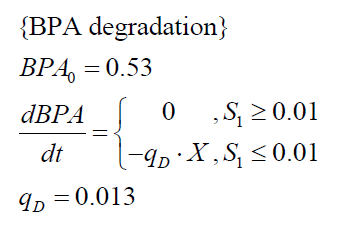Team:Bielefeld-Germany/Modell
From 2011.igem.org

Modelling of intracellular bisphenol A degradation
To model the BPA degradation by E. coli carrying BioBricks for BPA degradation (<partinfo>K123000</partinfo> and <partinfo>K123001</partinfo>) the cell growth has to be described first. The observed growth of E. coli on (our) LB medium was [http://en.wikipedia.org/wiki/Diauxie diauxic] with two different growth phases. Cell growth is a [http://en.wikipedia.org/wiki/First_order_kinetics#First-order_reactions first-order reaction] and is mathematically described as
with the specific growth rate µ and the cell count X. The specific growth rate is dependent on the concentration of the growth limiting substrate (e.g. glucose) and can be described as
with the substrate concentration S, the Monod constant KS and the maximal specific growth rate µmax ([http://www.annualreviews.org/doi/abs/10.1146/annurev.mi.03.100149.002103 Monod, 1949]). Because LB medium is a complex medium we cannot measure the substrate concentration so we have to assume an imaginary substrate concentration. Due to the diauxic growth two different substrates with different Monod constants and consumption rates are necessary to model the cell growth. The amount of a substrate S can be modelled as follows
with the specific substrate consumption rate per cell qS. The whole model for the diauxic growth of E. coli on LB medium with two not measurable (imaginary) substrates looks like:
The specific BPA degradation rate per cell qD is modelled with a Michaelis-Menten like kinetics (Michaelis and Menten, 1913). In the beginning of the cultivations, when E. coli growths on the "good" imaginary substrate S1, no BPA degradation is observed. When this substrate is consumed, the BPA degradation starts. The model for this diauxic behavior is as follows:
The modelling was done with the software [http://www.berkeleymadonna.com/ Berkeley Madonna] using the [http://en.wikipedia.org/wiki/Runge–Kutta_methods#Common_fourth-order_Runge.E2.80.93Kutta_method common fourth-order Runge-Kutta] method to solve the equations. The model was fitted to the measured data by the function "curve fit" in Berkeley Madonna to calculate the constants etc.
Fig. 1 shows a comparison between modelled and measured data for cultivations with BPA degrading E. coli. In Tab. 1 the parameters for the model are given obtained by curve fitting the model to the data.

Tab. 1: Parameters of the model.
| Parameter | K525512 | K525517 |
|---|---|---|
| X0 | 0.112 108 mL-1 | 0.138 108 mL-1 |
| µmax | 1.253 h-1 | 1.357 h-1 |
| KS,1 | 2.646 AU-1 | 1.92 AU-1 |
| KS,2 | 265.1 AU-1 | 103.1 AU-1 |
| S1,0 | 1.688 AU | 1.166 AU |
| qS,1 | 0.478 AU 10-8 cell-1 | 0.319 AU 10-8 cell-1 |
| S2,0 | 16.091 AU | 6.574 AU |
| qS,2 | 0.295 AU 10-8 cell-1 | 0.191 AU 10-8 cell-1 |
| BPA0 | 0.53 mM | 0.53 mM |
| qD,max | 8.77 10-11 mM cell-1 | 1.292 10-10 mM cell-1 |
| KM,BPA | 3.577 10-8 mM-1 | 2.173 10-8 mM-1 |
References
Michaelis L, Menten ML (1912) Die Kinetik der Invertin-Wirkung, Biochemische Zeitschrift 49:333-369.
Monod J (1949) The growth of bacterial cultures, Annu Rev Microbiol [http://www.annualreviews.org/doi/abs/10.1146/annurev.mi.03.100149.002103 3:371-394].
 "
"