Team:British Columbia/Notebook/Week 10
From 2011.igem.org
| Line 2: | Line 2: | ||
<b> Week 10: August 7-13 </b> | <b> Week 10: August 7-13 </b> | ||
| - | ==Lab Meeting - August 8 | + | ==Lab Meeting - August 8== |
| + | |||
| + | Today, we mostly discussed fundraising. We came up with a list of potential sponsors; each member is responsible for contacting their assigned sponsor. We need money! | ||
| + | |||
| + | Laura gave a very detailed update on her ideas for human practices, which includes videos, and a presentation at Science World. | ||
| + | |||
==UBC iGEM Logo== | ==UBC iGEM Logo== | ||
| Line 21: | Line 26: | ||
[[File:UBC IGEM_logo_7.JPG | thumb | center | bottom | Image B]] | [[File:UBC IGEM_logo_7.JPG | thumb | center | bottom | Image B]] | ||
| - | |||
| - | |||
[[File:ubcigemvickilb.jpg]] | [[File:ubcigemvickilb.jpg]] | ||
| Line 28: | Line 31: | ||
[[File:ubcigemrfp1.jpg]] | [[File:ubcigemrfp1.jpg]] | ||
| - | ==1,8- | + | ==1,8-Cineole== |
Jacob, now armed with what he believes are two versions of fully SDM'd 1,8-cineoles (still no sequencing results), decided to come in on the weekend and PCR, ligate, and grow everything. If everything worked perfectly, he could present some satisfying results at the Monday meeting. However, nothing ever seems to go as planned, and there were several setbacks. | Jacob, now armed with what he believes are two versions of fully SDM'd 1,8-cineoles (still no sequencing results), decided to come in on the weekend and PCR, ligate, and grow everything. If everything worked perfectly, he could present some satisfying results at the Monday meeting. However, nothing ever seems to go as planned, and there were several setbacks. | ||
| Line 34: | Line 37: | ||
Next came the ligations. | Next came the ligations. | ||
| - | Vicki prepared new pSB1C3 due to our previous stocks' odd behaviour (which included empty tubes that seemed to produce ligations and tubes that ran in two pieces without being cut), and cut it with ECORI and PSTI to visualize it on a gel. Jacob used some of this cut product for the biobrick ligation, along with | + | Vicki prepared new pSB1C3 due to our previous stocks' odd behaviour (which included empty tubes that seemed to produce ligations and tubes that ran in two pieces without being cut), and cut it with ECORI and PSTI to visualize it on a gel. The gel showed bands at the correct lengths (~1000bp) (Besides, the overnight culture turned pink/red - what else could it be if not pSB1C3 with RFP in it!). Jacob used some of this cut product for the biobrick ligation, along with ECORI and PSTI cut pgxe-tps-cin. |
| + | |||
Preparing the ligation products for the yeast was a bit harder. Concerning the restriction digest for the two synthases, the two sites on the ends of the synthases were SMAI and NOTI, the former of which only works in NEB buffer 4 at 25°, while the latter only had a 25% effectiveness in NEB buffer 4, and worked best at 37°. Jacob felt there was not enough product for a sequential digest, so he used NEB buffer 4 and let it sit at 25° for an hour then 37° for an hour. | Preparing the ligation products for the yeast was a bit harder. Concerning the restriction digest for the two synthases, the two sites on the ends of the synthases were SMAI and NOTI, the former of which only works in NEB buffer 4 at 25°, while the latter only had a 25% effectiveness in NEB buffer 4, and worked best at 37°. Jacob felt there was not enough product for a sequential digest, so he used NEB buffer 4 and let it sit at 25° for an hour then 37° for an hour. | ||
| + | |||
The biobrick was prepared by ligating the pSB1C3 with the synthases using T4 ligase, then transformed. The ligations for the yeast parts would have taken too long, so Jacob waited until the next day for the shuttle vector ligations. | The biobrick was prepared by ligating the pSB1C3 with the synthases using T4 ligase, then transformed. The ligations for the yeast parts would have taken too long, so Jacob waited until the next day for the shuttle vector ligations. | ||
Revision as of 05:31, 9 August 2011

 |
 |
 |
 |
 |
Lab Meeting - August 8
Today, we mostly discussed fundraising. We came up with a list of potential sponsors; each member is responsible for contacting their assigned sponsor. We need money!
Laura gave a very detailed update on her ideas for human practices, which includes videos, and a presentation at Science World.
UBC iGEM Logo
Joe is waiting for his primers to come. In the mean time, he is working on this year's iGEM logo on Adobe Illustrator. Living in the most beautiful place on Earth, with trees, mountains and serenity, it is hard not to be inspired by the scenic beauty of British Columbia, Canada. At the same, Joe wants to illustrate the effects of the mountain pine beetle epidemic on our forests, which is fueled by recent global climate changes. Even a beautiful place like this can't escape these changes, showing how our Earth's ecosystems are ultimately linked. Here is what he has so far...
Please Vote for one of these two:
1,8-Cineole
Jacob, now armed with what he believes are two versions of fully SDM'd 1,8-cineoles (still no sequencing results), decided to come in on the weekend and PCR, ligate, and grow everything. If everything worked perfectly, he could present some satisfying results at the Monday meeting. However, nothing ever seems to go as planned, and there were several setbacks.
The PCRs for preparing both the synthase genes for biobricks and 415 PAG GPD and GAL (both his-tagged and not) worked well, with the exception of the pg-tps-1,8-cineole for the biobrick. As a side note, Jacob ran duplicates of each reaction, one with 0.5 uL added magnesium per 50 uL reaction, and one without. Only the ones with magnesium worked.
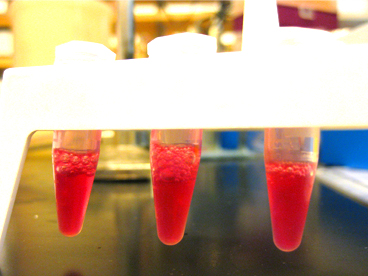
Next came the ligations. Vicki prepared new pSB1C3 due to our previous stocks' odd behaviour (which included empty tubes that seemed to produce ligations and tubes that ran in two pieces without being cut), and cut it with ECORI and PSTI to visualize it on a gel. The gel showed bands at the correct lengths (~1000bp) (Besides, the overnight culture turned pink/red - what else could it be if not pSB1C3 with RFP in it!). Jacob used some of this cut product for the biobrick ligation, along with ECORI and PSTI cut pgxe-tps-cin.
Preparing the ligation products for the yeast was a bit harder. Concerning the restriction digest for the two synthases, the two sites on the ends of the synthases were SMAI and NOTI, the former of which only works in NEB buffer 4 at 25°, while the latter only had a 25% effectiveness in NEB buffer 4, and worked best at 37°. Jacob felt there was not enough product for a sequential digest, so he used NEB buffer 4 and let it sit at 25° for an hour then 37° for an hour.
The biobrick was prepared by ligating the pSB1C3 with the synthases using T4 ligase, then transformed. The ligations for the yeast parts would have taken too long, so Jacob waited until the next day for the shuttle vector ligations.
The first biobrick ligation transformations were a failure.
The second biobrick ligation transformations were a success! That is to say, there were colonies, and they were not expressing the RFP protein.
The eventual transformations of the synthases on 415 PAG GPD and GAL grew colonies, but they are of dubious quality. The plates used supported colonies on the negative control plate, which had only competent cells. This raised the question of whether there was a problem with the plates or the cells themselves, and Jacob is running tests to check.
 "
"