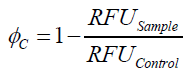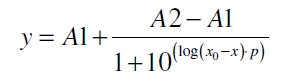Team:Bielefeld-Germany/Results/S-Layer/SgsE
From 2011.igem.org
(→SgsE from Geobacillus stearothermophilus NRS 2004/3a) |
(→Expression in E. coli) |
||
| Line 25: | Line 25: | ||
The SgsE gene under the control of a T7 / lac promoter (<partinfo>K525303</partinfo>) was fused to firefly luciferase (<partinfo>K525999</partinfo>) using Freiburg BioBrick assembly for characterization experiments. | The SgsE gene under the control of a T7 / lac promoter (<partinfo>K525303</partinfo>) was fused to firefly luciferase (<partinfo>K525999</partinfo>) using Freiburg BioBrick assembly for characterization experiments. | ||
| - | The SgsE|luciferase fusion protein was overexpressed in ''E. coli'' KRX after induction of T7 polymerase with 0.2 % L-rhamnose and induction of lac operator with 1 mM IPTG. | + | The SgsE|luciferase fusion protein was overexpressed in ''E. coli'' KRX after induction of T7 polymerase with 0.2 % L-rhamnose and induction of lac operator with 1 mM IPTG. |
| - | The following cultivation was carried out in a [http://www.gmi-inc.com/BioEngineering-KLF-Small-Laboratory-Fermenter.html#product_desc Bioengineering KLF] bioreactor with Bioengineering DCU and software. A sequencer which automatically pumped an inducer solution after 4 h cultivation time to start protein expression was implemented. Other parameters were: | + | The following cultivation was carried out in a [http://www.gmi-inc.com/BioEngineering-KLF-Small-Laboratory-Fermenter.html#product_desc Bioengineering KLF] bioreactor with Bioengineering DCU and software. A sequencer which automatically pumped an inducer solution after 4 h cultivation time to start protein expression was implemented. Other parameters were: |
* Medium: [https://2011.igem.org/Team:Bielefeld-Germany/Protocols/Materials#HSG_medium HSG medium] with 20 mg L<sup>-1</sup> chloramphenicol | * Medium: [https://2011.igem.org/Team:Bielefeld-Germany/Protocols/Materials#HSG_medium HSG medium] with 20 mg L<sup>-1</sup> chloramphenicol | ||
| Line 34: | Line 34: | ||
* DO: 60 % airsaturation (controlled with stirrer cascade starting with 200 rpm) | * DO: 60 % airsaturation (controlled with stirrer cascade starting with 200 rpm) | ||
* pH: 7.0 (controlled with 20 % phosphoric acid and 2 M NaOH) | * pH: 7.0 (controlled with 20 % phosphoric acid and 2 M NaOH) | ||
| - | * Antifoam: BASF | + | * Antifoam: BASF Pluronic PE-8100 |
* Induction solution: 0.2 % L-rhamnose and 1 mM IPTG | * Induction solution: 0.2 % L-rhamnose and 1 mM IPTG | ||
| - | The following figure shows the expression of the SgsE | luciferase S-layer fusion protein <partinfo>K525311</partinfo> in ''E. coli'' KRX in HSG medium with autoinduction sequencer as described above. Optical density, activity of the fused luciferase, dissolved oxygen and agitator speed are plotted against the cultivation time. | + | The following figure shows the expression of the SgsE | luciferase S-layer fusion protein <partinfo>K525311</partinfo> in ''E. coli'' KRX in HSG medium with autoinduction sequencer as described above. Optical density, activity of the fused luciferase, dissolved oxygen and agitator speed are plotted against the cultivation time. |
| - | [[Image:IGEM-Bielefeld2011-Cultivation311.jpg|750px|thumb|center|'''Fig. 3: Bioreactor cultivation of ''E. coli'' KRX expressing the fusion protein SgsE :: luciferase <partinfo>K525311</partinfo> under the control of a T7 / lac promoter. Induction with 1 mM IPTG and 0.2 % L-rhamnose after 4 h controlled by sequencer. Cultivation in [http://www.gmi-inc.com/BioEngineering-KLF-Small-Laboratory-Fermenter.html#product_desc Bioengineering KLF] bioreactor with 2.5 L [https://2011.igem.org/Team:Bielefeld-Germany/Protocols/Materials#HSG_medium HSG medium] with 20 mg L<sup>-1</sup> chloramphenicol starting with OD<sub>600</sub> = 0.4. Dissolved oxygen was regulated to 60 % airsaturation (controlled with stirrer cascade starting with 200 rpm) and the pH to 7.0 (controlled with 20 % phosphoric acid and 2 M NaOH). The dissolved oxygen sensor had to be recalibrated after about 140 min. ''']] | + | [[Image:IGEM-Bielefeld2011-Cultivation311.jpg|750px|thumb|center|'''Fig. 3: Bioreactor cultivation of ''E. coli'' KRX expressing the fusion protein SgsE :: luciferase <partinfo>K525311</partinfo> under the control of a T7 / lac promoter. Induction with 1 mM IPTG and 0.2 % L-rhamnose after 4 h controlled by sequencer. Cultivation in [http://www.gmi-inc.com/BioEngineering-KLF-Small-Laboratory-Fermenter.html#product_desc Bioengineering KLF] bioreactor with 2.5 L [https://2011.igem.org/Team:Bielefeld-Germany/Protocols/Materials#HSG_medium HSG medium] with 20 mg L<sup>-1</sup> chloramphenicol starting with OD<sub>600</sub> = 0.4. Dissolved oxygen was regulated to 60 % airsaturation (controlled with stirrer cascade starting with 200 rpm) and the pH to 7.0 (controlled with 20 % phosphoric acid and 2 M NaOH). The dissolved oxygen sensor had to be recalibrated after about 140 min. ''']] |
==Purification of SgsE fusion protein== | ==Purification of SgsE fusion protein== | ||
Revision as of 01:20, 29 October 2011


Contents |
SgsE from Geobacillus stearothermophilus NRS 2004/3a
SgsE monomers are naturally assembled in a lattice with oblique symmetry (p2) (compare figure on the right) exhibiting a well-defined periodicity and distances of 9.4 – 11.6 nm between the proteinaceous subunits. The S-layer protein SgsE of Geobacillus stearothermophilus NRS 2004/3a consists of 903 amino acids, including a 30 amino acid signal peptide (SLH-domain) at the amino-terminus. The carboxy-terminal part of SgsE is the larger part of the protein, encoding the self-assembly information. The protein is formed by the sgsE gene, has a calculated mass of 93.7 kDa and an isoelectric point of 6.1. When isolated, SgsE maintains its ability to self-assemble, and in dependence of salt concentration, duration of dialysis to remove the detergent and its amino acid sequence, it builds up five types of self-assembly products. These products are formed like flat sheets and cylinders ([http://onlinelibrary.wiley.com/doi/10.1002/smll.200700200/abstract Schäffer et al., 2007]).
