Team:ETH Zurich/Biology/Validation
From 2011.igem.org
(→References) |
(→Pu promoter) |
||
| Line 81: | Line 81: | ||
| - | |||
== References == | == References == | ||
<span id="Ref1">[1] [http://www.ncbi.nlm.nih.gov/pubmed/11550794 Beatrice Felenbok, Michel Flipphi and Igor Nikolaev: '''Ethanol Catabolism in ''Aspergillus nidulans'': A Model System for Studying Gene Regulation, Progress in Nucleic Acid Research and Molecular Biology''', 69: 149-204]</span> | <span id="Ref1">[1] [http://www.ncbi.nlm.nih.gov/pubmed/11550794 Beatrice Felenbok, Michel Flipphi and Igor Nikolaev: '''Ethanol Catabolism in ''Aspergillus nidulans'': A Model System for Studying Gene Regulation, Progress in Nucleic Acid Research and Molecular Biology''', 69: 149-204]</span> | ||
Revision as of 20:50, 3 October 2011
| Validation |
| |||
| Here we describe the experimental setups used to test the functionality of the parts: | ||||
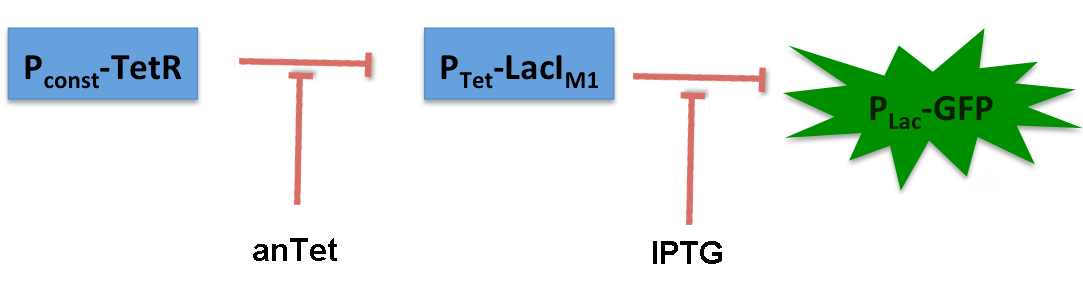
AlcR sensorTest systemAs described in the Smoke detectors section we designed two artificial AlcR-dependent promoters which get repressed upon binding of AlcR to acetaldehyde. For testing of our design we created a system with a sfGFP output in order to characterize the gene expression with different levels of AlcR production and acetaldehyde induction. The system works in the following way (see Figure 1): A) In normal medium conditions the TetR transcription factor is produced and binds to its cognate promoter, thus inhibiting the expression of alcR. In this case no AlcR is present and GFP is produced. B) Upon addition of increasing amounts of anhydrotetracycline, TetR gets released from the promoter and AlcR is produced. The amount of AlcR should increase with increasing amounts of anhydrotetracycline. C) By addition of acetaldehyde AlcR binds to the alc-promoter and the GFP response decreases.
