|
|
| (100 intermediate revisions not shown) |
| Line 1: |
Line 1: |
| - | {{css}} | + | {{main}} |
| | | | |
| - | <html>
| |
| - | <head>
| |
| - | <title>UNIPV TEAM 2011</title>
| |
| - | <meta name="description" content="Description" />
| |
| - | <meta name="keywords" content="Keywords" />
| |
| | | | |
| | | | |
| - | <link rel="stylesheet" href="https://2011.igem.org/Team:UNIPV-Pavia/Templates/nivo_default_css?action=raw&ctype=text/css" type="text/css"/> | + | <html> |
| - | <link rel="stylesheet" href="https://2011.igem.org/Team:UNIPV-Pavia/Templates/nivo_css?action=raw&ctype=text/css" type="text/css"/> | + | <h2 class="art-postheader"> Results </h2> |
| | + | <div class="cleared"></div> |
| | + | <div class="art-postcontent"> |
| | + | |
| | + | <!--------------------------------> |
| | + | <!--------------------------------> |
| | + | <!--------------------------------> |
| | + | <!-----------MENU-----------------> |
| | + | <!--------------------------------> |
| | + | <!--------------------------------> |
| | + | <!--------------------------------> |
| | + | <br> |
| | + | <a name='indice'></a> |
| | + | <div class="cleared"></div> |
| | + | <div class="art-postcontent"> |
| | + | <table id="toc" class="toc"> |
| | + | <tr> |
| | + | <td><div id="toctitle"> |
| | + | <h2>Contents</h2> |
| | + | </div> |
| | + | <ul> |
| | + | <li class="toclevel-1"><a href="#assembly"><span class="tocnumber">1</span> <span class="toctext">Part assembly</span></a> |
| | + | <li class="toclevel-1"><a href="#characterization"><span class="tocnumber">1</span> <span class="toctext">Characterization of basic modules</span></a> |
| | + | <ul> |
| | + | <li class="toclevel-2"><a href="#promoters"><span class="tocnumber">2.1</span> <span class="toctext">Promoter characterization</span></a></li> |
| | + | <li class="toclevel-2"><a href="#enzymes"><span class="tocnumber">2.2</span> <span class="toctext">Characterization of the activity of the enzymes AiiA and LuxI</span></a></li> |
| | + | <li class="toclevel-2"><a href="#rbs"><span class="tocnumber">2.3</span> <span class="toctext">Characterization of RBS efficiency</span></a></li> |
| | + | </ul> |
| | + | <li class="toclevel-1"><a href="#growth"><span class="tocnumber">3</span> <span class="toctext">Identification of bacterial growth parameters</span></a></li> |
| | + | <li class="toclevel-1"><a href="#HSL"><span class="tocnumber">4</span> <span class="toctext">Estimation of the spontaneous degradation of HSL in M9 medium and in cultures at different pH values</span></a></li> |
| | + | <li class="toclevel-1"><a href="#t9002"><span class="tocnumber">5</span> <span class="toctext">Characterization of BBa_T9002 biosensor</span></a></li> |
| | + | </ul></td> |
| | + | </tr> |
| | + | </table> |
| | + | </div> |
| | + | <script>if (window.showTocToggle) { var tocShowText = "show"; var tocHideText = "hide"; showTocToggle(); } </script> |
| | + | <em> NB: unless differently specified, all tests were performed in <a href='https://2011.igem.org/Team:UNIPV-Pavia/Protocols#MG1655Z1'><em>E. coli</em> MGZ1</a> in M9 supplemented medium at 37°C. For the cloning of the parts, <a href='https://2011.igem.org/Team:UNIPV-Pavia/Protocols#TOP10'><em>E. coli</em> TOP10</a> was used. </em> <br> |
| | + | <br> |
| | + | |
| | + | <!--------------------------------> |
| | + | <!--------------------------------> |
| | + | <!--------------------------------> |
