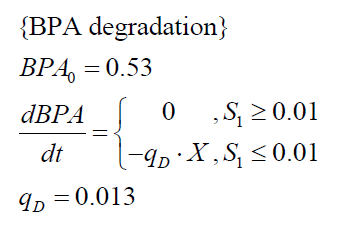Team:Bielefeld-Germany/Results/BPA
From 2011.igem.org
(→References) |
(→Sequencing results) |
||
| Line 3: | Line 3: | ||
==Sequencing results== | ==Sequencing results== | ||
| - | The iGEM team from the University of Alberta sent in BioBricks for BPA degradation in 2008 (<partinfo>K123000</partinfo> and <partinfo>K123001</partinfo>). Sequencing of these BioBricks by the iGEM HQ and by us led to negative results so the sequences entered into the partsregistry are not correct although these sequences are the ones from ''Sphingomonas bisphenolicum'' AO1 (compare [http://www.ncbi.nlm.nih.gov/nuccore/AB255167.1 genbank entry]). In addition, there are "illegal" AgeI and NgoMIV restriction sites in the BioBrick which are used for Freiburg BioBrick assembly standard (RFC 25). After translating and comparing the original sequences from the partsregistry and the sequences from our sequencing results ''in silico'' we saw that the amino acid sequences were identical. These BioBricks were probably synthesized in the Freiburg assembly standard 25 because they have the accordant restriction sites and they were codon optimized for ''Escherichia coli''. But the original sequence from ''S. bisphenolicum'' AO1 was entered into the registry because amino acid sequence of the real sequence and the sequence that was entered are identical. The alignments are shown in figures 1 - 4. | + | The iGEM team from the University of Alberta sent in BioBricks for BPA degradation in 2008 (<partinfo>K123000</partinfo> and <partinfo>K123001</partinfo>). Sequencing of these BioBricks by the iGEM HQ and by us led to negative results so the sequences entered into the partsregistry are not correct although these sequences are the ones from ''Sphingomonas bisphenolicum'' AO1 (compare [http://www.ncbi.nlm.nih.gov/nuccore/AB255167.1 genbank entry]). In addition, there are "illegal" AgeI and NgoMIV restriction sites in the BioBrick which are used for Freiburg BioBrick assembly standard (RFC 25). After translating and comparing the original sequences from the partsregistry and the sequences from our sequencing results ''in silico'' we saw that the amino acid sequences were identical. These BioBricks were probably synthesized in the Freiburg assembly standard 25 because they have the accordant restriction sites and they were codon optimized for ''Escherichia coli''. But the original sequence from ''S. bisphenolicum'' AO1 was entered into the registry because amino acid sequence of the real sequence and the sequence that was entered are identical. The alignments are shown in figures 1 - 4. |
<center><gallery perrow=4> | <center><gallery perrow=4> | ||
Revision as of 23:19, 21 September 2011


Contents |
Sequencing results
The iGEM team from the University of Alberta sent in BioBricks for BPA degradation in 2008 (<partinfo>K123000</partinfo> and <partinfo>K123001</partinfo>). Sequencing of these BioBricks by the iGEM HQ and by us led to negative results so the sequences entered into the partsregistry are not correct although these sequences are the ones from Sphingomonas bisphenolicum AO1 (compare [http://www.ncbi.nlm.nih.gov/nuccore/AB255167.1 genbank entry]). In addition, there are "illegal" AgeI and NgoMIV restriction sites in the BioBrick which are used for Freiburg BioBrick assembly standard (RFC 25). After translating and comparing the original sequences from the partsregistry and the sequences from our sequencing results in silico we saw that the amino acid sequences were identical. These BioBricks were probably synthesized in the Freiburg assembly standard 25 because they have the accordant restriction sites and they were codon optimized for Escherichia coli. But the original sequence from S. bisphenolicum AO1 was entered into the registry because amino acid sequence of the real sequence and the sequence that was entered are identical. The alignments are shown in figures 1 - 4.