Expression in E. coli
The sgsE gene under the control of a T7 / lac promoter (<partinfo>K525303</partinfo>) was fused to mCitrine ([http://partsregistry.org/Part:BBa_J18931 BBa_J18931]) using Freiburg BioBrick assembly for characterization experiments.
The SgsE|mCitrine fusion protein was overexpressed in E. coli KRX after induction of T7 polymerase by supplementation of 0.1 % L-rhamnose and 1 mM IPTG using the autoinduction protocol by Promega.

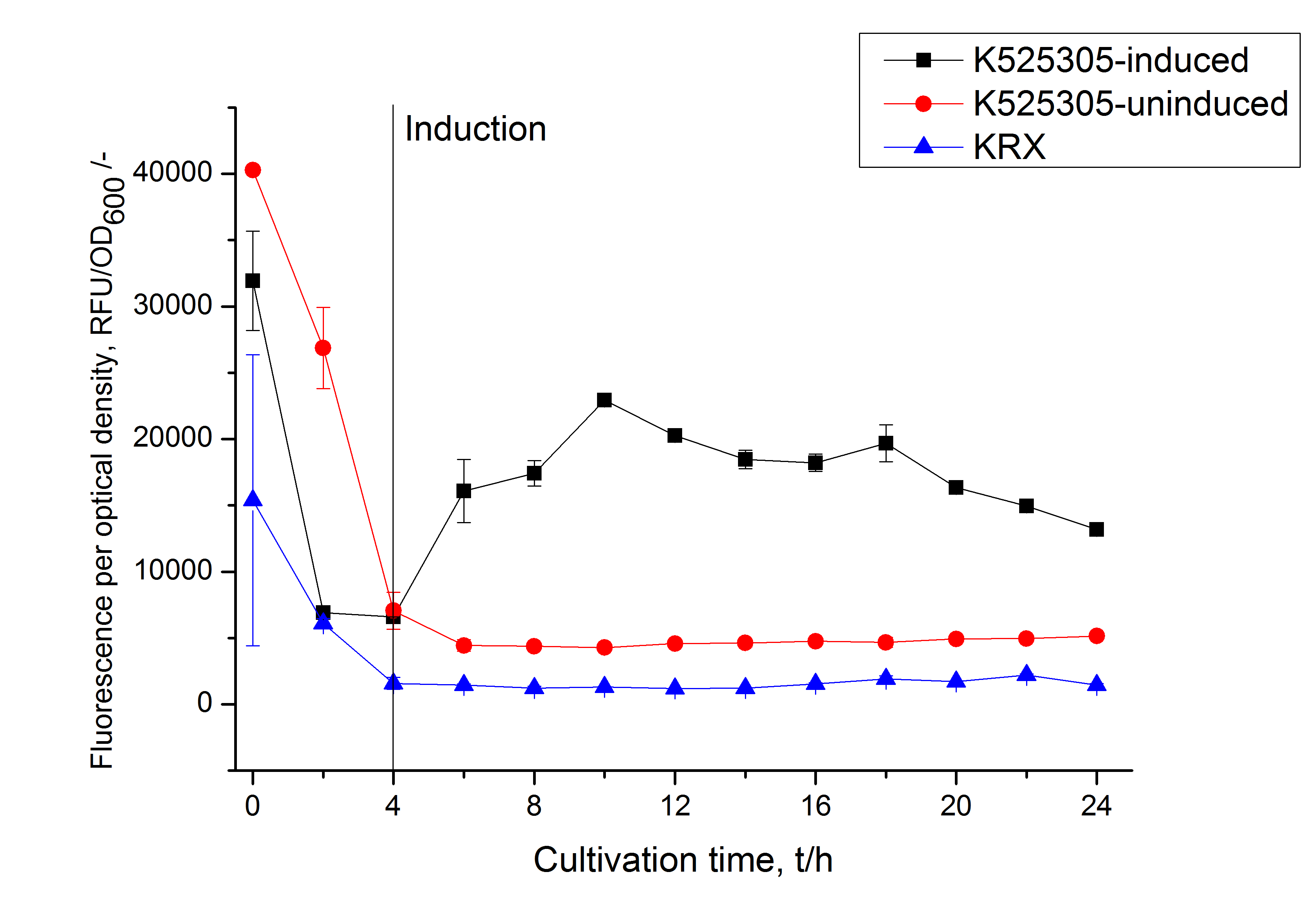
The SgsE gene under the control of a T7 / lac promoter (<partinfo>K525303</partinfo>) was fused to firefly luciferase (<partinfo>K525999</partinfo>) using Freiburg BioBrick assembly for characterization experiments.
