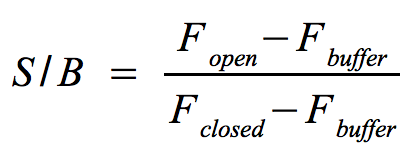Team:Bielefeld-Germany/Protocols/Downstream-processing
From 2011.igem.org
| Line 99: | Line 99: | ||
===His-tag affinity chromatography=== | ===His-tag affinity chromatography=== | ||
| + | |||
| + | ==Bradford Protein Assay== | ||
==Measuring of [http://partsregistry.org/Part:BBa_E1010 mRFP]== | ==Measuring of [http://partsregistry.org/Part:BBa_E1010 mRFP]== | ||
Revision as of 11:57, 10 September 2011

Production Protocols: These are the protocols for the cultivation and the downstream processing.
Cultivation
Expression of [http://expasy.org/sprot/hamap/CORGL.html Brevibacterium flavum] and [http://ijs.sgmjournals.org/cgi/content/abstract/54/3/779 Corynebacterium halotolerans] S-layer genes
Expression of bisphenol A degrading BioBricks in E. coli

- Used BioBricks: <partinfo>K525502</partinfo>, <partinfo>K525512</partinfo>, <partinfo>K525517</partinfo>, <partinfo>K525552</partinfo>
- Chassis: Promega's [http://www.promega.com/products/cloning-and-dna-markers/cloning-tools-and-competent-cells/bacterial-strains-and-competent-cells/single-step-_krx_-competent-cells/ E. coli KRX]
- Medium: LB medium supplemented with 100 mg L-1 Ampicillin and 120 mg L-1 bisphenol A
- BPA is thermally stable -> you can autoclave it together with the medium
- 100 mL culture in 300 mL shaking flask without baffles (Schott) with silicon plugs
- Cultivation temperature: 24 °C, 30 °C or 37 °C, tempered with Infors AG AQUATRON at 120 rpm
- for characterizations: automatic sampling every three hours with Gilson fraction controller F2XX cooled (< 4 °C) with Julabo F10 water bath BURMA-SHAVE!!
- the characterization experiment setup is shown on the picture on the right
Purification methods
Release of periplasmic protein fraction from E. coli by cold osmotic shock
Modified protocol from [http://www.jbc.org/content/240/9/3685.full.pdf+html?sid=4a90c176-0ec3-489f-8c82-4734274cebf5 Neu & Heppel, 1965].
- Centrifuge E. coli cell suspension for 5 min at 14000 g (4 °C) to collect the cells.
- Discard the entire supernatant.
- Resuspend the cells ice-cold cell fractionating buffer #1. The resulting volume should be 1/4 of the former suspension volume.
- Incubate for 20 min on ice. Ivert the suspension at regular intervals to counteract sedimentation.
- Centrifuge the cell suspension for 15 min at 14000 g (4 °C).
- Discard the entire supernatant.
- Resuspend the cells ice-cold cell fractionating buffer #2. The resulting volume should be 1/4 of the former suspension volume.
- Incubate for 10 min on ice under regular invertion.
- Centrifuge the cell suspension for 15 min at 14000 g (4 °C).
- Save the supernatant, which contains the periplasmatic proteins.
- If the periplasmatic protein fraction is turbid, re-centrifuge and filter it through a 0.2 µm filter.