Bisphenol A degradation with E. coli
The bisphenol A degradation with the BioBricks <partinfo>K123000</partinfo> and <partinfo>K123001</partinfo> works in E. coli KRX in general. Because [http://onlinelibrary.wiley.com/doi/10.1111/j.1365-2672.2008.03843.x/full Sasaki et al. (2008)] reported problems with protein folding in E. coli which seem to inhibit a complete BPA degradation, we did not cultivate at 37 °C and we did not use the strong T7 promoter as [http://onlinelibrary.wiley.com/doi/10.1111/j.1365-2672.2008.03843.x/full Sasaki et al. (2008)] did for expressing these BioBricks. Therefore we cultivated at 30 °C and we used a medium strong constitutive promoter (<partinfo>J23110</partinfo>). Simultaneously, 30 °C is announced as the cultivation temperature of S. bisphenolicum AO1. With the chosen promoter upstream of a polycistronic bisdAB gene we were able to completely degrade 120 mg L-1 BPA in about 30 - 33 h. By fusing <partinfo>K123000</partinfo> and <partinfo>K123001</partinfo> (using Freiburg BioBrick assembly standard) we could improve the BPA degradation of E. coli even further so that 120 mg L-1 BPA can be degraded in 21 - 24 h. This data is shown in the following figure:
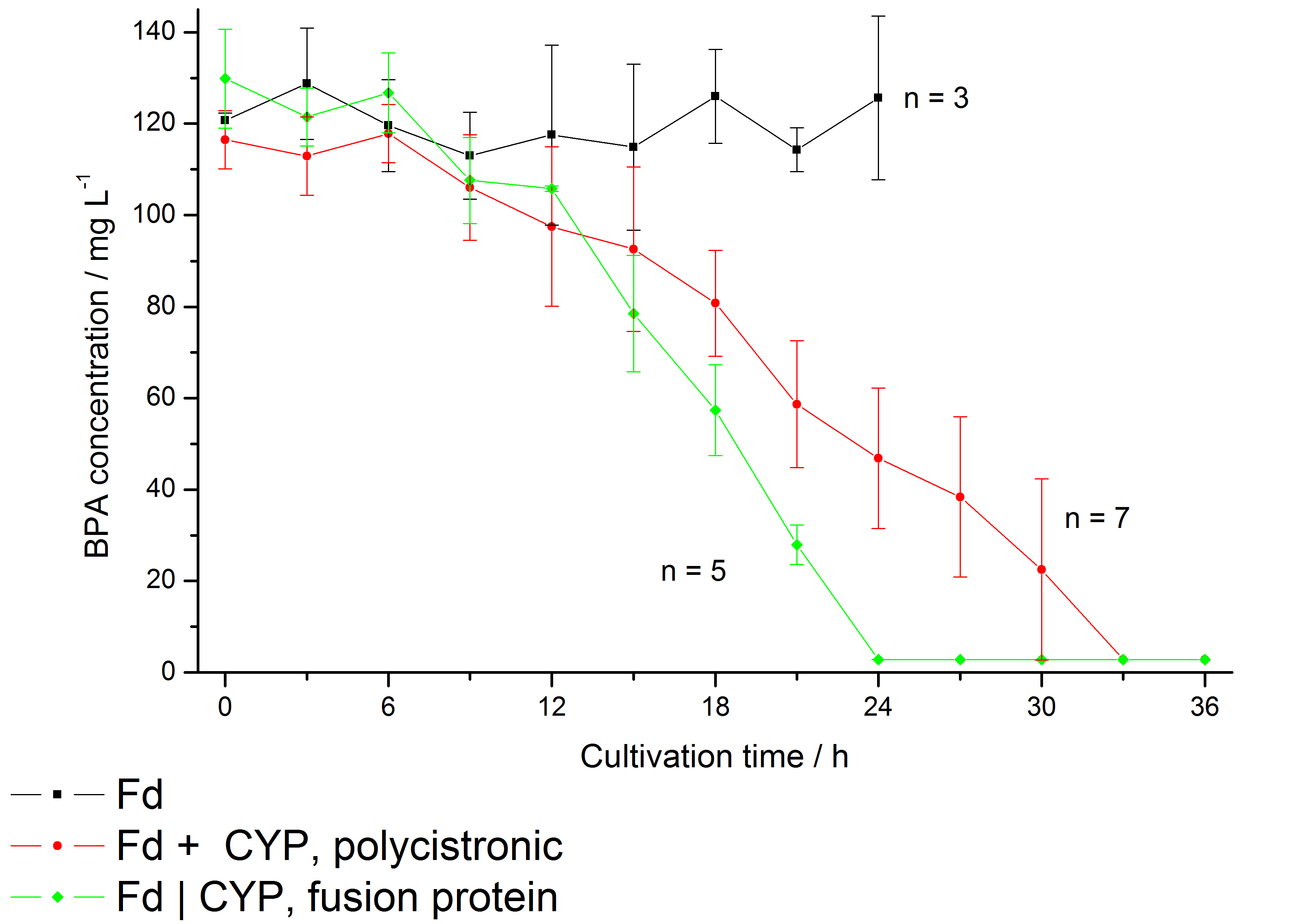
Additionally, we carried out these cultivations at different temperatures and BPA concentrations, but the chosen conditions (30 °C and 120 mg L-1 BPA) seemed to be the best. Higher BPA concentrations had an effect on the growth of E. coli and higher temperature led to a worse BPA degradation (probably due to misfolding of the enzymes). Lower temperature also led to less BPA degradation (probably due to slower growth, expression and reaction rate at lower temperatures). The data dealing with the effect of the temperature on the BPA degradation is shown in fig. 6.
