Team:Bielefeld-Germany/Data Page
From 2011.igem.org
(Difference between revisions)
| Line 2: | Line 2: | ||
<html><img src="https://static.igem.org/mediawiki/2011/f/ff/Bielefeld-header-data-page.png"/><p></p></html> | <html><img src="https://static.igem.org/mediawiki/2011/f/ff/Bielefeld-header-data-page.png"/><p></p></html> | ||
| - | This page gives a basic overview about our cell-free bisphenol A biosensor system and the BioBricks we have used. A more detailed description of the biosensor system can be found in our [[Team:Bielefeld-Germany/Project/Description | project description]] | + | This page gives a basic overview about our cell-free bisphenol A biosensor system and the BioBricks we have used. A more detailed description of the biosensor system can be found in our [[Team:Bielefeld-Germany/Project/Description | project description]] as well as in the [[Team:Bielefeld-Germany/Project/Background/BPA | bisphenol A]], [[Team:Bielefeld-Germany/Project/Background/S-Layer | S-layer]] and [[Team:Bielefeld-Germany/Project/Background/NAD | NAD<sup>+</sup> detection]] background subsections. |
==How our system works== | ==How our system works== | ||
Revision as of 11:40, 21 September 2011


This page gives a basic overview about our cell-free bisphenol A biosensor system and the BioBricks we have used. A more detailed description of the biosensor system can be found in our project description as well as in the bisphenol A, S-layer and NAD+ detection background subsections.
Contents |
How our system works
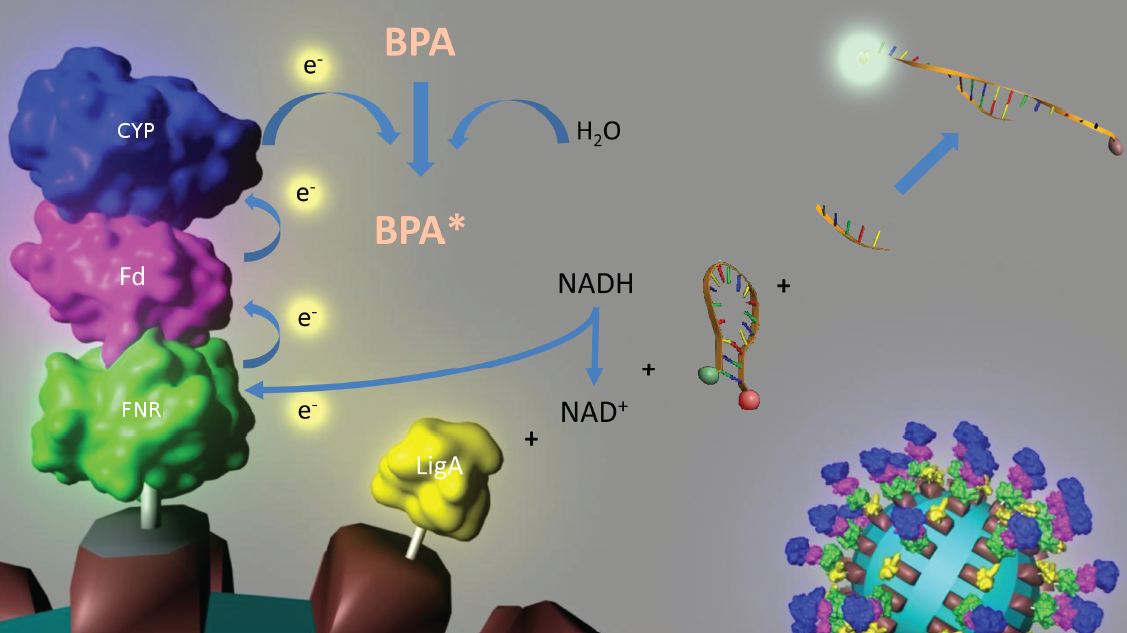
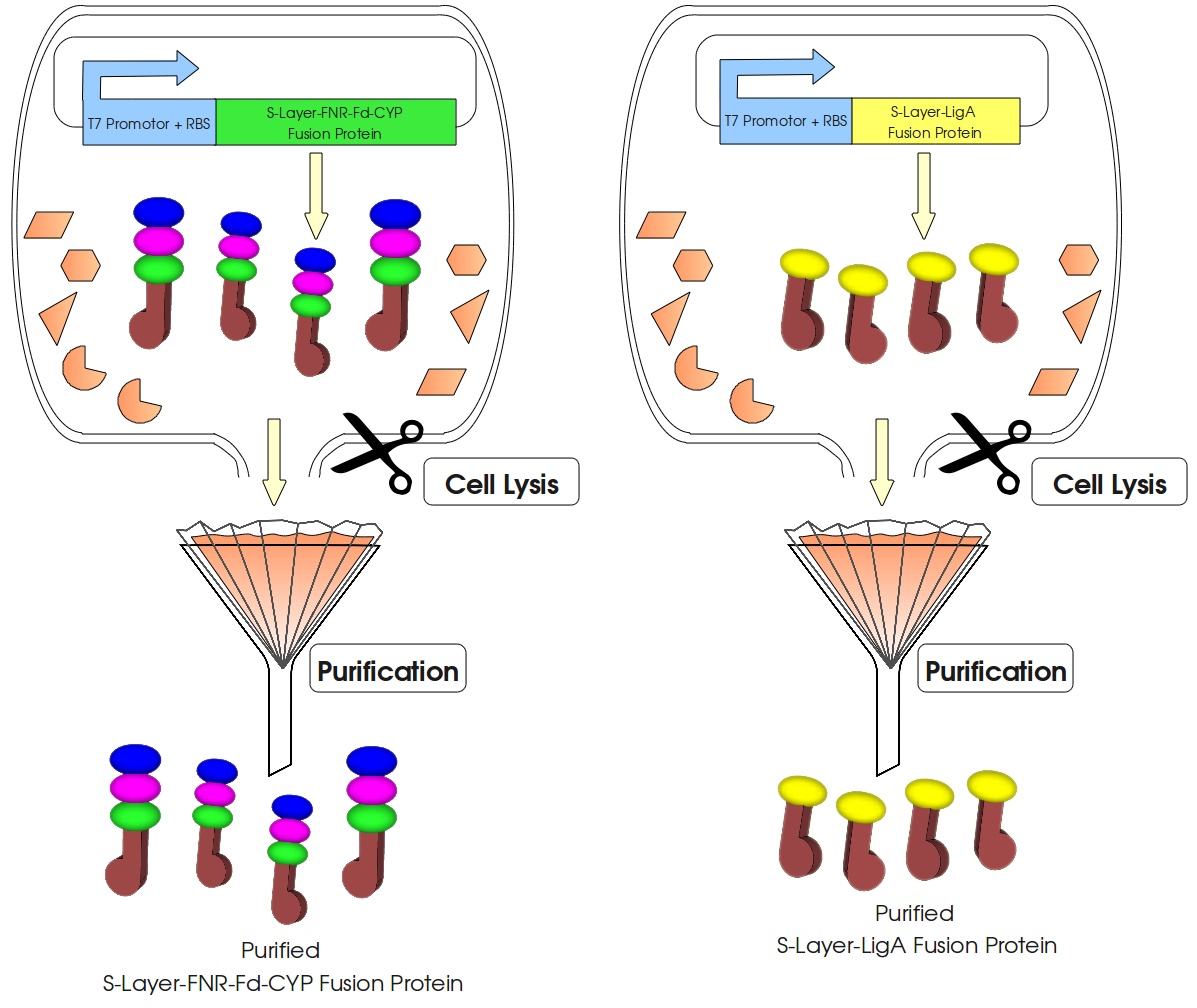