| | + | <!-----------ASSEMBLY-------------> |
| | + | <!--------------------------------> |
| | + | <!--------------------------------> |
| | + | <!--------------------------------> |
| | + | |
| | + | <a name='assembly'></a> |
| | + | <h1>Parts assembly</h1> |
| | + | All the parts have been cloned with success. The part name, plasmids and quality controls are reported in the <a href='https://2011.igem.org/Team:UNIPV-Pavia/Freezer'>Freezer section</a>. |
| | + | <div align="right"><small><a href="#indice" title="">^top</a></small></div> |
| | + | |
| | + | <!--------------------------------> |
| | + | <!--------------------------------> |
| | + | <!--------------------------------> |
| | + | <!-------CHARACTERIZATION---------> |
| | + | <!--------------------------------> |
| | + | <!--------------------------------> |
| | + | <!--------------------------------> |
| | + | |
| | + | <a name='characterization'></a> |
| | + | <h1>Characterization of basic modules</h1> |
| | + | |
| | + | <!--------------------------------> |
| | + | <!--------------------------------> |
| | + | <!--------------------------------> |
| | + | <!--------pTet and pLux-----------> |
| | + | <!--------------------------------> |
| | + | <!--------------------------------> |
| | + | <!--------------------------------> |
| | + | |
| | + | <a name='promoters'></a> |
| | + | <h2>Characterization of promoters pTet and pLux</h2> |
| | + | <p align='justify'> |
| | + | <p>Inducible and constitutive promoters were assembled upstream of different coding sequences containing an RBS from the Community collection.</p> |
| | + | <p>The assembled RBSs are:</p> |
| | + | <br> |
| | + | <div align='center'> |
| | + | <table class='data'> |
| | + | <tr> |
| | + | <td class="row"><b>BioBrick code</b></td> |
| | + | <td><b> Declared efficiency</b></td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td class="row">BBa_B0030 </td> |
| | + | <td class="row"> 0,6</td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td class="row">BBa_B0031 </td> |
| | + | <td class="row"> 0,07</td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td class="row">BBa_B0032 </td> |
| | + | <td class="row"> 0,3</td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td class="row">BBa_B0034 </td> |
| | + | <td class="row"> 1</td> |
| | + | </tr> |
| | + | </table> |
| | + | </div> |
| | + | <br> |
| | + | <div align="justify"> |
| | + | <p>For an inducible device, the RBS variation has the purpose to stretch the induction curve, thus modulating its PoPs-OUT range.</p> |
| | + | <p>The complex RBS-promoter acts as a whole regulatory element and determines the amount of translated protein. |
| | + | RBSs have been reported to have an un-modular behavior, since the translational efficiency is not independent on the coding sequences, but variates as an effect of different mRNA structure stability [Salis et al., Nat Biotec, 2009]. It is not possible to separate the effects of the sole promoter and of the sole RBS on the total amount/activity of gene product (in this case study, mRFP).</p> |
| | + | <p>For this reason, every combination 'Promoter+RBS' was studied as a different regulatory element. Regulatory elements were characterized using mRFP reporter protein for different RBSs in terms of Synthesis rate per Cell (<b>S<sub>cell</sub></b>) and <b>R.P.U.s</b> (Relative Promoter Units) as explained in <a href='https://2011.igem.org/Team:UNIPV-Pavia/Measurements'>measurements</a> section.</p> |