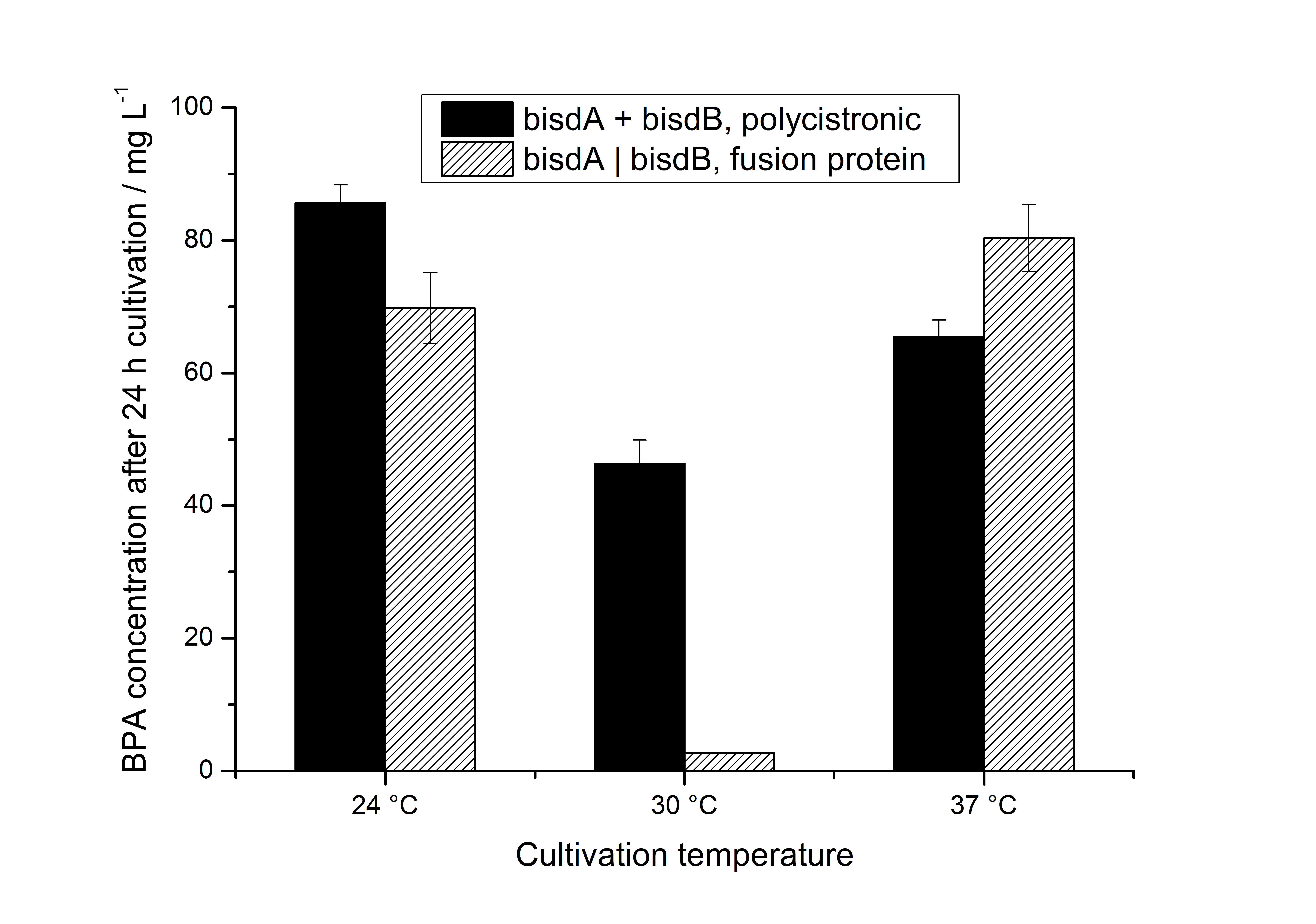
As shown by [http://onlinelibrary.wiley.com/doi/10.1111/j.1365-2672.2008.03843.x/full Sasaki et al. (2008)], BisdB expressed in E. coli leads to hardly no BPA degradation. In our experiments we could not detect the BPA degradation products 1,2-Bis(4-hydroxyphenyl)-2-propanol and 2,2-Bis(4-hydroxyphenyl)-1-propanol in cultivations with E. coli expressing <partinfo>K123000</partinfo> or <partinfo>K123001</partinfo> alone (neither via UV- nor MS-detection). The BPA degradation products 1,2-Bis(4-hydroxyphenyl)-2-propanol and 2,2-Bis(4-hydroxyphenyl)-1-propanol were identified via MS-MS (m/z: 243 / 225 / 211 / 135) and only occured in cultivations with E. coli expressing BisdA and BisdB simultaneously. [http://www.springerlink.com/content/q7864l02734wg32m/ Sasaki et al. (2005)] reported the same MS-MS results for 1,2-Bis(4-hydroxyphenyl)-2-propanol and 2,2-Bis(4-hydroxyphenyl)-1-propanol when degrading BPA with S. bisphenolicum AO1 as we observed in our BPA degradation experiments.
We could also identify the BPA degradation products when working with E. coli TOP10 and MACH1 (data not shown). But because we aim to fuse BisdA and BisdB to S-layer proteins which are supposed to be expressed in E. coli KRX we carried out the characterizations of <partinfo>K123000</partinfo> and <partinfo>K123001</partinfo> in this strain.
Modelling of intracellular bisphenol A degradation
The modelling was done with the software [http://www.berkeleymadonna.com/ Berkeley Madonna] using the [http://en.wikipedia.org/wiki/Runge–Kutta_methods#Common_fourth-order_Runge.E2.80.93Kutta_method common fourth-order Runge-Kutta] method to solve the equations. The model was fitted to the measured data shown above by the function "curve fit" in Berkeley Madonna to calculate the parameters, constants etc.
