Team:SJTU-BioX-Shanghai/Project/Subproject1-3
From 2011.igem.org
(Difference between revisions)
ChobitParrot (Talk | contribs) (→Location of Rare Codons) |
(→Number of Rare Codons) |
||
| (8 intermediate revisions not shown) | |||
| Line 9: | Line 9: | ||
__NOTOC__ | __NOTOC__ | ||
| - | ==Number of Rare Codons== | + | ==Part I: Number of Rare Codons== |
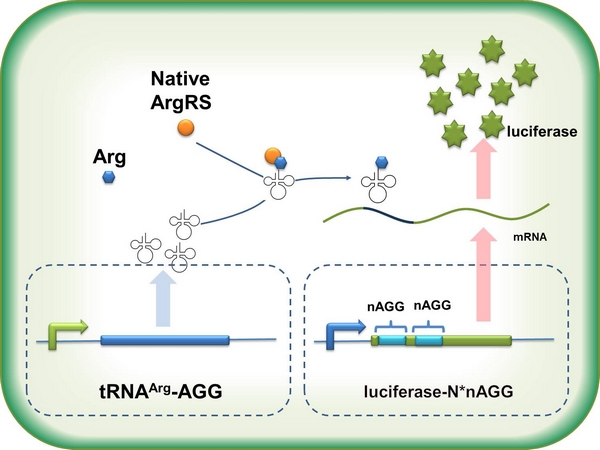
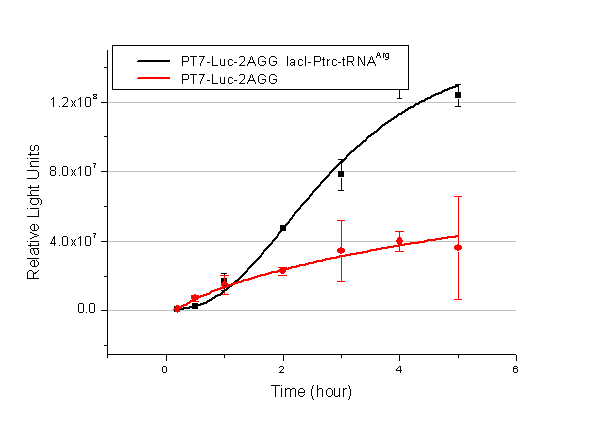
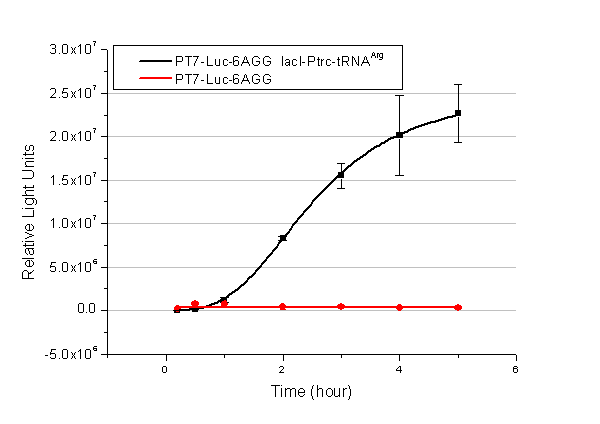
In this part we want to explore the influence of the number of rare codons inserted in the mRNA. We have inserted 2, 4, 6, 8 AGG codons respectively after the start codon in luciferase gene. T7 promoter or ''bla'' promoter<sup>[1]</sup> are used to control target protein mRNA amount. We use different combinations of number of AGG codons and strength of promoters to characterize regulation<sup>[1]</sup>. | In this part we want to explore the influence of the number of rare codons inserted in the mRNA. We have inserted 2, 4, 6, 8 AGG codons respectively after the start codon in luciferase gene. T7 promoter or ''bla'' promoter<sup>[1]</sup> are used to control target protein mRNA amount. We use different combinations of number of AGG codons and strength of promoters to characterize regulation<sup>[1]</sup>. | ||
| Line 63: | Line 63: | ||
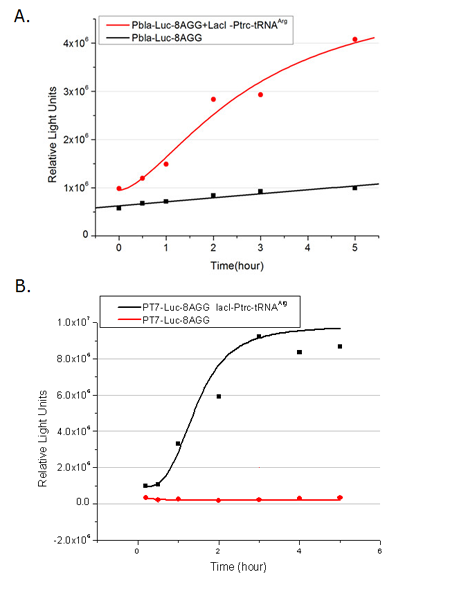
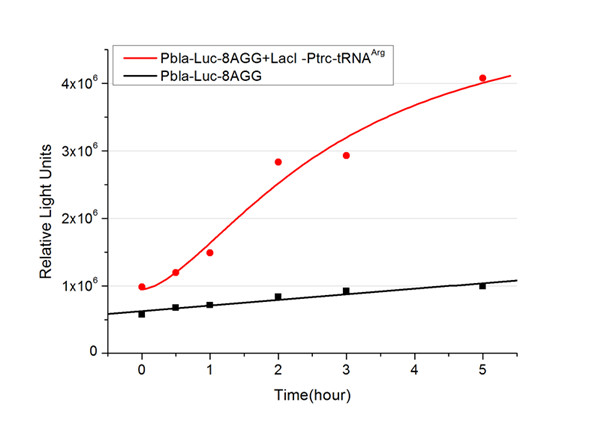
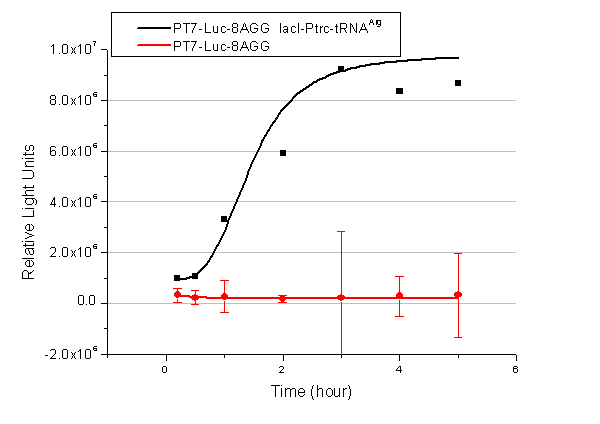
[[image:11sjtu_Compare_8.png|frame|center|''Fig.3'' (A) Luciferase (P''bla''-Luc-8AGG) production in cells overexpressing rare tRNA with ''lacI''-Ptrc-tRNA<sup>Arg</sup>. Luciferase production is reflected by bioluminescence emitted from the luciferin reaction. (B) Luciferase production in cells with PT7-Luc-8AGG