Experiments and resultsIn a first experiment, the alc-promoters were tested by using a non-codon optimized natural variant of alcR. Because of this we tested the compatibility of the codon usage in the variant from Aspergillus nidulans with the one from E. coli. We received a codon adaptation index (CAI) of 0.7 [3]. Nevertheless, five rare arginine codons in E.coli were present within the first 20 amino acids of the gene. After a first experiment without any significant results (data not shown), the first three rare arginine codons of alcR were exchanged by PCR and a fully codon-optimized variant was ordered. All of the following experiments were performed with alcR codon-optimized within the first 20 amino acids. The AlcR test was established as a two plasmid system containing ampicillin and kanamycin resistances and pMB1 respectively pSC101 origins of replication. These plasmids were co-transformed into E. coli JM101 strain. M9 medium was inoculated with an overnight culture of the strain. The bacteria were induced with different amounts of acetaldehyde (0, 10, 1000 µM) and anhydrotetracycline (0, 1, 100 ng/mL) during the exponential growth phase. In order to avoid evaporation of acetaldehyde, all equipment was cooled down to -20 degrees prior to preparation of the stock solutions. All solutions were afterwards prepared in tubes at 4°C. Finally, the cell suspension was added to the solutions and the tubes were tightly closed to prevent loss of acetaldehyde. Fluorescence and OD measurements were performed in a 96-well plate several hours after induction. 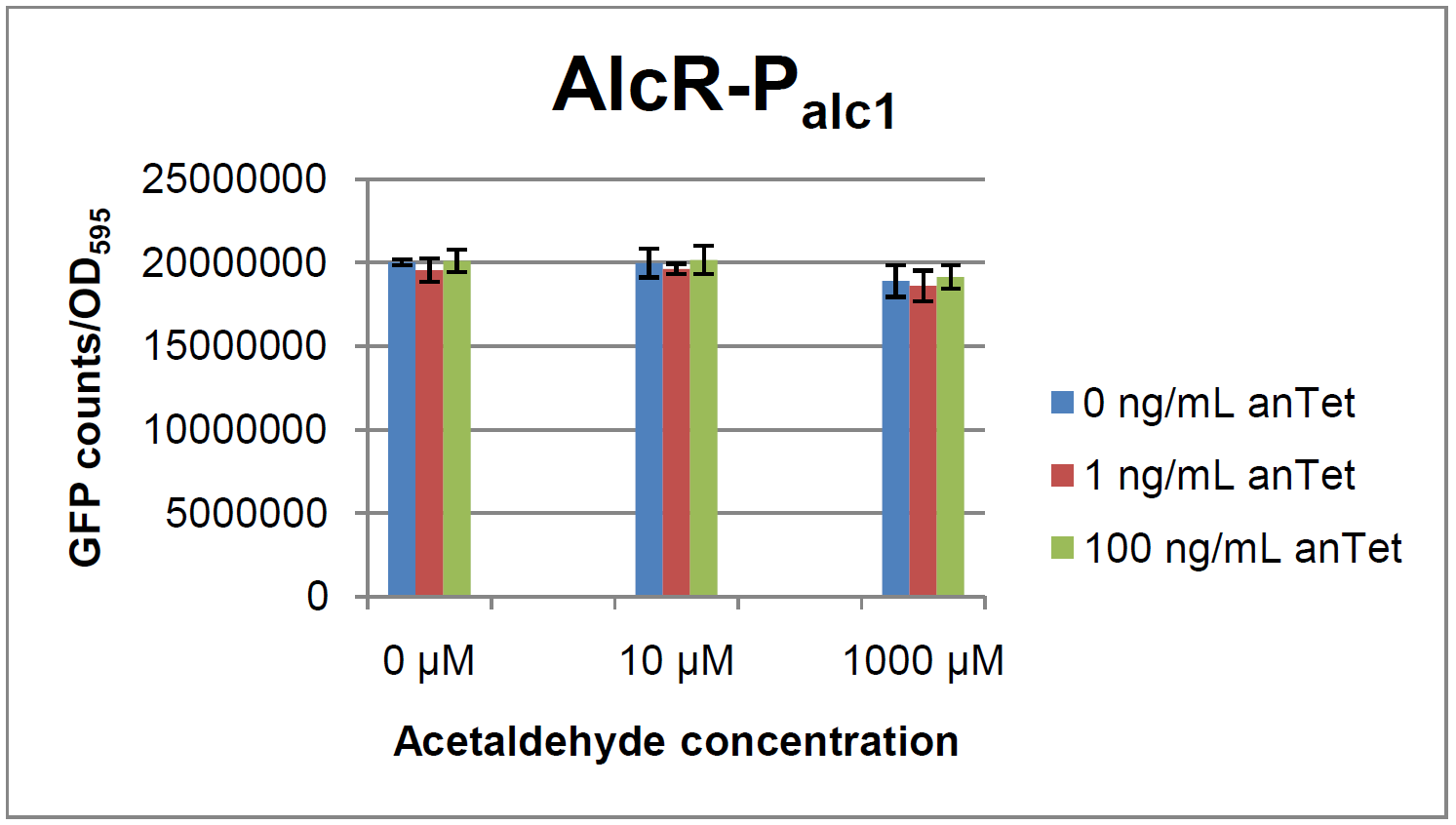 Figure 2: Test results for Palc1 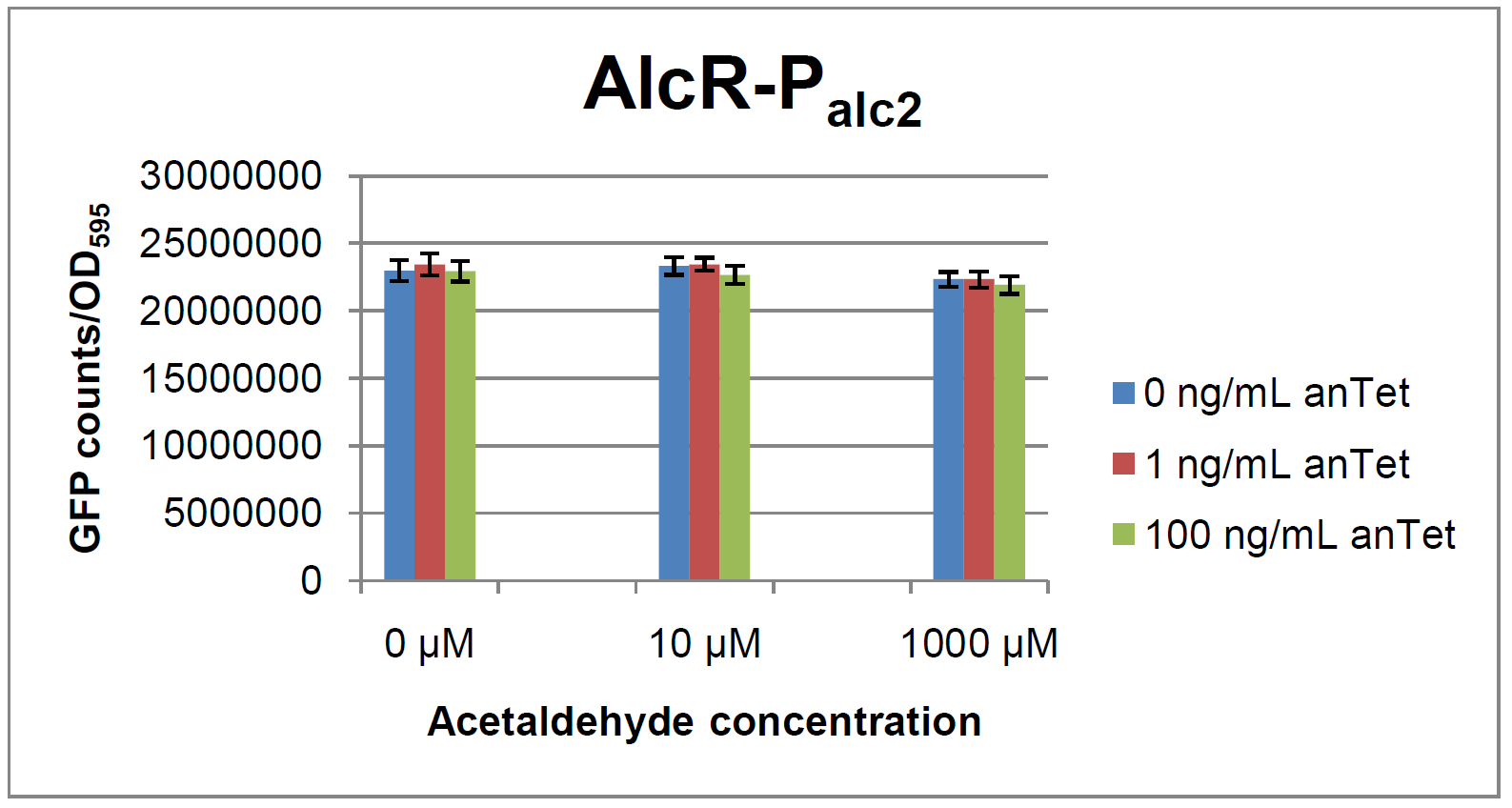 Figure 3: Test results for Palc2
As visible on the two images above, no significant change in GFP fluorescence could be observed neither between different acetaldehyde concentrations nor between different AlcR induction levels. Because of these negative results and the non codon-optimized alcR, we performed expression tests for the protein in E. coli strain JM101. In these experiments, gene expression