Inclusion body clean-up
- harvest the cells by centrifugation (30 min, 10,000 g, 4 °C)
- resuspend pellet and disrupt cells
- centrifuge lysate (60 min, >17,000 g, 4 °C)
- wash pellet at least two times with water to remove water-soluble proteins
- after washing the pellet: incubate the pellet in denaturation buffer for 60 min, 4 °C with vertical rotator
- final concentration in denaturation buffer: 0.5 mg wet biomass per mL
- centrifuge (60 min, >17,000 g, 4 °C)
- the higher the speed, the better the result
- collect supernatant and incubate the pellet again in denaturation buffer (60 min, 4 °C, vertical rotator)
- centrifuge (60 min, >17,000 g, 4 °C)
- collect supernatant and discard pellet
Ultra-/Diafiltration

- Milipore ... XL 50 and 100 kDa membranes
- pump
- bla bla
Ion exchange chromatography (IEX) for S-layer proteins from Corynebacterium
- used column: DEAE HiTrap 1 mL with GE Healthcare ÄKTA
- flow rate: 1 mL min-1
- equilibrate column with 5 column volumes of binding buffer
- inject sample and wash column with 5 column volumes of binding buffer
- elute S-layer with 40 % binding / elution buffer mix
- elute remaining proteins with 100 % elution buffer
Ni-NTA spin columns
denaturating
non-denaturating
His-tag affinity chromatography
Bradford Protein Assay
Measuring of [http://partsregistry.org/Part:BBa_E1010 mRFP]
- Take at least 500 µL sample for each measurement (200 µL is needed for one measurement) so you can perform a repeat determination
- Freeze samples at -80 °C for storage
- To measure the samples thaw at room temperature and fill 200 µL of each sample in one well of a black, flat bottom 96 well microtiter plate (perform at least a repeat determination)
- Measure the fluorescence in a platereader (we used a [http://www.tecan.com/platform/apps/product/index.asp?MenuID=1812&ID=1916&Menu=1&Item=21.2.10.1 Tecan Infinite® M200 platereader]) with following settings:
- 20 sec orbital shaking (1 mm amplitude with a frequency of 87.6 rpm)
- Measurement mode: Top
- Excitation: 584 nm
- Emission: 620 nm
- Number of reads: 25
- Manual gain: 100
- Integration time: 20 µs
Sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE)
This analytical Method can be used for separation and identification of proteins according to their electrophoretic mobility. The mobility is a function of length of the molecular weight. Proteins that have identical charge per unit mass due to binding of SDS results in an equal electrophoretic mobility.
Pouring the polyacrylamide gel
- Make a master mix for the stacking and separating gel without adding ammonium persulfate and TEMED.
- Aliquote 6,5 mL for each separating and 2,5 mL for each stacking gel.
- Add ammonium persulfate and TEMED to each separating gel aliquote and pour the solution quickly into your gel casting form. Leave about 2 centimeters below the bottom of the comb for the stacking gel.
- Layer isopropanol on top of the gel.
- Leave the separating gel at room temperature for >60 minutes to polymerize.
- Remove isopropanol and wait until the surface is dry.
- Add ammonium persulfate and TEMED to each separating gel aliquote and pour the solution quickly into your gel casting form.
- Insert comb without getting bubbles stuck underneath
- Leave the gel at room temperature for >60 minutes to polymerize.
- For storage
- Remove sealing and store the gel wrapped in moistened paper towel at 4°C.
Preparing the sample
- Mix your protein mixture 4:1 with Laemmli-buffer (30 mL protein solution + 10 mL Laemmli-buffer)
- Heat for 5 minutes at 95 °C.
Running the gel
- Remove sealing, put the polymerized gel into gel box and pour SDS running buffer into the negative and positive electrode chamber.
- Remove comp without destroying the gel pocket.
- Pipet the sample into the gel pockets, adjusting the volume according to the amount of protein in your sample. Make sure to include a lane with molecular weight standards (PageRuler Prestained Protein Ladder™ (Fa. Fermentas)) to determinate the molecular weight of your sample.
- Connect the power lead and run the stacking gel with 10 mA until the blue dye front enters the separating gel.
- Raise amperage up to 20 mA for running the separating gel.
- When the distance of the lowest molecular weight standard lane to the gel end is down to 0,5 cm stop the electrophoresis by turning off the power supply.
Polyacrylamide gel staining
Colloidal Coomassie Blue staining
Modified staining protocol from [http://http://newjournal.kcsnet.or.kr/main/j_search/j_abstract_view.htm?code=B021105&qpage=j_search&spage=b_bkcs&dpage=ar Kang et al., 2002].