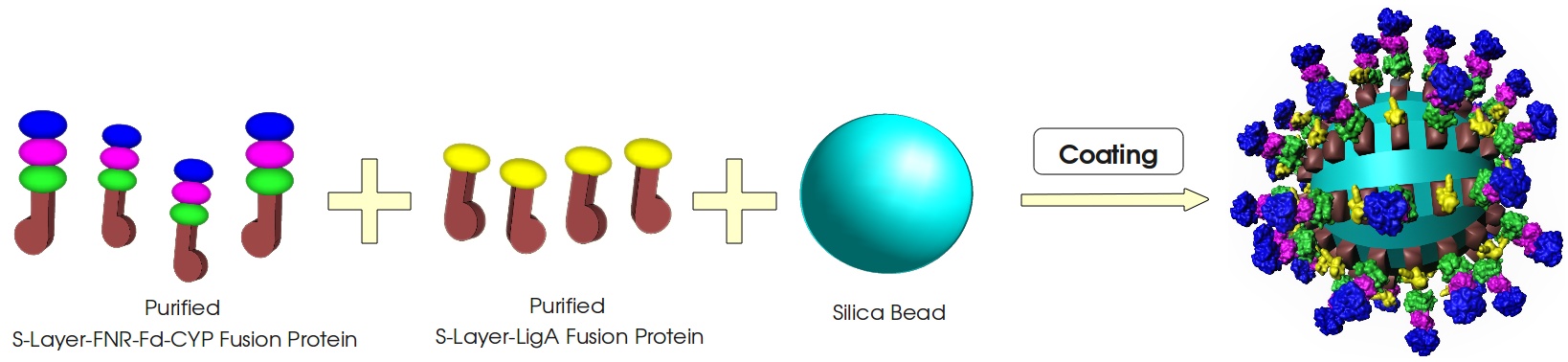
Figure 3: Visualization of our cell-free bisphenol A biosensor system with all essential components. Bisphenol A (BPA) is reduced by the electrons from NADH transferred by the ferredoxin-NADP+ oxidoreductase (FNR, <partinfo>BBa_K525499</partinfo>), ferredoxin (Fd, <partinfo>BBa_K123000</partinfo>) and cytochrome P450 (CYP, <partinfo>BBa_K123001</partinfo>) which are fused to a S-layer protein. The molecular beacon (hairpin structure) binds two short DNA oligos. The NAD+-dependent ligase (LigA), which is also fused to S-layer proteins, ligates the two oligos so that the hairpin structure opens up and the fluorophore is able to emit light after extinction.
Data for our favorite new parts
- [http://partsregistry.org/Part:BBa_K525305 Main Page] - Fusion protein of S-layer SgsE and mCitrine: This fluorescent S-layer fusion protein is used to characterize purification methods and to demonstrate the S-layers ability to self-assemble on surfaces.
- [http://partsregistry.org/Part:BBa_K525515 Main Page] - Fusion protein of BisdA and BisdB: This fusion protein improves the bisphenol A degradation in E. coli compared to the so far in the partsregistry existing BPA degrading BioBricks.
- [http://partsregistry.org/Part:BBa_K525710 Main Page] - NAD+-dependent DNA ligase from E. coli (LigA) : This enzyme enables determination of NAD+ even in very low concentrations by introducing it to a molecular beacon based bioassay.
Data for pre-existing parts