| | + | </p> |
| | + | <p>Operative parameters of the promoter are derived from the estimated Hill equations obtained by <em>nonlinear least squares</em> fitting (<em>lsqnonlin</em> Matlab routine) of the <a href='https://2011.igem.org/Team:UNIPV-Pavia/Project/Modelling#Equations_for_gene_networks'>Hill function</a> expressed in RPUs:</p> |
| | + | <p></p> |
| | + | <p> |
| | + | <ol> |
| | + | <ul> |
| | + | <li><b> RPU<sub>max</sub></b> is equal to the α and represents the maximum promoter activity |
| | + | </p> |
| | + | </li> |
| | + | <p> |
| | + | <li><b> RPU<sub>min</sub></b> is equal to the α * δ represents the minimum promoter activity |
| | + | </p> |
| | + | </li> |
| | + | <p> |
| | + | <li> <b>Switch point</b> is computed as the abscissa of the inflection point of the Hill curve and it is representative of the position of linear region |
| | + | </p> |
| | + | </li> |
| | + | <p> |
| | + | <li> <b>Linearity boundaries</b> are determined as the intersection between the tangent line to the inflection point and the upper and lower horizontal boundaries of the Hill curve. |
| | + | |
| | + | </li> |
| | + | </p> |
| | + | </ul> |
| | + | </ol> |
| | + | </p> |
| | + | <p align='justify'> The estimated parameters for the Hill functions of pLux are summarized in the table below. For more details on parameter estimation, see the <a href='https://2011.igem.org/Team:UNIPV-Pavia/Project/Modelling#Ptet_&_Plux'>model section</a>. </p> |
| | + | <table class='data' width='100%'> |
| | + | <tr> |
| | + | <td class="row"><b>RBS</b></td> |
| | + | <td class="row"><b>α<sub>p<sub>Lux</sub></sub> [(AUr/min)/cell]</b></td> |
| | + | <td class="row"><b>δ<sub>p<sub>Lux</sub></sub> [-]</b></td> |
| | + | <td class="row"><b>η<sub>p<sub>Lux</sub></sub> [-]</b></td> |
| | + | <td class="row"><b>k<sub>p<sub>Lux</sub></sub> [ng/ml]</b></td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td class="row">BBa_B0030</td> |
| | + | <td class="row">438 [10]</td> |
| | + | <td class="row">0.05 [>100]</td> |
| | + | <td class="row">2 [47]</td> |
| | + | <td class="row">1.88 [27]</td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td class="row">BBa_B0031</td> |
| | + | <td class="row">9.8 [7]</td> |
| | + | <td class="row">0.11 [57]</td> |
| | + | <td class="row">1.2 [29]</td> |
| | + | <td class="row">1.5 [26]</td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td class="row">BBa_B0032</td> |
| | + | <td class="row">206 [3]</td> |
| | + | <td class="row">0 [>>100]</td> |
| | + | <td class="row">1.36 [10]</td> |
| | + | <td class="row">1.87 [9]</td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td class="row">BBa_B0034</td> |
| | + | <td class="row">1105 [6]</td> |
| | + | <td class="row">0.02 [>100]</td> |
| | + | <td class="row">1.33 [19]</td> |
| | + | <td class="row">2.34 [18]</td> |
| | + | </tr> |
| | + | </table> |
| | + | <div align="center">Data are provided as average [CV%].</div> |
| | + | <br> |
| | + | <p>The operative parameters are summarized in the table below:</p> |
| | + | </div> |
| | + | <table align='center' class='data' width='100%'> |
| | + | <tr> |
| | + | <td class='row'><b>RBS</b></td> |
| | + | <td class='row'><b>RPU<sub>max</sub></b></td> |
| | + | <td class='row'><b>RPU<sub>min</sub></b></td> |
| | + | <td class='row'><b>Switch point [nM]</b></td> |