- agitate the staining solution at 37 °C over night to form the colloids
- After finishing the SDS-PAGE remove gel from gel casting form and tranfer it in to a box.
- Add 100 mL of the stainig solution to your polyacrylamid gel.
- Incubate the gel in the solution at room temperature until the protein bands got an intensive blue color. Shake the gel continuously during incubation.
- Remove the staining solution
- wash the gel with dH2O
- Incubate the gel in ddH2O (2-6h) for bleaching the background. Shake the gel continuously during incubation. If necessary replace the colored water with new one.
Silver staining
Modified staining protocol from
- After finishing the SDS-PAGE remove gel from gel casting form and tranfer it in to a box.
- Add 50 mL of fixation solution and incubate the gel 1 to 24 h. The formaldehyde (37 %) must be added to the solution short time before.
- Remove the fixation solution and wash the gel 3 times (1-20 min) with 50 mL of 50 % (v/v) ethanol.
- Add 50 mL of thiosulfate solution and incubate the gel exactly for 1 min. If the gel incubates to long the background becomes to dark.
- Remove thiosulfate solution and wash the gel 3 times (20 s) with dH2O.
- Add 50 mL of impregnation solution and icubate the gel for 15 to 20 min. The formaldehyde (37 %) must be added to the solution short time before.
- Remove impregnation solution and wash the gel 2 times (20 s) with dH2O.
- Add 50 mL of developing solution and incubate the gel until protein bands become visible.
- Remove the developing solution immediately and wash the gel for 10 to 20 s with dH2O.
- Add 50 mL of stop solution and incubate the gel 10 to 20 s.
- Remove the stop solution and wash the gel for 1 min with dH2O.
Tryptic digest of gel lanes for analysis with MALDI-TOF
Note:
- Make sure to work under a fume hood.
- Do not work with protective gloves to prevent contamination of your sample with platicizers.
Reaction tubes have to be cleaned with 60% (v/v) CH3CN, 0,1% (v/v) TFA. Afterwards the solution has to be removed completely followed by evaporation of the tubes under a fume hood. Alternatively microtiter plates from Greiner® (REF 650161) can be used without washing.
- Cut out the protein lanes of a Coomassie-stained SDS-PAGE using a clean scalpel. Gel parts are transferred to the washed reaction tubes/microtiter plate. If necessary cut the parts to smaller slices.
- Gel slices should be washed two times. Therefore add 200 µL 30% (v/v) acetonitrile in 0,1 M ammonium hydrogen carbonate each time and shake lightly for 10 minutes. Remove supernatant and discard to special waste.
- Dry gel slices at least 30 minutes in a Speedvac.
- Rehydrate gel slices in 15 µL Trypsin-solution followed by short centrifugation.
- Gel slices have to be incubated 30 minutes at room temperature, followed by incubation at 37 °C overnight.
- Dry gel slices at least 30 minutes in a Speedvac.
- According to the size of the gel slice, add 5 – 20 µL 50% (v/v) ACN / 0,1% (v/v) TFA.
- Samples can be used for MALDI measurement or stored at -20 °C.
Trypsin-solution: 1 µL Trypsin + 14 µL 10 mM NH4HCO3
- Therefore solubilize lyophilized Trypsin from Promega® in 200 µL of provided buffer and incubate for 15 minutes at 30 °C for activation. For further use it can be stored at -20 °C.
