Team:British Columbia/Model3
From 2011.igem.org

Monoterpene Synthase Structural Modeling
Three-dimensional Structure of the 3 Terpene Synthase Proteins
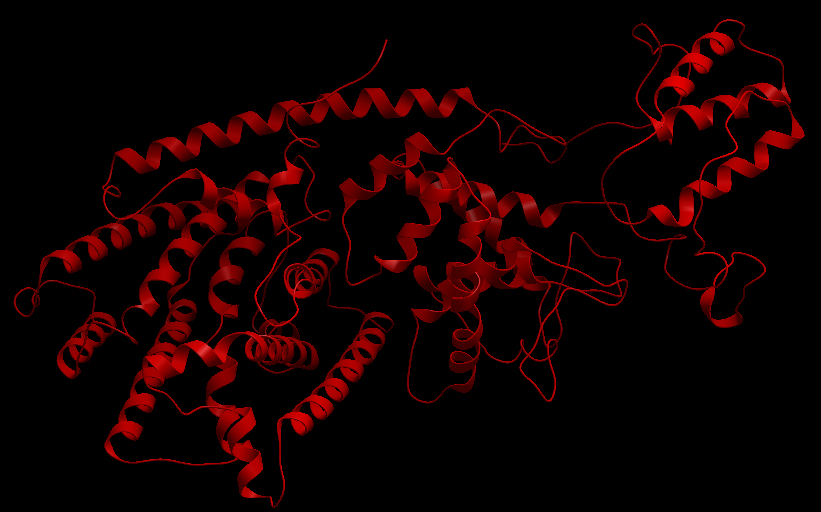
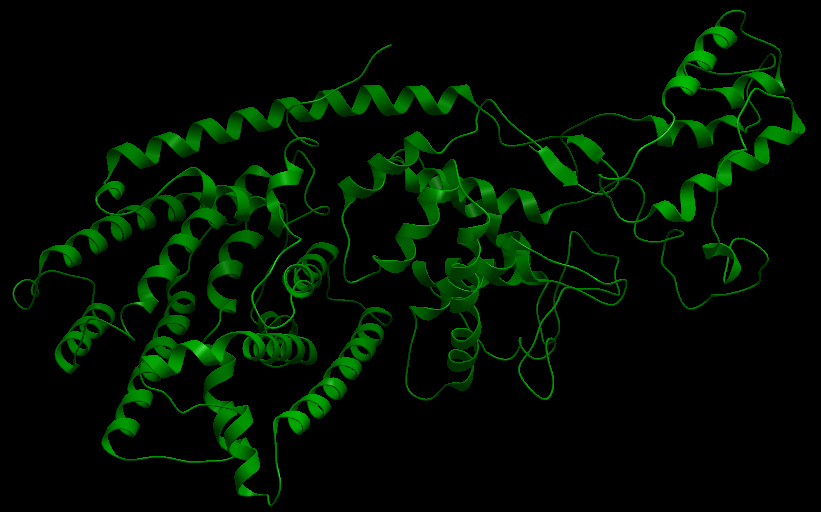
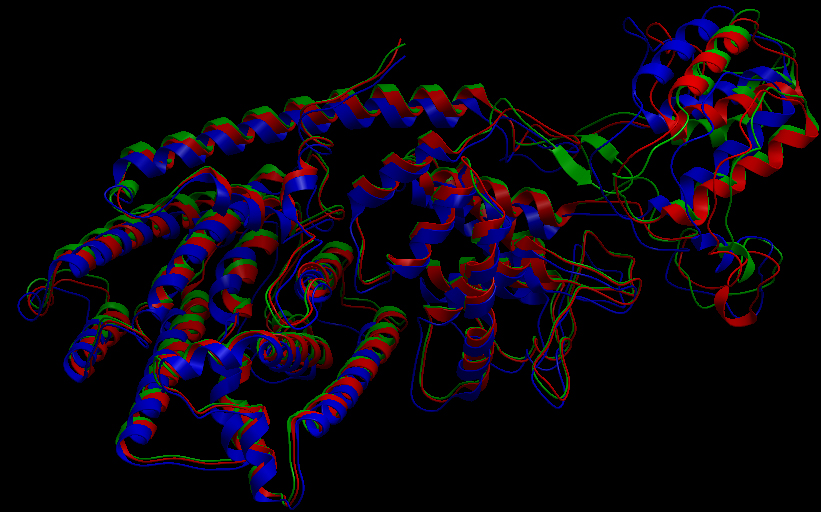
Ribbon representation of the alpha-pinene synthase. (HQ426166). Ribbon representation of the beta-pinene synthase (HQ426154). Ribbon representation of the limonene synthase (AY473624). Superimposition of the 3D structures of the three synthase (blue), beta-pinene synthase (red), and limonene synthase (green). The synthases exhibit high structural similarity. This observation allows us to identify homologous amino acid residues that may affect terpene synthesis efficiency.Reactive site of the alpha-pinene synthase
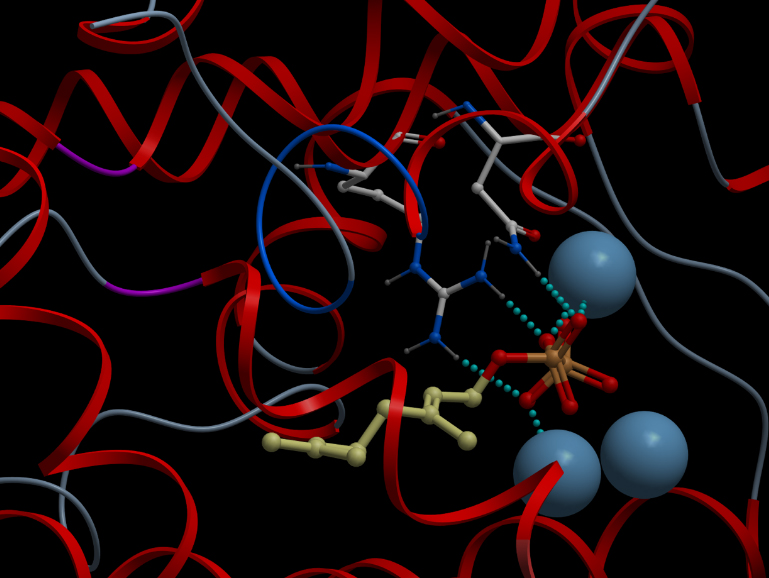
Skin representation of the reactive site of alpha-pinene synthase. The substrate, geranyl diphosphate (GDP), is docked inside the reactive pocket. The oxygen atoms (red spheres) belonging to the phosphate groups of GDP and the carbon atoms (yellow spheres) forming the backbone of GDP are shown. Magnesium ion cofactors (blue spheres) interact with GDP. Alternative representation of the reactive site of alpha-pinene synthase. The synthase structure is shown in ribbon model and the substrate GDP in ball-and-stick model. The magnesium ion cofactors are represented by blue spheres. The dotted light blue lines indicate hydrogen bonds.List of Amino Acid Residues in Alpha-Pinene Synthase Within 3.5 Angstrom of the Reactive Site