The SgsE|luciferase fusion protein was overexpressed in E. coli KRX after induction of T7 polymerase with 0.2 % L-rhamnose and induction of lac operator with 1 mM IPTG.
The following cultivation was carried out in a [http://www.gmi-inc.com/BioEngineering-KLF-Small-Laboratory-Fermenter.html#product_desc Bioengineering KLF] bioreactor with Bioengineering DCU and software. A sequencer which automatically pumped an inducer solution after 4 h cultivation time to start protein expression was implemented. Other parameters were:
- Medium: HSG medium with 20 mg L-1 chloramphenicol
- Culture volume: 2.5 L
- Starting OD600: 0.4
- DO: 60 % airsaturation (controlled with stirrer cascade starting with 200 rpm)
- pH: 7.0 (controlled with 20 % phosphoric acid and 2 M NaOH)
- Antifoam: BASF Pluronic PE-8100
- Induction solution: 0.2 % L-rhamnose and 1 mM IPTG
The following figure shows the expression of the SgsE | luciferase S-layer fusion protein <partinfo>K525311</partinfo> in E. coli KRX in HSG medium with autoinduction sequencer as described above. Optical density, activity of the fused luciferase, dissolved oxygen and agitator speed are plotted against the cultivation time.

Purification of SgsE fusion protein
Purification of SgsE | mCitrine without His-tag
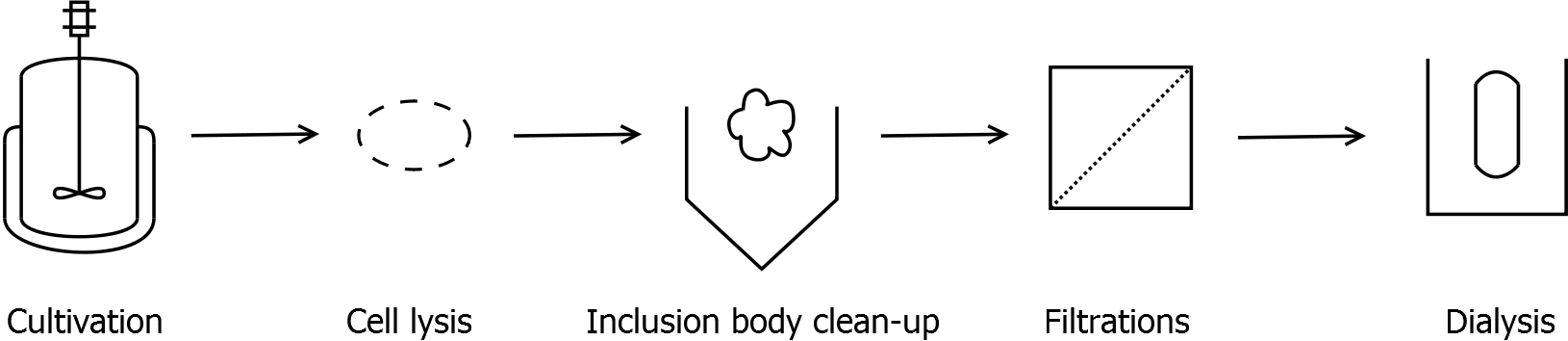
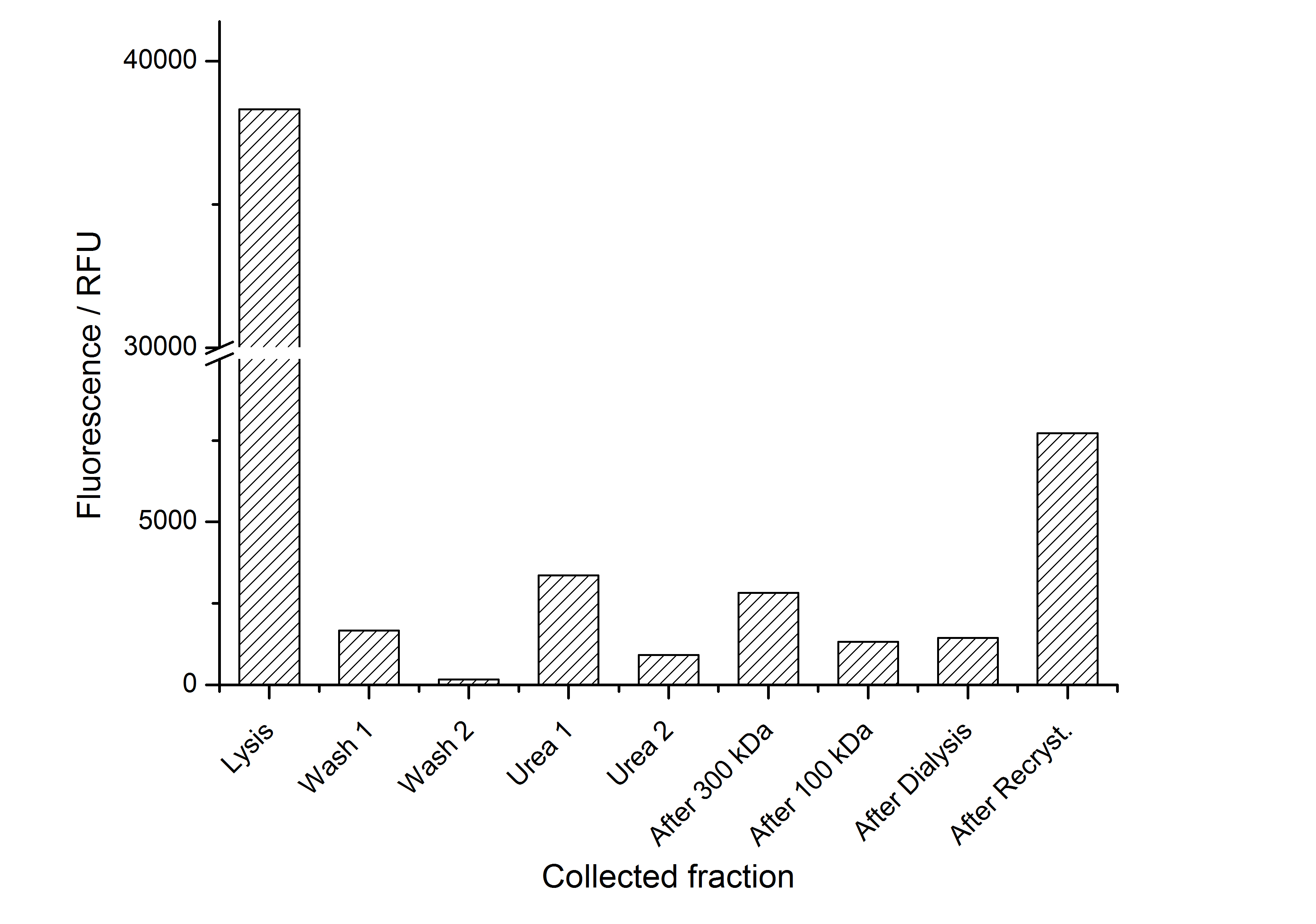
As observed in the analysis of the cultivations with expression of SgsE | mCitrine fusion proteins, these proteins form inclusion bodies in E. coli. Inclusion bodies have the advantage that they are relatively easy to clean-up and are resistant to proteases. So the