as reporter. Here we analyze the influences of strong/weak promoter in luciferase production. Strong promoter (T7) of target gene can improve the titration curve, indicating that our device works better under strong target protein promoters. ]] | [[image:11sjtu_Compare_8.png|frame|center|''Fig.3'' (A) Luciferase (P''bla''-Luc-8AGG) production in cells overexpressing rare tRNA with ''lacI''-Ptrc-tRNA<sup>Arg</sup>. Luciferase production is reflected by bioluminescence emitted from the luciferin reaction. (B) Luciferase production in cells with PT7-Luc-8AGG as reporter. Here we analyze the influences of strong/weak promoter in luciferase production. Strong promoter (T7) of target gene can improve the titration curve, indicating that our device works better under strong target protein promoters. ]] | ||
| - | |||
==Location of Rare Codons== | ==Location of Rare Codons== | ||
| Line 81: | Line 80: | ||
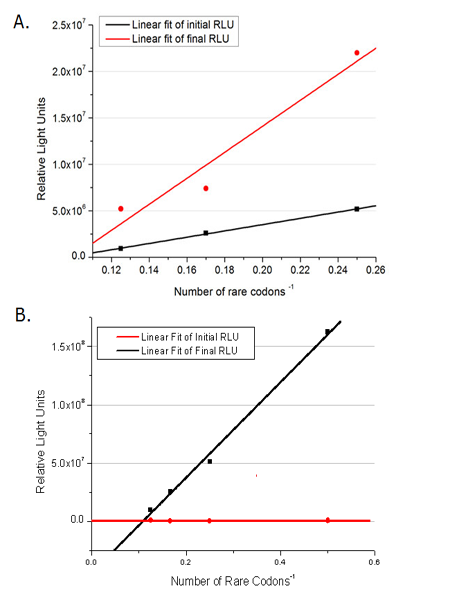
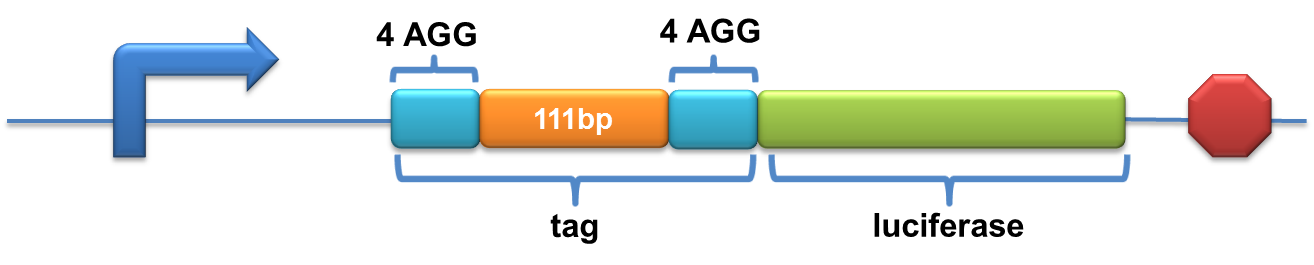
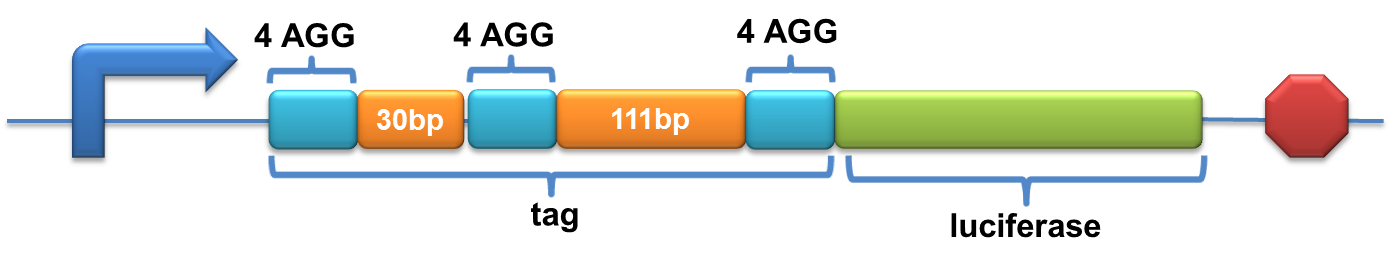
[[image:11SJTU-3X4AGG.png|600px]] | [[image:11SJTU-3X4AGG.png|600px]] | ||
| - | Experiment results showed that | + | Experiment results showed that when there is 111bp interval between the two-copy 4AGG, background is lower than there is a 30bp interval. In another analysis, protein production can be induced to a higher level when there are two copies of AGG tandems. |
| - | + | ||
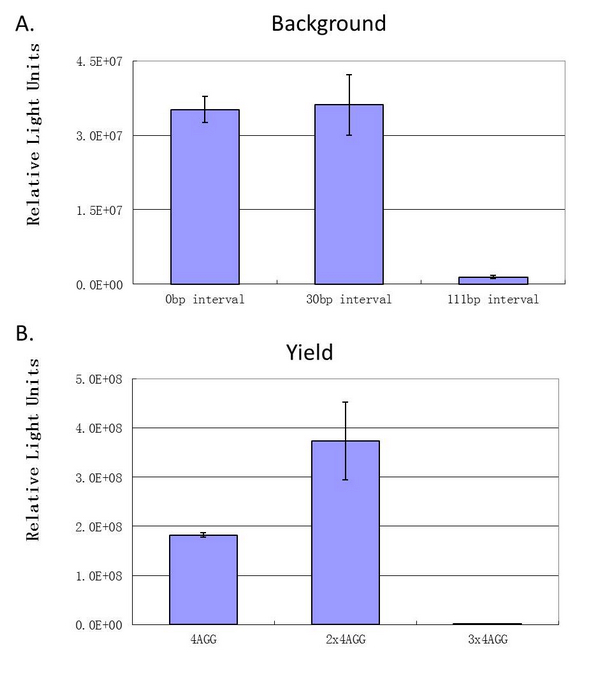
| - | When there | + | [[image:11SJTU_rare_27.jpg|frame|center|''Fig.4 A. When there is 111bp interval between the two-copy 4AGG, background is lower than there is a 30bp interval. B. Protein production can be induced to a higher level when there are two copies of AGG tandems.'']] |