| | + | <td class='row'><b>Linear boundaries [MIN; MAX] [nM]</b></td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td class='row'>B0030</td> |
| | + | <td class='row'>4.28</td> |
| | + | <td class='row'>0.20</td> |
| | + | <td class='row'>1.08</td> |
| | + | <td class='row'>[0.36; 3.27]</td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td class='row'>B0031</td> |
| | + | <td class='row'>4.93</td> |
| | + | <td class='row'>0.55</td> |
| | + | <td class='row'>0.25</td> |
| | + | <td class='row'>[0.03; 2.30]</td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td class='row'>B0032</td> |
| | + | <td class='row'>9.49</td> |
| | + | <td class='row'>0.02</td> |
| | + | <td class='row'>0.47</td> |
| | + | <td class='row'>[0.07; 3.07]</td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td class='row'>B0034</td> |
| | + | <td class='row'>21.53</td> |
| | + | <td class='row'>0.51</td> |
| | + | <td class='row'>0.53</td> |
| | + | <td class='row'>[0.08; 3.77]</td> |
| | + | </tr> |
| | + | </table> |
| | + | <table width='100%'> |
| | + | <tr> |
| | + | <td width='50%'><div style="text-align:justify"> |
| | + | <div class="thumbinner" width='100%'><a href="https://static.igem.org/mediawiki/2011/5/50/E17_RPU_80.jpg" class="image"><img alt="" src="https://static.igem.org/mediawiki/2011/5/50/E17_RPU_80.jpg" class="thumbimage" width="100%"></a></div> |
| | + | </div></td> |
| | + | <td width='50%'><div style="text-align:justify"> |
| | + | <div class="thumbinner" width='100%'><a href="https://static.igem.org/mediawiki/2011/c/c4/E18_RPU_80.jpg" class="image"><img alt="" src="https://static.igem.org/mediawiki/2011/c/c4/E18_RPU_80.jpg" class="thumbimage" width="100%"></a></div> |
| | + | </div></td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td width='50%'><div style="text-align:justify"> |
| | + | <div class="thumbinner" width='100%'><a href="https://static.igem.org/mediawiki/2011/7/78/E19_RPU_80.jpg" class="image"><img alt="" src="https://static.igem.org/mediawiki/2011/7/78/E19_RPU_80.jpg" class="thumbimage" width="100%"></a></div> |
| | + | </div></td> |
| | + | <td width='50%'><div style="text-align:justify"> |
| | + | <div class="thumbinner" width='100%'><a href="https://static.igem.org/mediawiki/2011/5/55/E20_RPU_80.jpg" class="image"><img alt="" src="https://static.igem.org/mediawiki/2011/5/55/E20_RPU_80.jpg" class="thumbimage" width="100%"></a></div> |
| | + | </div></td> |
| | + | </tr> |
| | + | </table> |
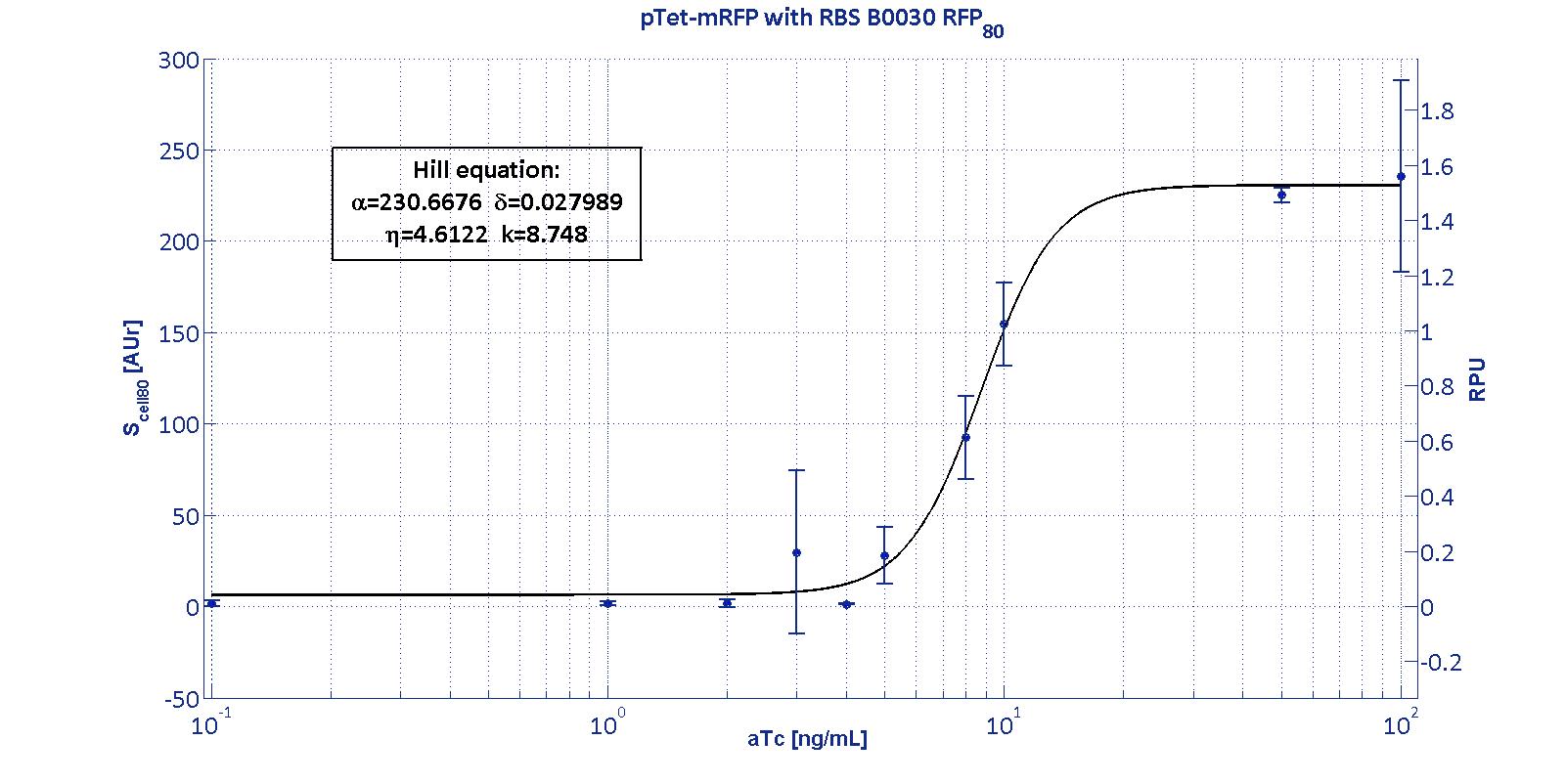
| | + | <p>The protocols for the characterization of p<sub>Tet</sub> promoter are reported in the <a href='https://2011.igem.org/Team:UNIPV-Pavia/Measurements#pTet_protocol'>p<sub>Tet</sub> measurement section</a>.</p> |
| | + | <p> Here we have characterized its transcriptional strength as a function of aTc induction (ng/ul) for different RBSs. Three different induction curves were obtained and are reported in figure:</p> |
| | | | |
| | + | <center> |
| | + | <a href="https://static.igem.org/mediawiki/2011/a/af/E32_RPU_80.jpg" class="image"> <img alt="" src="https://static.igem.org/mediawiki/2011/a/af/E32_RPU_80.jpg" class="thumbimage" width="50%"></a> |
| | + | </center> |
| | + | <table width='100%'> |
| | + | <tr> |
| | + | <td width='50%'><div style="text-align:justify"> |
| | + | <div class="thumbinner" width='100%'><a href="https://static.igem.org/mediawiki/2011/9/99/E34_RPU_80.jpg" class="image"><img alt="" src="https://static.igem.org/mediawiki/2011/9/99/E34_RPU_80.jpg" class="thumbimage" width="100%"></a></div> |