was induced with different amounts of anhydrotetracycline during the exponential growth phase of the bacteria. After cultivation, cells were harvested and lysed either by using a Retsch mill or by lysozyme. After resuspension of the cell extract we performed both SDS-PAGE gels and western blots to check for alcR expression. Buffer was added to the cell debris and heated to 95°C for 10 minutes in order to resolubilize potential inclusion bodies. AlcR has a molecular mass of about 96 kDa and thus its band was suspected to be on the same level as the red box marked in the SDS-PAGE gel (see Figure 4) [2]. Despite increasing amounts of anhydrotetracycline (wells 1 and 3: 0 ng/mL, 2 and 4: 1 ng/mL, 3 and 6: 100 ng/mL), no change could be observed in this area of the gel for any of the probes. A reason for this could be that alcR is not expressed at all. To check again whether alcR is expressed at all we performed a western blot. Therefore, a 6x His-tag was added to the C-terminal end of AlcR and we used an anti-His antibody kit in order to detect it. A probe with another concentrated 6x His-tagged protein was added as positive control so that we could exclude systematic errors during the blotting procedure (see Figure 5). Because we could not show any expression of alcR at all we suppose that there is no AlcR produced in the cells, probably due to the usage of rare codons in the gene. Due to a long delay in the synthesis of the codon-optimized part, we were not able to test our system with this variant until now. |
Xylene degradationBy the upper tol pathway indole is metabolized to indigo which is a blue pigment. To show that the upper tol pathway is active cells were incubate on LB plates containing 1 mM indole for 6 h at 37 °C. A atmosphere of m-xylene is created in the plate to induce PU promoter. If colonies became blue this is an indication for an active upper tol pathway. |