first purification step is to isolate and solubilize the inclusion bodies. This step is followed by two filtrations (300 kDa UF and 100 kDa DF/UF) to further concentrate and purify the S-layer proteins. After the filtrations, the remaining protein solution is dialyzed against ddH2O for 18 h at 4 °C in the dark. The dialysis leads to a precipitation of the water-insoluble proteins. After centrifugation of the dialysate the water-soluble S-layer monomers remain in the supernatant and can be used for recrystallization experiments.
The fluorescence of the collected fractions of this purification strategy is shown in the following Figure 4:
A lot of protein is lost during the purification especially after centrifugation steps. The fluorescence in the urea containing fractions is lowered due to denaturation of the fluorescent protein. Some fluorescence could be regenerated by the recrystallization in HBSS. This purification strategy is very simple and can be carried out by nearly everyone in any lab being one first step to enable real do it yourself nanobiotechnology.
Purification of SgsE | mCitrine with His-tag
By fusing the SgsE | mCitrine with a C terminal [http://partsregistry.org/Part:BBa_K157011 His-6-tag] the S-layer protein could be simply purified by using a denaturating His-tag affinity chromatography. This purification strategie has the advantage that no time-consuming and complex inclusion body purification and filtration is necessary to decrease the amount of native E. coli proteins. Additional to the simplification a higher purity could of the S-layer protein could be reached.