- [http://partsregistry.org/Part:BBa_K123000:Experience Experience] - BisdA degrades Bisphenol A when used with BisdB, BBa_K123000 (University of Alberta, iGEM 2008): Complete degradation of 120 mg L-1 Bisphenol A with polycistronic bisdAB gene in 30-33 h. Even faster (21-24 h) when using a fusion protein of BisdA and BisdB.
- [http://partsregistry.org/Part:BBa_K123001:Experience Experience] - BisdB degrades Bisphenol A when used with BisdA, BBa_K123001 (University of Alberta, iGEM 2008): Complete degradation of 120 mg L-1 Bisphenol A with polycistronic bisdAB gene in 30-33 h. Even faster (21-24 h) when using a fusion protein of BisdA and BisdB.
We have also characterized the following parts
- [http://partsregistry.org/Part:BBa_K525405 Main Page] - Fusion protein of S-layer SbpA and mCitrine: This fluorescent S-layer fusion protein is used to characterize purification methods and to demonstrate the S-layers ability to self-assemble on surfaces.
- [http://partsregistry.org/Part:BBa_K525512 Main Page] - Polycistronic expression of BisdA and BisdB: This is the version of BPA degrading BioBricks found in the partsregistry - comparison to our fusion protein <partinfo>K525515</partinfo>.
- [http://partsregistry.org/Part:BBa_K525517 Main Page] - Fusion Protein of BisdA and BisdB (expressed): This fusion protein improves the bisphenol A degradation in E. coli compared to the so far in the partsregistry existing BPA degrading BioBricks.
- [http://partsregistry.org/Part:BBa_K525234 Main Page] - Fusion protein of S-layer CspB and mRFP: This fluorescent S-layer fusion protein is used to characterize the intracellular localisation as well as purification methods for CspB S-layers.
 "
"