'''Experiment results showed that luciferase tagged with two-copy 4AGG insertions with an interval of 111 bp can increase yield and lower background noise of target protein.''' We further conducted experiments to test its characters. | '''Experiment results showed that luciferase tagged with two-copy 4AGG insertions with an interval of 111 bp can increase yield and lower background noise of target protein.''' We further conducted experiments to test its characters. | ||
| Line 89: | Line 88: | ||
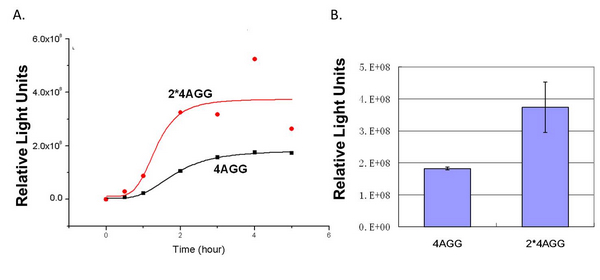
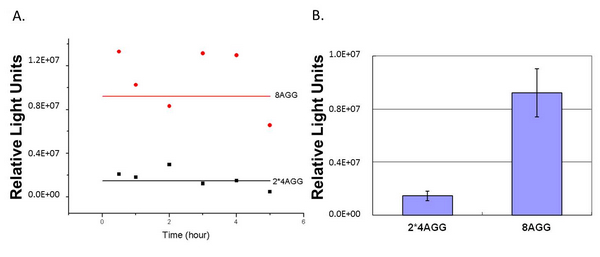
When we compare the two-copy 4AGG tagged luciferase with the single copy one, we find that two-copy 4AGG tagged luciferase has a higher yield. | When we compare the two-copy 4AGG tagged luciferase with the single copy one, we find that two-copy 4AGG tagged luciferase has a higher yield. | ||
| - | [[image:11SJTU_rare_25.jpg|frame|center | + | [[image:11SJTU_rare_25.jpg|frame|center|''fig.5 Two-copy 4AGG tagged luciferase has a higher yield than single-copy'']] |
When we compare the two-copy 4AGG tagged luciferase with 8AGG tagged luciferase, we find that two-copy 4AGG tagged luciferase has a lower background. | When we compare the two-copy 4AGG tagged luciferase with 8AGG tagged luciferase, we find that two-copy 4AGG tagged luciferase has a lower background. | ||
| - | [[image:11SJTU_rare_26.jpg|frame|center | + | [[image:11SJTU_rare_26.jpg|frame|center|''fig.6 Two-copy 4AGG tagged luciferase has a lower background than 8AGG tagged luciferase'']] |
==Note: Our device can be used as a regulating tool== | ==Note: Our device can be used as a regulating tool== | ||
| Line 99: | Line 98: | ||