| | + | </div></td> |
| | + | <td width='50%'><div style="text-align:justify"> |
| | + | <div class="thumbinner" width='100%'><a href="https://static.igem.org/mediawiki/2011/e/e4/E35_RPU_80.jpg" class="image"><img alt="" src="https://static.igem.org/mediawiki/2011/e/e4/E35_RPU_80.jpg" class="thumbimage" width="100%"></a></div> |
| | + | </div></td> |
| | + | </tr> |
| | + | </table> |
| | + | <p align='justify'> The estimated parameters of the Hill curves described in the figures are summarized in the table below: </p> |
| | + | <br> |
| | + | <table class='data' width='100%' title='parameter value'> |
| | + | <tr> |
| | + | <td class="row"><b>RBS</b></td> |
| | + | <td class="row"><b>α<sub>p<sub>Tet</sub></sub> [(AUr/min)/cell]</b></td> |
| | + | <td class="row"><b>δ<sub>p<sub>Tet</sub></sub> [-]</b></td> |
| | + | <td class="row"><b>η<sub>p<sub>Tet</sub></sub> [-]</b></td> |
| | + | <td class="row"><b>k<sub>p<sub>Tet</sub></sub> [nM]</b></td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td class="row">BBa_B0030</td> |
| | + | <td class="row">230.67 [3.7]</td> |
| | + | <td class="row">0.028 [91.61]</td> |
| | + | <td class="row">4.61 [23.73]</td> |
| | + | <td class="row">8.75 [4.16]</td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td class="row">BBa_B0031</td> |
| | + | <td class="row">ND</td> |
| | + | <td class="row">ND</td> |
| | + | <td class="row">ND</td> |
| | + | <td class="row">ND</td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td class="row">BBa_B0032</td> |
| | + | <td class="row">55.77 [12]</td> |
| | + | <td class="row">1.53E-11 [>>100]</td> |
| | + | <td class="row">4.98 [57.62]</td> |
| | + | <td class="row">7.26 [14.98]</td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td class="row">BBa_B0034</td> |
| | + | <td class="row">120 [5.95]</td> |
| | + | <td class="row">0.085 [40.6]</td> |
| | + | <td class="row">24.85 [47.6]</td> |
| | + | <td class="row">9 [5.43]</td> |
| | + | </tr> |
| | + | </table> |
| | + | Data are provided as average [CV%] <br> |
| | + | <br> |
| | + | <p align='justify'> The operative parameters are summarized in the table below: </p> |
| | + | <table align='center' class='data' width='100%'> |
| | + | <tr> |
| | + | <td class='row'><b>RBS</b></td> |
| | + | <td class='row'><b>RPU<sub>max</sub></b></td> |
| | + | <td class='row'><b>RPU<sub>min</sub></b></td> |
| | + | <td class='row'><b>Switch point [ng/ml]</b></td> |
| | + | <td class='row'><b>Linear boundaries [MIN; MAX] [ng/ml]</b></td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td class='row'>B0030</td> |
| | + | <td class='row'>1.53</td> |
| | + | <td class='row'>~0</td> |
| | + | <td class='row'>7.95</td> |
| | + | <td class='row'>[4.66;11.99]</td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td class='row'>B0031</td> |
| | + | <td class='row'>ND</td> |
| | + | <td class='row'>ND</td> |
| | + | <td class='row'>ND</td> |
| | + | <td class='row'>ND</td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td class='row'>B0032</td> |
| | + | <td class='row'>3.16</td> |
| | + | <td class='row'>~0</td> |
| | + | <td class='row'>6.7</td> |
| | + | <td class='row'>[4.45;10.05]</td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td class='row'>B0034</td> |
| | + | <td class='row'>2.73</td> |
| | + | <td class='row'>0.23</td> |
| | + | <td class='row'>8.96</td> |