Bisphenol A analysis
Extraction with ethylacetate
- mix 100 µL culture supernatant with 100 µL internal standard ( bisphenol F, 100 µg L-1)
- add 200 µL ethylacetate (HPLC grade) for extraction
- vortex (30 s)
- centrifuge for phase separation (5 min, 5000 g)
- take a bit from upper phase and put it in a clean eppi
- SpeedVac at 40 °C to remove ethlyacetate
- solve remaining BPA in water (HPLC grade), vortex (30 s)
- solubility of BPA in water only 300 mg L-1
- for LC-MS analysis of BPA, 300 mg BPA L-1 is definitely enough
- if you want to detect or expect higher concentrations of BPA, solve it in an acetonitrile-water-mix
HPLC method
- C18 reverse phase column
- Isocratic method: 45 % Acetonitrile
- Flow = 0.6 mL min-1
- UV-detection at 227 nm
- Internal standard: 100 mg L-1 Bisphenol F (BPF)
- Column:
- Eurospher II 100-5 C18p by [http://www.knauer.net/ Knauer]
- Dimensions: 150 x 4.6 mm with precolumn
- Particle size: 5 µm
- Pore size: 100 Å
- Material: silica gel
- Software:
- Clarity (Version 3.0.5.505) by [http://www.dataapex.com/ Data Apex]
- Autosampler:
- Midas by [http://www.spark.nl/ Spark Holland]
- Tray cooling: 10 °C
- Pump:
- L-6200A Intelligent Pump by [http://www.hitachi.com/ Hitachi]
- UV-Detector:
- Series 1050 by [http://www.hp.com/ Hewlett Packard]
NAD+ detection
For the [http://www.molecular-beacons.com/MB_SC_design.html design of molecular beacons] different available tools may help you so that a correct functionality is ensured under defined assay conditions. All measurements were made in the NAD+ bioassay buffer.
Characterisation of molecular beacons
Preferentially, use a complementary oligonucleotide (target) as well as two enshortened oligonucleotides (split target) for [http://www.molecular-beacons.com/MB_SC_protocol.html#cap3 characterising the molecular beacon] in its closed and open state. In this way you can verify simultaneously the ineffectiveness of the split target to reach the molecular beacon`s open state.
- Optimal wavelengths:
- Determine the emission spectrum of your molecular beacon by initially using a literature value for the exctintion wavelength of the labeled fluorophore. For this add 10 µL of 1 µM molecular beacon in 200 µL NAD+ bioassay buffer, mix thoroughly and wait 2 min until measuring the fluorescence with a spectrofluorometer at an defined temperature (optimally 37 °C) .
- Add 10 µL of 2 µM either target or split target and shake at least 6 min until measuring the fluorescence again (when equilibrium is reached).
- Repeat the procedure to determine the extinction spectrum using the self-calculated optimal emission wavelength for your fluorophore/quencher combination. For further fluorescence measurements use the extinction wavelength for which the difference between the molecular beacons's closed state and the open state signals is maximal.
- Thermal profile analysis:
- Prepare a 30 µL reaction mix composed of NAD+ bioassay buffer and 1 µM molecular beacon. In a separate sample add 2 µM either target or split target.
- Use a spectrofluorometric thermal cylcer to dertemine the fluorescence as a function of temperature. Start at 80 °C and decrease the temperature stepwise in 1 °C intervals holding each temperature for 1 min and measuring the fluorescence at the end of each step.
- Signal-to-background ratio (S/B):
- The approach is quite similar to the one that determines the optimal wavelengths which should be used to monitor the fluorescence of the NAD+ bioassay buffer (Fbuffer) and the added molecular beacon (Fclosed) as well as the target (Fopen) or split target. Monitor the fluorescence at the optimal wavelengths until the equilibrium is reached each time before you add a new component.
- The signal-to-background ratio can be calculated as shown on the right. Finally, compare the signal-to-background ratios after adding the target and split target to check whether the split target has an effect on the melting of the molecular beacon's hairpin structure.
- Imaging:
- Prepare a 200 µL reaction mix composed of NAD+ bioassay buffer and 500 nM molecular beacon in a PCR tube. In a separate sample add 600 µM either target or split target. Excitate the samples by a UV transilluminator and take images preferably with a camera that can detect different colours.
Purification of DNA ligase
The purification of the overexpressed NAD+-dependent DNA ligase gene (BBa_K525710) in E. coli was performed under native conditions and Ni-NTA columns were used utilizing the recombinant protein`s C-terminal 6xHis-tag.
