated (we had trouble with this particular tube of dNTP from last year), the concentrations for the reagents were not optimized so... Marianne looked into some protocols and optimized the SDM-PCR protocol. Let the failures begin.
In a group, Vicki, Marianne, Gurpal and Jacob again attempted to perform SDM to remove an internal cut-site. ThermoPol was used as a buffer and PFU was used. After adding the primers and other constituents, we each ended up with ~ 25uL samples. We set our samples to run in a thermocycler for about 4 hours with specified conditions. When the cycling was completed, DpnI enzyme was added to each PCR tube and incubated overnight at 37oC.
To check if our SDM worked, we performed gel electrophoresis. Unfortunately, our gel showed that our site directed mutagenesis did not work as our product was still cut in two by the restriction enzyme. But, each of us learnt what not to do for next time which is always good.
 "
"