BandpassCharacterization of lacIM1LacIM1 (Bba_K62500) is a LacI variant with the same protein sequence as the wild-type LacI, but with a completely codon-modified DNA sequence. This enables to use both sequences on two different plasmids (with two different promoters) without recombination happening between them, which is necessary for in our system to establish the bandpass filter. By entering this BioBrick into the registry, we wanted to check its functionality. Thus we engineered a test system for our BioBrick by adding a lac-dependent GFP reporter. A tet-promoter was put in front of lacIM1 and TetR constitutively produced, thus preventing lacIM1 from expression. In the non-induced state of the system, lacIM1 expression is repressed and the cells produce GFP. If anhydrotetracycline is added, TetR is released from Ptet and LacI is produced. LacI inhibits gfp transcription and the fluorescence decreases. Addition of IPTG although leads to the opposite case: LacI gets inhibited and the fluorescence increases. The test system for Bba_K62500 was established as a single plasmid system. In the experiments we used the two different plasmids pSB1AK3 and pSB3C5 as backbones of the designed network. Experiments were performed in E.coli strain JM101, which was grown in M9 medium until exponential growth and then induced with increasing amounts of anhydrotetracyline (0 ng/mL, 1 ng/mL, 100 ng/mL) and IPTG (0 µM, 1 µM, 10 µM, 100 µM, 1000 µM). The cells were incubated in a 96-well plate at 37°C and every 15 minutes OD600 and GFP fluorescence was measured. The results for the characterization of lacIM1 can be found in the Experimental results section. |
|
References[1] [http://www.ncbi.nlm.nih.gov/pubmed/11550794 Beatrice Felenbok, Michel Flipphi and Igor Nikolaev: Ethanol Catabolism in Aspergillus nidulans: A Model System for Studying Gene Regulation, Progress in Nucleic Acid Research and Molecular Biology, 69: 149-204] [2] [http://www.ncbi.nlm.nih.gov/pubmed/3072264 Beatrice Felenbok et al. The ethanol regulon in Aspergillus nidulans: characterization and sequence of the positive regulatory gene alcR. Gene. 1988 ;73(2):385-96.] [3] [http://openwetware.org/wiki/M9_medium/minimal OpenWetWare contributors, M9 medium/minimal 20 Apr 2007, 17:20 UTC. 21 Sep 2011, 17:51] |
 "
"