- Cultivation
- Prepare an overnight culture of [http://www.promega.com/products/cloning-and-dna-markers/cloning-tools-and-competent-cells/bacterial-strains-and-competent-cells/single-step-_krx_-competent-cells/ E. coli KRX] carrying the plasmid with DNA ligase (BBa_K525710) in 30 ml LB medium containing 20 μg mL-1 chloramphenicol at 37 °C.
- Dilute the overnight culture in 100 ml autoinduction medium (20 μg mL-1 chloramphenicol added) to an OD600 = 0,1 and harvest the cells after 4 h growth at 37 °C.
- Use the centrifuged cells immediately for protein purification or store them at -20 °C.
- Small-scale purification
- Resuspend a pellet derived from 5 ml cell culture volume in 630 μl NPI-10.
- Add 70 μl lysozyme (10 mg/ml) as well as 25 Units benzonase nuclease and mix thoroughly.
- Incubate on ice for 30 min.
- Centrifuge the lysate for 30 min at 12000 g (4 °C). Collect the supernatant (cleared lysate).
- Prepare a [http://www.qiagen.com/products/protein/purification/qiaexpressproteinpurificationsystem/ni-ntaspincolumns.aspx?ShowInfo=1 Ni-NTA spin column] ([http://www.qiagen.com/default.aspx QIAGEN]) by equilibration with 600 μl NPI-10 and centrifugation for 2 min at 890 g (4 °C).
- Load up to 600 μl cleared lysate onto the [http://www.qiagen.com/products/protein/purification/qiaexpressproteinpurificationsystem/ni-ntaspincolumns.aspx?ShowInfo=1 Ni-NTA spin column] and centrifuge for 5 min at 270 g (4 °C). Collect the flow-through.
- Wash the [http://www.qiagen.com/products/protein/purification/qiaexpressproteinpurificationsystem/ni-ntaspincolumns.aspx?ShowInfo=1 Ni-NTA spin column] with 600 μl NPI-20 and centrifuge for 2 min at 890 g (4 °C). Repeat this step and collect flow-through each time.
- Elute the protein with 300 μl NPI-500 and centrifuge for 2 min at 890 g (4 °C). Repeat this step and collect flow-through each time.
- For centrifugal ultrafiltration use [http://www.sartorius-stedim.com/DE/en/Centrifugal-Ultrafiltration--Vivaspin-%26-Centrisart/Vivaspin-500/VS0111-VIVASPIN-500-5%2C000-MCWO-PES-25-BOX/6htcki6uex7/m8qc5rabaox/n8uj9p1cyz4/article.htm Vivaspin 500] concentrators ([http://www.sartorius.de/index.php?id=156&no_cache=1 Sartorius]) with a 5000 molecular weight cut-off PES membrane. Repeat the procedure three times by adding DNA ligase buffer (4 °C) for buffer exchange (remove of imidazole).
- Check each purification step and especially the purity of the protein in the final condition by SDS-PAGE analysis.
- Determine the protein concentration with a Bradford Protein Assay.
- For long-time storage keep the protein at -20 °C.
- If the binding conditions are not effective enough you can reduce the imidazole concentration in NPI-10 to 1-5 mM. If the eluate contains unspecifically bound proteins maybe a washing step with NPI-10 containing 50 mM or 100 mM imidazole could help to get higher purity.
- Large-scale purification
- Resuspend a cell culture pellet in 5 ml binding buffer per gramm biomass.
- Add 10 mM MgCl2, 1 mg/ml lysozyme] as well as 5 Units benzonase nuclease per 1 ml cell culture volume and mix thoroughly.
- Incubate at 4 °C for 30 min.
- Centrifuge the lysate for 30 min at 10000 g (4 °C). Collect the supernatant (cleared lysate).
- Prepare a [http://www.gelifesciences.com/aptrix/upp01077.nsf/Content/Products?OpenDocument&moduleid=165904 HisTrap™ FF crude column] ([http://www.gelifesciences.com/aptrix/upp01077.nsf/Content/life-sciences_homepage GE Healthcare]) by equilibration with 5 column volumes of binding buffer. The following steps have to be performed at 4 °C.