To model the BPA degradation by E. coli carrying BioBricks for BPA degradation (<partinfo>K123000</partinfo> and <partinfo>K123001</partinfo>) the cell growth has to be described first to calculate a specific BPA degradation rate per cell. The observed growth of E. coli on (our) LB medium was [http://en.wikipedia.org/wiki/Diauxie diauxic] with two different growth phases. Cell growth is a [http://en.wikipedia.org/wiki/First_order_kinetics#First-order_reactions first-order reaction] and is mathematically described as
with the specific growth rate µ and the cell count X. The specific growth rate is dependent on the concentration of the growth limiting substrate (e.g. glucose) and can be described as
with the substrate concentration S, the Monod constant KS and the maximal specific growth rate µmax ([http://www.annualreviews.org/doi/abs/10.1146/annurev.mi.03.100149.002103 Monod, 1949]). Because LB medium is a complex medium we cannot measure the substrate concentration so we have to assume an imaginary substrate concentration. Due to the diauxic growth two different substrates with different Monod constants and consumption rates are necessary to model the cell growth. The amount of a substrate S can be modelled as follows
with the specific substrate consumption rate per cell qS. The whole model for the diauxic growth of E. coli on LB medium with two not measurable (imaginary) substrates looks like:
The specific BPA degradation rate per cell qD is modelled with an equation like eq. (3). In the beginning of the cultivations, when E. coli growths on the "good" imaginary substrate S1, no BPA degradation is observed. When this substrate is consumed, the BPA degradation starts. The model for this diauxic behavior is as follows:
Figure 7 shows a comparison between modelled and measured data for cultivations with BPA degrading E. coli. In Tab. 1 the parameters for the model are given, obtained by curve fitting the model to the data.