We have tested luciferin reaction in cells. We examined the changes in luciferase enzyme activity over time after rare tRNA expression is induced. The amount of luciferase is reflected indirectly by the bioluminescence emitted from the luciferin reaction. Results are shown below: | We have tested luciferin reaction in cells. We examined the changes in luciferase enzyme activity over time after rare tRNA expression is induced. The amount of luciferase is reflected indirectly by the bioluminescence emitted from the luciferin reaction. Results are shown below: | ||
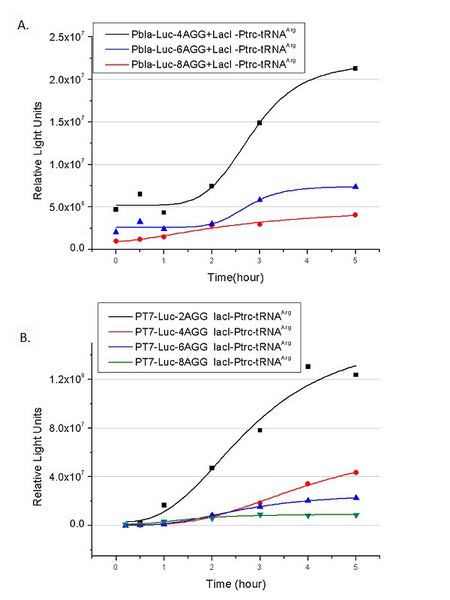
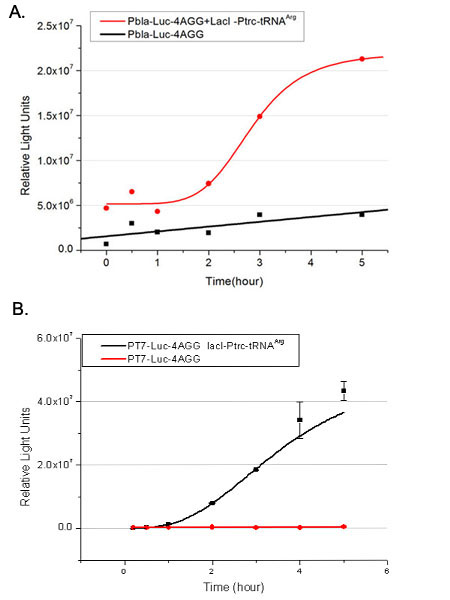
| - | [[image:11-SJTU-compare-4.jpg|frame|center|''Fig. | + | [[image:11-SJTU-compare-4.jpg|frame|center|''Fig.7'' (A)Luciferase (P''bla''-Luc-4AGG([http://partsregistry.org/wiki/index.php?title=Part:BBa_K567006 BBa_K567006])) production reflected by bioluminescence emitted from the luciferin reaction. (B)Luciferase (PT7-Luc-4AGG ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K567009 BBa_K567009])) production reflected by bioluminescence emitted from the luciferin reaction. ]] |
Here we use the above two curves as examples to characterize the working curve of our device. Both curves fits typical titration curve, indicating that our device can function as a regulating tool. | Here we use the above two curves as examples to characterize the working curve of our device. Both curves fits typical titration curve, indicating that our device can function as a regulating tool. | ||
| Line 106: | Line 105: | ||
<gallery caption="Pbla"> | <gallery caption="Pbla"> | ||
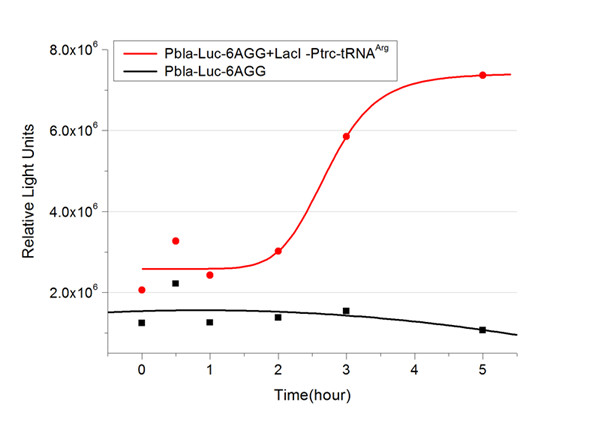
| - | image:11SJTUResultPlbla6.jpg|''Fig. | + | image:11SJTUResultPlbla6.jpg|''Fig.8'' |
| - | image:11SJTUResultPbla8.jpg|''Fig. | + | image:11SJTUResultPbla8.jpg|''Fig.9'' |