606G, 609Q, 610V, 613D, 710Y, 714V, 715G, 719C, 753W, 754R, 757N, 765E, 770Q
We propose that these sites are important to terpene synthesis.
Materials and Methods
We used MODELLER (1) to automate homology-based 3D structure prediction. We identified an experimentally determined 3D structure of a taxadiene synthase from Pacific Yew (PDB ID: 3P5R) as an appropriate template for modeling. The substrate GDP was docked using functions implemented in ICM by Molsoft LLC (2). All images were taken using the ICM by Molsoft LLC.
Future Directions
In the current study, we identified putative amino acid residues that may be important to the efficient of terpene synthesis. We propose that these amino acid sites can be mutated in the future to create mutant versions of the terpene synthases that exhibit greater terpene production output.References
1. N. Eswar, M. A. Marti-Renom, B. Webb, M. S. Madhusudhan, D. Eramian, M. Shen, U. Pieper, A. Sali. Comparative Protein Structure Modeling With MODELLER. Current Protocols in Bioinformatics, John Wiley & Sons, Inc., Supplement 15, 5.6.1-5.6.30, 2006.2. Abagyan, R.A., Totrov, M.M., and Kuznetsov, D.A. Icm: A New Method For Protein Modeling and Design: Applications To Docking and Structure Prediction From The Distorted Native Conformation. J. Comp. Chem. 15, 488-506. 1994.
Acknowledgement
We are very grateful to Yvonne Y. Li from the BC Genome Sciences Centre for valuable technical advice.
 "
"