Tab. 1: Parameters of the model.
| Parameter | <partinfo>K525512</partinfo> | <partinfo>K525517</partinfo> |
|---|---|---|
| X0 | 0.112 108 mL-1 | 0.138 108 mL-1 |
| µmax | 1.253 h-1 | 1.357 h-1 |
| KS,1 | 2.646 AU-1 | 1.92 AU-1 |
| KS,2 | 265.1 AU-1 | 103.1 AU-1 |
| S1,0 | 1.688 AU | 1.166 AU |
| qS,1 | 0.478 AU 10-8 cell-1 | 0.319 AU 10-8 cell-1 |
| S2,0 | 16.091 AU | 6.574 AU |
| qS,2 | 0.295 AU 10-8 cell-1 | 0.191 AU 10-8 cell-1 |
| BPA0 | 0.53 mM | 0.53 mM |
| qD | 8.76 10-11 mM cell-1 | 1.29 10-10 mM cell-1 |
The specific BPA degradation rate per cell qD is about 50 % higher when using the fusion protein compared to the polycistronic bisdAB gene. This results in an average 9 hours faster, complete BPA degradation by E. coli carrying <partinfo>K525517</partinfo> compared to <partinfo>K525512</partinfo> as observed during our cultivations. The fusion protein between BisdA and BisdB improves the BPA degradation by E. coli.
Interpretation of the results
Misfolding seems to be a problem when expressing BisdA and BisdB in E. coli. To reduce this, the cultivation conditions were improved for the BPA degradation with the polycistronic bisdAB gene in E. coli first, compared to the literature ([http://onlinelibrary.wiley.com/doi/10.1111/j.1365-2672.2008.03843.x/full Sasaki et al., 2008]). Problems with misfolding of BisdA and BisdB could be reduced by lowering the temperature and growth rate and by using a weaker promoter for expression.
When degrading BPA with E. coli using the BisdA | BisdB fusion protein, both domains (BisdA and BisdB) are active and correctly folded because otherwise there would be no BPA degradation measured and no BPA degradation products 1,2-Bis(4-hydroxyphenyl)-2-propanol and 2,2-Bis(4-hydroxyphenyl)-1-propanol could be detected, which could definetely be identified via MS-MS. The about 50 % higher specific BPA degradation rate in E. coli expressing BisdA | BisdB fusion protein could be explained either by improved folding properties of the fusion protein or by the closer distance of BisdA and BisdB in the fusion protein leeding to a faster electron transfer and therefore a more efficient reaction. In this case, which has to be further analysed by cell-free enzyme assays, a natural cytochrome P450 class I electron transport system was converted into a more effective class V electron transport system, demonstrating the new possibilities of synthetic biology. In addition, this new class V system would work without alanine-rich linker, potentially changing the view on cytochrome P450 depending electron transport chains.
Summary of results
Tab. 2: Important parameters of <partinfo>K525512</partinfo> and <partinfo>K525517</partinfo>.
| Experiment | Characteristic | Result K525512 | Result K525517 |
|---|---|---|---|
| Expression in E. coli | Compatibility | E. coli KRX, TOP10, MACH1, BL21(DE3) | |
| Expression | Constitutive | ||
| Optimal temperature | 30 °C | ||
| BPA working concentration | 120 mg L-1 (0.53 mM) | ||
| Purification | Molecular weight | 59.3 kDa | 11.2 and 48.3 kDa |
| Theoretical pI | 4.99 | 4.31 and 5.27 | |
| High absorbtion | 450 nm (due to CYP) | ||
| Degradation of BPA | Completely degradation of 0.53 mM BPA | 21 - 24 h | 30 - 33 h |
| Specific BPA degradation rate | 1.29 10-10 mM cell-1 | 8.76 10-11 mM cell-1 | |
References
Monod J (1949) The growth of bacterial cultures, [http://www.annualreviews.org/doi/abs/10.1146/annurev.mi.03.100149.002103 Annu Rev Microbiol 3:371-394].
Sasaki M, Maki J, Oshiman K, Matsumura Y, Tsuchido T (2005) Biodegradation of bisphenol A by cells and cell lysate from Sphingomonas sp. strain AO1, [http://www.springerlink.com/content/q7864l02734wg32m/ Biodegradation 16(5):449-459].
Sasaki M, Tsuchido T, Matsumura Y (2008) Molecular cloning and characterization of cytochrome P450 and ferredoxin genes involved in bisphenol A degradation in Sphingomonas bisphenolicum strain AO1, [http://onlinelibrary.wiley.com/doi/10.1111/j.1365-2672.2008.03843.x/full J Appl Microbiol 105(4):1158-1169].
 "
"