| | + | <td class='row'>[8.27;9.71]</td> |
| | + | </tr> |
| | + | </table> |
| | + | <div align="right"><small><a href="#indice" title="">^top</a></small></div> |
| | | | |
| - | <script type="text/javascript" src="http://ajax.googleapis.com/ajax/libs/jquery/1.3.2/jquery.min.js"></script> | + | <!--------------------------------> |
| - | <script type="text/javascript" src="https://2011.igem.org/Team:UNIPV-Pavia/Templates/nivo_slider_pack?action=raw& | + | <!--------------------------------> |
| | + | <!--------------------------------> |
| | + | <!--------AiiA and LuxI-----------> |
| | + | <!--------------------------------> |
| | + | <!--------------------------------> |
| | + | <!--------------------------------> |
| | | | |
| - | ctype=text/javascript"></script>
| + | <a name='enzymes'></a> |
| - | | + | <h2>Characterization of enzymes AiiA and LuxI</h2> |
| - | <script type="text/javascript">
| + | <p align='justify'> LuxI has been characterized through the Biosensor BBa_T9002 (see <a href='https://2011.igem.org/Team:UNIPV-Pavia/Project/Modelling#introduction_to_T9002'>modelling section</a>) using the <a href='https://2011.igem.org/Team:UNIPV-Pavia/Parts/Characterized#pTetLuxI'>p<sub>Tet</sub>-RBSx-LuxI-TT</a> measurement systems. <br> |
| - | $(window).load(function() {
| + | The parameters V<sub>max</sub>, k<sub>M,LuxI</sub> and α<sub>RBSx</sub> were estimated with a simultaneous fitting of the data collected as described in <a href='https://2011.igem.org/Team:UNIPV-Pavia/Measurements#LuxI'>measurement section</a> for the four measurement parts <a href='https://2011.igem.org/Team:UNIPV-Pavia/Parts/Characterized#pTetLuxI'>p<sub>Tet</sub>-RBSx-LuxI-TT</a> assayed by <a href='https://2011.igem.org/Team:UNIPV-Pavia/Measurements#T9002'>BBa_T9002 biosensor</a> section. |
| - | $('#slider').nivoSlider({
| + | <div align="justify"> |
| - | effect:'fade',
| + | <div class="thumbinner" style="width: 850px;"> <a href="https://static.igem.org/mediawiki/2011/4/40/Luxi_prod.jpg"> <img alt="" src="https://static.igem.org/mediawiki/2011/4/40/Luxi_prod.jpg" class="thumbimage" width="85%" height="50%"></a></div> |
| - | pauseOnHover:true
| + | |
| - | });
| + | |
| - | });
| + | |
| - | | + | |
| - | | + | |
| - | </script> | + | |
| - | | + | |
| - | | + | |
| - | </head> | + | |
| - | | + | |
| - | <body> | + | |
| - | <div class="igem_logo" onclick="location.href='https://2011.igem.org';" style="cursor: pointer;"></div> | + | |
| - | <div id="art-main">
| + | |
| - | <div class="art-sheet">
| + | |
| - | <div class="art-sheet-tl"></div>
| + | |
| - | <div class="art-sheet-tr"></div>
| + | |
| - | <div class="art-sheet-bl"></div>
| + | |
| - | <div class="art-sheet-br"></div>
| + | |
| - | <div class="art-sheet-tc"></div>
| + | |
| - | <div class="art-sheet-bc"></div>
| + | |
| - | <div class="art-sheet-cl"></div>
| + | |
| - | <div class="art-sheet-cr"></div>
| + | |
| - | <div class="art-sheet-cc"></div>
| + | |
| - | <div class="art-sheet-body">
| + | |
| - | <div class="art-header">
| + | |
| - | <div class="art-header-clip">
| + | |
| - | <div class="art-header-jpeg" onclick="location.href='https://2011.igem.org/Team:UNIPV-Pavia';" style="cursor: pointer;"></div>
| + | |
| - | </div>
| + | |
| - | <div class="art-logo"></div>
| + | |
| - | </div>
| + | |
| - | | + | |
| - | | + | |
| - | | + | |
| - | <div class="cleared reset-box"></div>
| + | |
| - | <div class="art-nav">