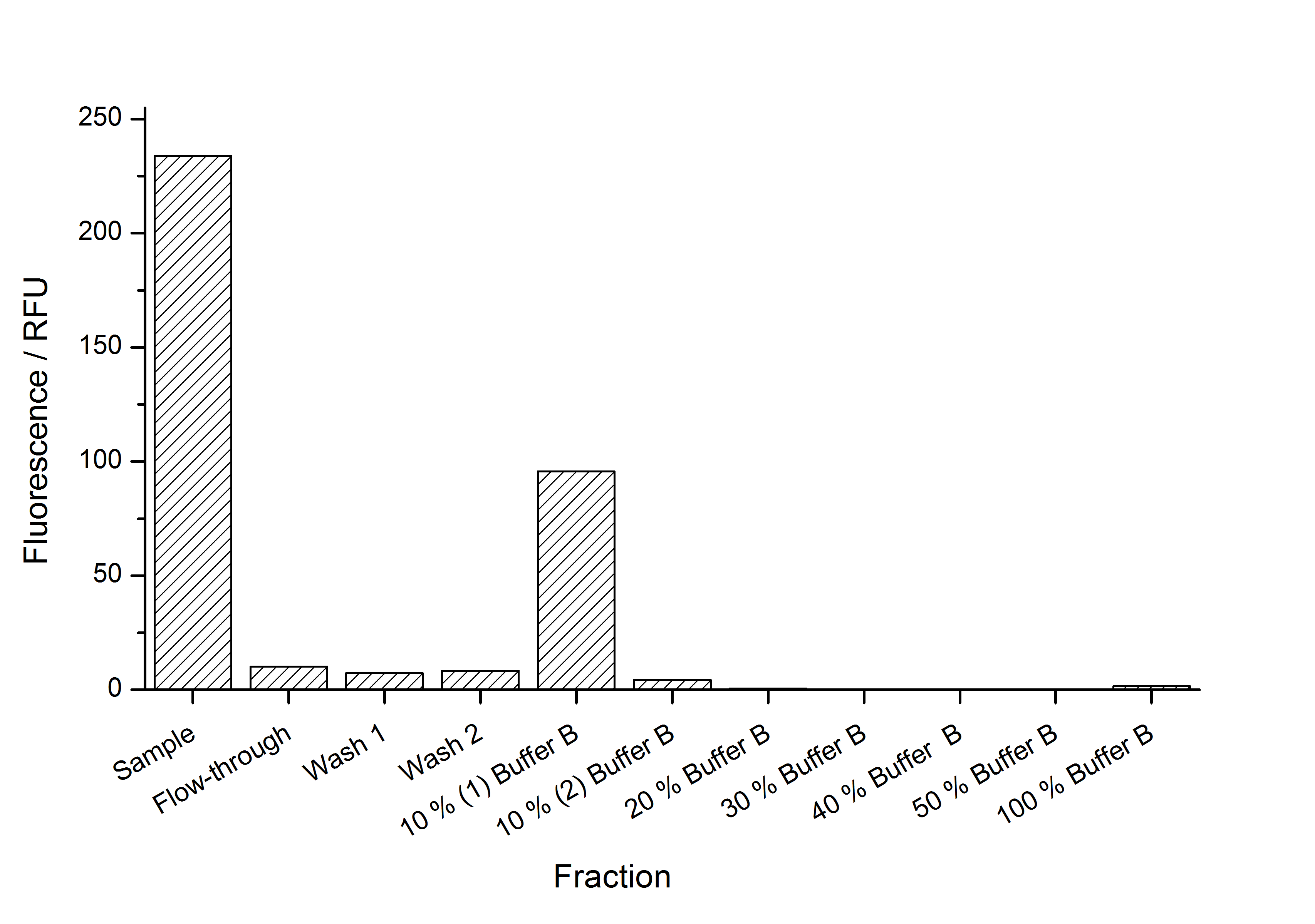
This purification was made using the SgsE fusion protein containing a N-terminal mCitrine,for identification and a C-terminal His-6-tag. The SDS-PAGE gel (Fig. 5) and the fluorescence (Fig. 6) in the collected fractions showed that the majority of fusion protein was eluated with a imidazole concentration of ca. 50 mM. There was also fluorescence measurable in the flowthrough and wash fractions. This indicates that the used 1 mL HisTrap FF crude (GE Healthcare) was overloded or the protein was bound weakly at to the affinity matrix. The resulting purity in the elution fraction, the saving of time and the easiness makes this procedure to the prefered purification method.

Final purification strategies for SgsE | mCitrine
Strategy with His-tag Scheme of purification strategy for SgsE (fusion) proteins with His-tag:
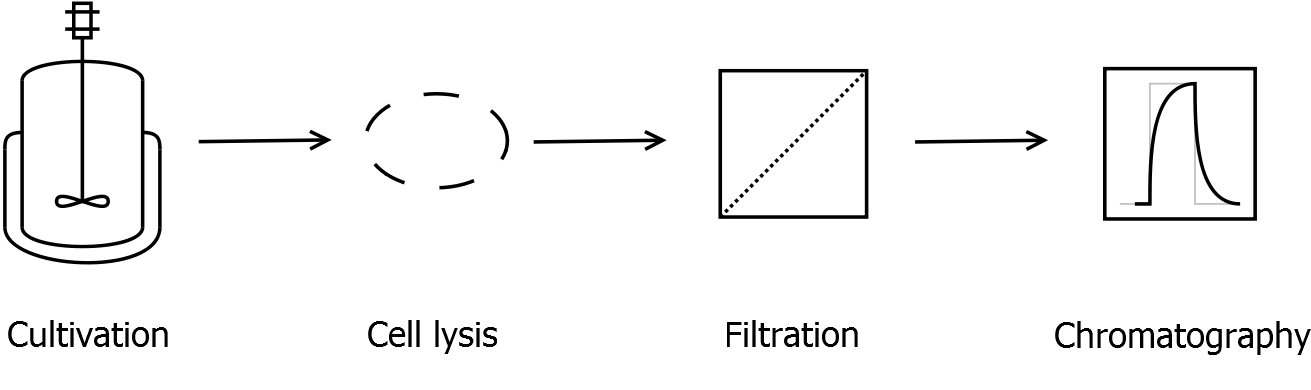
First, SgsE is expressed in E. coli under the control of a T7 / lac promoter for separation of growth and production phase due to metabolic stress of the S-layer expression. Because the SgsE protein is forming inclusion bodies in E. coli, the cells were mechanically disrupted (Sonification on ice) in binding buffer containing 6 M urea. After centrifugation the supernatant is loaded as sample onto a nickel-nitrilotriacetic acid (Ni-NTA) metal-affinity column. The S-layer containing elution fraction of the denaturing His-tag affinity chromatography is afterwards dialysed against water. This leads to the precipitation of water-insoluble proteins. The supernatant contains the monomeric SgsE solution.