</gallery> | </gallery> | ||
<gallery caption="PT7"> | <gallery caption="PT7"> | ||
| - | image:11SJTUT7-2.png|''Fig. | + | image:11SJTUT7-2.png|''Fig.10'' |
| - | image:11SJTUT7-6.png|''Fig. | + | image:11SJTUT7-6.png|''Fig.11'' |
| - | image:11SJTUT7-8.png|''Fig. | + | image:11SJTUT7-8.png|''Fig.12'' |
</gallery> | </gallery> | ||
'''Note''':Click to see large figures. | '''Note''':Click to see large figures. | ||
| Line 120: | Line 119: | ||
From this experiment, we noticed that the typical working curve of our device can be better observed under IPTG induced ''lacI''-Ptrc-tRNA<sup>Arg</sup> ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K567001 BBa_K567001]) compared with UV excitation induced sulA promoter-tRNA<sup>Arg</sup>([http://partsregistry.org/wiki/index.php?title=Part:BBa_K567002 BBa_K567002]), though sulA promoter-tRNA<sup>Arg</sup> responded quicker to signals. So in the above experiments, we test with ''lacI''-Ptrc-tRNA<sup>Arg</sup>. | From this experiment, we noticed that the typical working curve of our device can be better observed under IPTG induced ''lacI''-Ptrc-tRNA<sup>Arg</sup> ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K567001 BBa_K567001]) compared with UV excitation induced sulA promoter-tRNA<sup>Arg</sup>([http://partsregistry.org/wiki/index.php?title=Part:BBa_K567002 BBa_K567002]), though sulA promoter-tRNA<sup>Arg</sup> responded quicker to signals. So in the above experiments, we test with ''lacI''-Ptrc-tRNA<sup>Arg</sup>. | ||
| + | |||
| + | ===Reference=== | ||
| + | [1]Ulrich Deuschlel., et al., Promoters of Escherichia coli: a hierarchy of in vivo strength indicates alternate structures The EMBO Journal vol.5 no. 11 pp.2987-2994, 1986 | ||
Latest revision as of 04:03, 29 October 2011
 "
"