| + | |
| - | <div class="art-nav-l"></div>
| + | |
| - | <div class="art-nav-r"></div>
| + | |
| - | <div class="art-nav-outer">
| + | |
| - | <ul class="art-hmenu">
| + | |
| - | <li>
| + | |
| - | <a><span class="l"></span><span class="r"></span><span class="t">Project</span></a>
| + | |
| - | <ul>
| + | |
| - | <li>
| + | |
| - | <a href="./project/motivation.html">Motivation</a>
| + | |
| - | | + | |
| - | </li>
| + | |
| - | <li>
| + | |
| - | <a href="./project/solution.html">Solution</a>
| + | |
| - | | + | |
| - | </li>
| + | |
| - | <li>
| + | |
| - | <a href="./project/modelling.html">Modelling</a>
| + | |
| - | | + | |
| - | </li>
| + | |
| - | <li>
| + | |
| - | <a href="./project/results.html">Results</a>
| + | |
| - | | + | |
| - | </li>
| + | |
| - | <li>
| + | |
| - | <a href="./project/references.html">References</a>
| + | |
| - | | + | |
| - | </li>
| + | |
| - | </ul>
| + | |
| - | </li>
| + | |
| - | <li>
| + | |
| - | <a><span class="l"></span><span class="r"></span><span class="t">Lab</span></a>
| + | |
| - |
| + | |
| - | <ul>
| + | |
| - | <li>
| + | |
| - | <a>Materials & Methods</a>
| + | |
| - | <ul>
| + | |
| - | <li>
| + | |
| - | <a href="https://2011.igem.org/Team:UNIPV-Pavia/Protocols">Protocols</a>
| + | |
| - | | + | |
| - | </li>
| + | |
| - | <li>
| + | |
| - | <a href="./lab/materials-methods/instruments.html">Instruments</a>
| + | |
| - | | + | |
| - | </li>
| + | |
| - | </ul>
| + | |
| - | | + | |
| - | </li>
| + | |
| - | <li>
| + | |
| - | <a>Notebook</a>
| + | |
| - | <ul>
| + | |
| - | <li>
| + | |
| - | <a href="https://2011.igem.org/Team:UNIPV-Pavia/Calendar">Calendar</a>
| + | |
| - | | + | |
| - | </li>
| + | |
| - | <li>
| + | |
| - | <a href="https://2011.igem.org/Team:UNIPV-Pavia/Freezer">Freezer Management</a>
| + | |
| - | | + | |
| - | </li>
| + | |
| - | </ul>
| + | |
| - | | + | |
| - | </li>
| + | |
| - | <li>
| + | |
| - | <a href="https://2011.igem.org/Team:UNIPV-Pavia/Biosafety">Biosafety</a>
| + | |
| - | | + | |
| - | </li>
| + | |
| - | </ul>
| + | |
| - | </li>
| + | |
| - | <li>
| + | |
| - | <a><span class="l"></span><span class="r"></span><span class="t">Parts</span></a>
| + | |
| - | <ul>
| + | |
| - | <li>
| + | |
| - | <a href="./parts/submitted.html">Submitted</a>
| + | |
| - | | + | |
| - | </li>
| + | |
| - | <li>
| + | |
| - | <a href="./parts/characterized.html">Characterized</a>
| + | |
| - | | + | |
| - | </li>
| + | |
| - | </ul>
| + | |
| - | </li>
| + | |
| - | <li>
| + | |
| - | <a><span class="l"></span><span class="r"></span><span class="t">About us</span></a>
| + | |
| - | <ul>
| + | |
| - | <li>
| + | |
| - | <a href="./about-us/team.html">Team</a>
| + | |
| - | | + | |
| - | </li>
| + | |
| - | <li>
| + | |
| - | <a href="./about-us/pavia.html">Pavia</a>
| + | |
| - | | + | |
| - | </li>
| + | |
| - | <li>
| + | |
| - | <a href="./about-us/gallery.html">Gallery</a>
| + | |
| - | | + | |
| - | </li>
| + | |
| - | <li>
| + | |
| - | <a href="./about-us/sponsors.html">Sponsors</a>
| + | |
| - | | + | |
| - | </li>
| + | |
| - | </ul>
| + | |
| - | </li>
| + | |
| - | </ul>
| + | |
| | </div> | | </div> |
| | + | <div align="justify"> |
| | + | <p>The estimated parameters for the enzymatic activity of LuxI are reported in the table below:</p> |
| | </div> | | </div> |
| - | | + | <div align=center"> |
| - | | + | <table align="center" class='data'> |
| - | </html> | + | <tr> |