Click for detailed information
Strategy without His-tag
Scheme of purification strategy for SgsE (fusion) proteins without His-tag:
First, SgsE is expressed in E. coli under the control of a T7 / lac promoter for separation of growth and production phase due to metabolic stress of the S-layer expression. Because the SgsE protein is forming inclusion bodies in E. coli, an inclusion body purification with urea follows the cell lysis. The S-layers are further concentrated and purified by two ultrafiltration / diafiltration steps (300 kDa and 100 kDa) and afterwards dialysed against water leading to the precipitation of water-insoluble proteins. The supernatant contains the monomeric SgsE solution.
Click for detailed information
Purification of SgsE | luciferase fusion protein
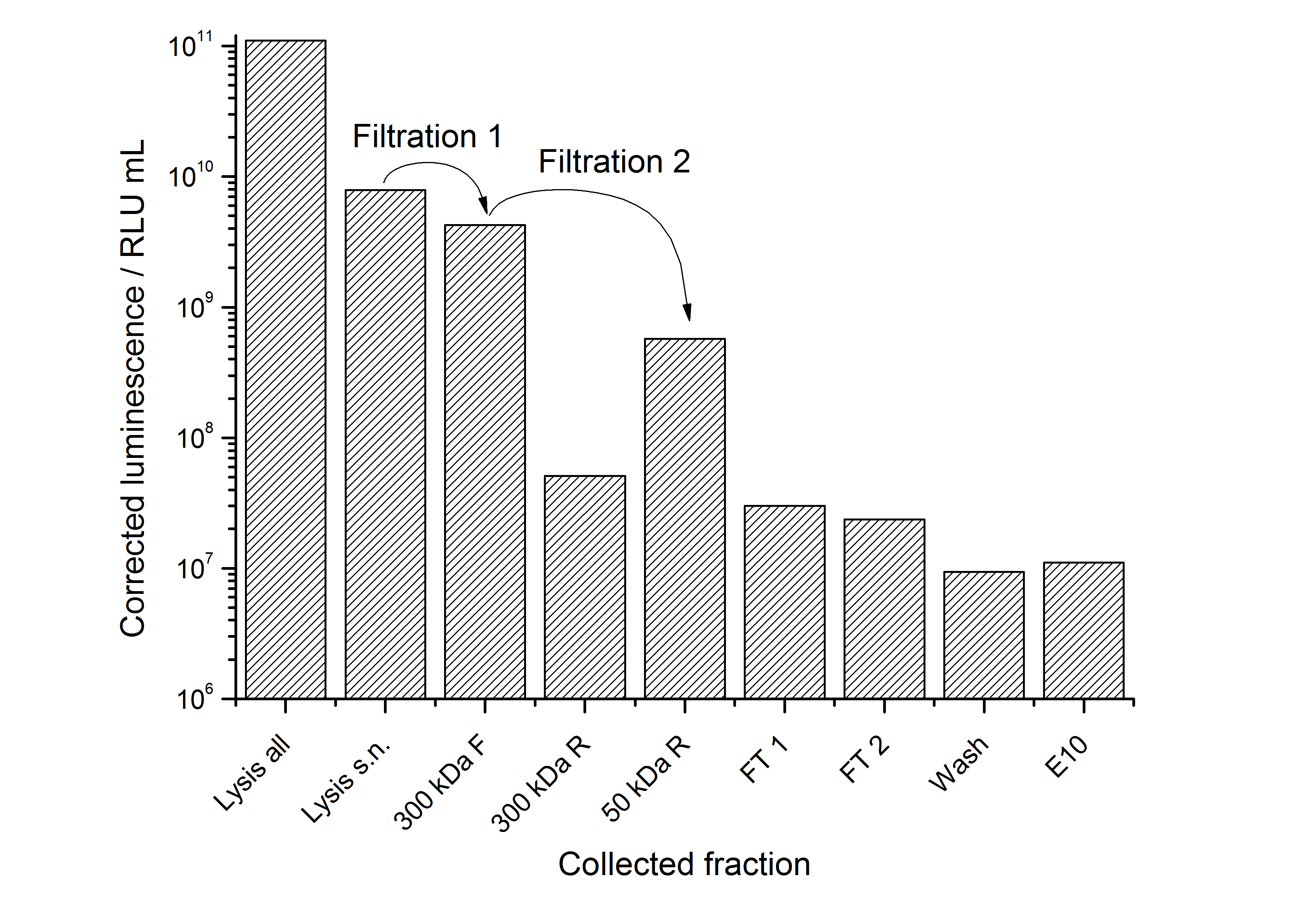
After the analysis of cultivations with expression of SgsE | luciferase fusion proteins, different cell fractions were analyzed. It could be seen that the proteins form inclusion bodies in E. coli but that there are some soluble proteins left. This has the advantage that the proteins carrying an enzyme as fusion proteins do not have to be treated with denaturating agents like urea which destroys the enzyme (data not shown).
To capture the protein from the cell lysate an ion exchange chromatography (IEX) was carried out (binding with pH 7.0, 25 mM NaCl, quaternary amine beads, elution with 100 mM NaCl). A lot of protein was found in the flow-through. When concentrating and rebuffering the proteins with PES (polyethylene sulfone) membranes a lot of protein was lost. The S-layer proteins stuck to the membrane. Some could be removed again from the membrane after cutting out the filter and incubate it in ddH2O over-night. This problem has to be kept in mind when using this S-layer.