- Load the cleared lysate onto the [http://www.qiagen.com/products/protein/purification/qiaexpressproteinpurificationsystem/ni-ntaspincolumns.aspx?ShowInfo=1 HisTrap™ FF crude column] and collect flow-through.
- Wash the [http://www.qiagen.com/products/protein/purification/qiaexpressproteinpurificationsystem/ni-ntaspincolumns.aspx?ShowInfo=1 HisTrap™ FF crude column] with 10 column volumes of binding buffer and collect flow-through.
- Elute the protein successively with 5 ml of buffers that contain different concentrations of imidazole (e.g. binding buffer with 40, 60, 100, 300 and 500 mM imidazole) and collect flow-through in 1 ml fractions.
- For centrifugal ultrafiltration use [http://www.sartorius-stedim.com/DE/de/Zentrifugal-Ultrafiltration--Vivaspin-%26-Centrisart/Vivaspin-15/VS1501-VIVASPIN-15-10%2C000-MWCO-PES-10-BOX/6htcki6uex7/1b11mt9gskl/kn4oouwu6g3/article.htm Vivaspin 15] concentrators ([http://www.sartorius.de/index.php?id=156&no_cache=1 Sartorius]) with a 10000 molecular weight cut-off PES membrane. Repeat the procedure three times by adding DNA ligase buffer (4 °C) for buffer exchange (remove of imidazole).
- Check each purification step and especially the purity of the protein in the final condition by SDS-PAGE analysis.
- Determine the protein concentration with a Bradford Protein Assay.
- For long-time storage keep the protein at -20 °C.
Deadenylation
The majority of purified DNA ligase from [http://www.promega.com/products/cloning-and-dna-markers/cloning-tools-and-competent-cells/bacterial-strains-and-competent-cells/single-step-_krx_-competent-cells/ E. coli KRX] is usually in the adenylated form. But only the apoenzyme is really useful for the [http://pubs.acs.org/doi/abs/10.1021/ac102742k NAD+ bioassay] so that the AMP moiety has to be removed.
- Mix the purified DNA ligase with deadenylation buffer and incubate for 10 min at 37 °C.
- Use [http://www.sartorius-stedim.com/DE/de/Zentrifugal-Ultrafiltration--Vivaspin-%26-Centrisart/Vivaspin-15/VS1501-VIVASPIN-15-10%2C000-MWCO-PES-10-BOX/6htcki6uex7/1b11mt9gskl/kn4oouwu6g3/article.htm Vivaspin 15] concentrators ([http://www.sartorius.de/index.php?id=156&no_cache=1 Sartorius]) with a 10000 molecular weight cut-off PES membrane for buffer exchange at 4 °C. Repeat the procedure at least three times by adding DNA ligase buffer (remove of NMN and formed NAD+).
- Check the deadenylation process by SDS-PAGE analysis. If the deadenylation was successful there should not be a double band indicating both forms of DNA ligase.
- Determine the protein concentration with a Bradford Protein Assay.
- For long-time storage keep the protein at -20 °C.
NAD+ bioassay
- Prepare a 80 μl aliquot composed of 250 nM molecular beacon and 250 nM split target in NAD+ bioassay buffer.
- Incubate for 8 min at 37 °C.
- Add 6 Units of deadenylated DNA ligase (approximately ... ng).
- Monitor the fluorescence with a spectrofluorometer at 37 °C and wait until it reaches equilibrium.
- Add NAD+ in different final concentrations (e.g. 0, 0.3, 0.5, 5, 10, 30, 40, 80, 200, 300, 500, 1000 nM), stirr shortly and monitor the fluorescence at 37 °C again.
- Determine the initial enhancement rate of flourescence intensity for each NAD+ concentration.
- For the calibration curve plot the initial velocity against the NAD+ concentration.
 "
"