The results of the purification approach are shown in Fig. 7:
The purification strategy has to be improved. The inclusion bodies cannot be purified because urea damages the luciferase irreversible (data not shown). The loss due to adsorption of the SgsE | luciferase fusion protein to PES membranes could be avoided by using different membranes. The binding conditions of the IEX have to be improved as well. Anyway, the idea behind this purification strategy could be a starting point for a better strategy. Possibilities for improvement are:
- Different membranes for ultra- / diafiltration
- Other binding conditions for the IEX capturing step (higher pH)
- Hydrophobic interaction chromatography as purification step after IEX (works in general, data not shown)
- Size exclusion chromatography (SEC) for polishing
Immobilization behaviour
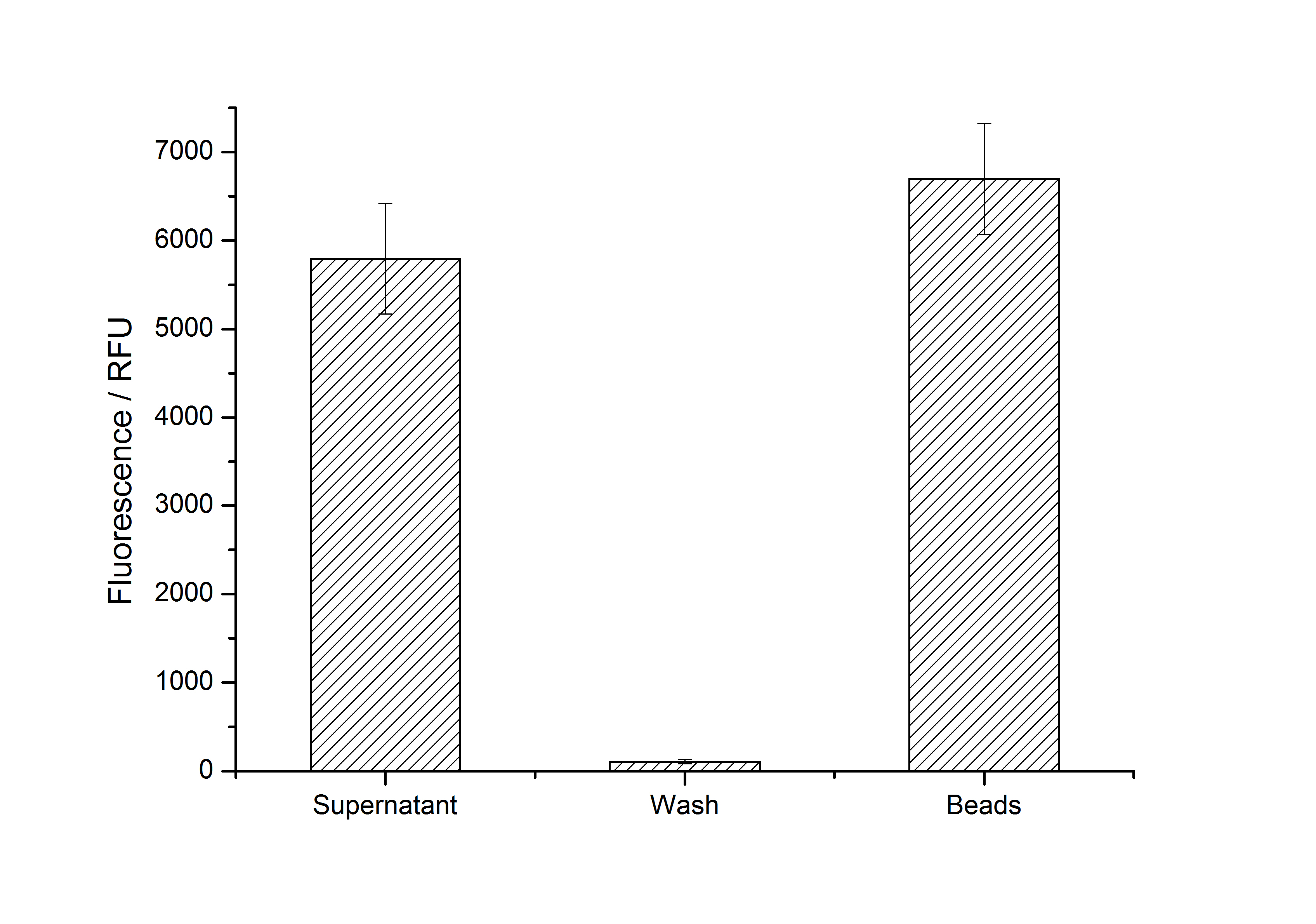
After purification, solutions of monomeric SgsE S-layer proteins can be recrystallized and immobilized on silicon dioxide beads in HBSS (Hank's buffered saline solution). After the recrystallization procedure the beads are washed with and stored in ddH2O at 4 °C in the dark. The fluorescence of the collected fractions of a recrystallization experiment with <partinfo>K525305</partinfo> are shown in Fig. 8. 100 mg beads were coated with 100 µg of protein. The figure shows, that not all of the protein is immobilized on the beads (supernatant fraction) but the immobilization is pretty stable (very low fluorescence in the wash). After the immobilization, the beads show a high fluorescence indicating the binding of the SgsE | mCitrine fusion protein.
Optimal bead to protein ratio for immobilization
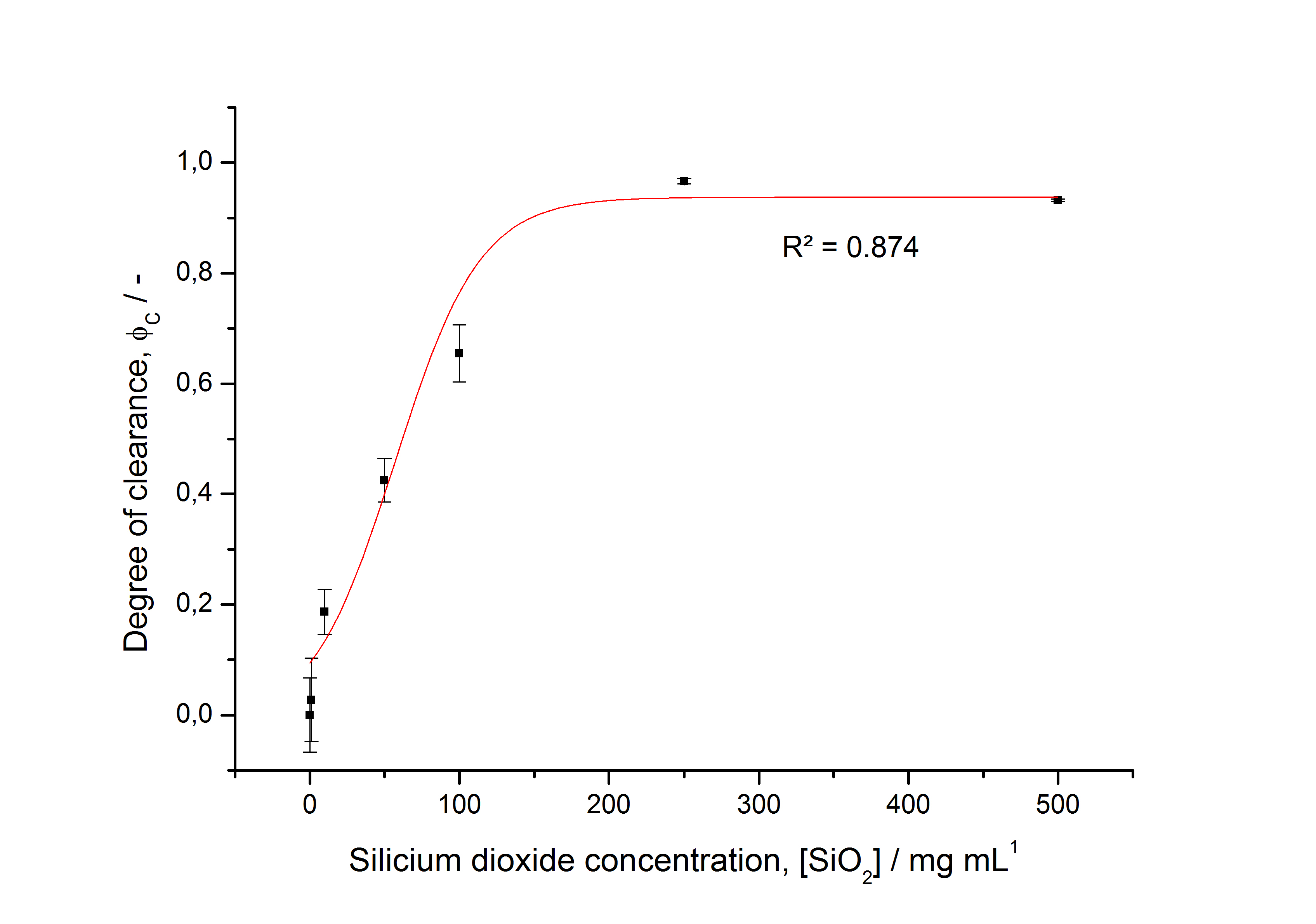
To determine the optimal ratio of silica beads to protein for immobilization, the degree of clearance ϕC in the supernatant is calculated and plotted against the concentration of silica beads used in the accordant immobilization experiment (compare Fig. 9):
The data was collected in three independent experiments. The fluorescence of the samples was measured in the supernatant of the immobilization experiment after centrifugation of the silica beads. The fluorescence of the control was measured in a sample which was treated exactly like the others but no silica beads were added. 100 µg protein was used for one immobilization experiment. The data was fitted with a sigmoidal dose-response function of the form
with the Hill coefficient p, the bottom asymptote A1, the top asymptote A2 and the switch point log(x0) (R² = 0.874).
The fit indicates that a good silica concentration for 100 µg of protein is 150 - 200 mg mL-1. This set-up leads to saturated beads with low waste of protein. So a good protein / bead ratio to work with is 5 - 7 * 10-4.
Effect on enzyme stability
To test whether the S-layer protein enhances the half-life of the firefly luciferase, immobilization experiments were carried out. After the IEX described above, the elution fraction was immobilized on silicon dioxide beads or just diluted with HBSS buffer which is used for the immobilization / recrystallization of SgsE S-layer proteins. The immobilization is carried out at room temperature for 4 h. It could be seen that the luciferase activity nearly expired during this time (in the positive and the negative control). So the S-layer SgsE could not stabilize the luciferase at room temperature. The results of this experiment is shown in the figure below:
 "
"