| - | {{article}}
| + | <td class='row'><b>V<sub>max</sub></b></td> |
| - | | + | <td class='row'><b>k<sub>M,LuxI</sub></b></td> |
| - | | + | <td class='row'><b>α<sub>B0030</sub></b></td> |
| - | | + | <td class='row'><b>α<sub>B0031</sub></b></td> |
| - | | + | <td class='row'><b>α<sub>B0032</sub></b></td> |
| - | | + | <td class='row'><b>α<sub>B0034</sub></b></td> |
| - | <html> | + | </tr> |
| - | | + | <tr> |
| - | | + | <td class='row'>3.56*10<sup>-9</sup></td> |
| - | | + | <td class='row'>6.87*10<sup>3</sup></td> |
| - | <div style="width:700px;margin:40px auto;">
| + | <td class='row'> 87 </td> |
| - | <div class="slider-wrapper theme-default"> | + | <td class='row'>8.5</td> |
| - | <div class="ribbon"></div> | + | <td class='row'>ND</td> |
| - | | + | <td class='row'> 252 </td> |
| - | <div id="nivoslider-125" class="nivoSlider" style="width:700px;height:300px;"><img src="http://nivo.dev7studios.com/wp-content/plugins/nivo-slider-pro/timthumb.php?src=http://nivo.dev7studios.com/wp-content/uploads/2011/08/nemo83.png&h=300&w=700&zc=1&q=100" alt="" /><img src="http://nivo.dev7studios.com/wp-content/plugins/nivo-slider-pro/timthumb.php?src=http://nivo.dev7studios.com/wp-content/uploads/2011/08/slider65.png&h=300&w=700&zc=1&q=100" title="#nivoslider-125-caption-0" alt="" /><img src="http://nivo.dev7studios.com/wp-content/plugins/nivo-slider-pro/timthumb.php?src=http://nivo.dev7studios.com/wp-content/uploads/2011/08/walle12.png&h=300&w=700&zc=1&q=100" alt="" /></div> | + | </tr> |
| | + | </table> |
| | </div> | | </div> |
| - | <div id="nivoslider-125-caption-0" class="nivo-html-caption">You can add captions too…</div>
| |
| - | <p><script type="text/javascript">
| |
| - | jQuery(window).load(function(){
| |
| - | jQuery("#nivoslider-125").nivoSlider({
| |
| - | effect:"random",
| |
| - | slices:15,
| |
| - | boxCols:8,
| |
| - | boxRows:4,
| |
| - | animSpeed:500,
| |
| - | pauseTime:3000,
| |
| - | startSlide:0,
| |
| - | directionNav:true,
| |
| - | directionNavHide:true,
| |
| - | controlNav:true,
| |
| - | controlNavThumbs:false,
| |
| - | controlNavThumbsFromRel:true,
| |
| - | keyboardNav:true,
| |
| - | pauseOnHover:true,
| |
| - | manualAdvance:false
| |
| - | });
| |
| - | });
| |
| - | </script>
| |
| - | </div>
| |
| - |
| |
| - |
| |
| - |
| |
| - |
| |
| - |
| |
| - |
| |
| - |
| |
| | <br> | | <br> |
| - |
| + | </p> |
| - | <h2 class="art-postheader">
| + | </p> |
| - | CTRL + <em>E</em>
| + | <p align='justify'> The provided parameters k<sub>M</sub> and V<sub>max</sub> represent the enzymatic activity of LuxI, described by our model. They must not be confused with the operative parameters of the Michaelis-Menten relation. |
| - | </h2>
| + | These synthetic parameters have a great importance, since they can be used in more complicated models in order to predict the behavior of complex circuits. </p> |
| - | <div class="cleared"></div>
| + | <p align='justify'> The AiiA enzyme activity has been characterized under the regulation of p<sub>tet</sub> promoter, assaying its enzymatic activity. </p> |
| - | <div class="art-postcontent">
| + | <p align='justify'> The parameters k<sub>cat</sub>, k<sub>M,AiiA</sub> and α<sub>RBSx</sub> would have been estimated with a simultaneous fitting of the data collected as described in <a href='https://2011.igem.org/Team:UNIPV-Pavia/Measurements#AiiA'>measurement section</a> for the four measurement parts <a href='https://2011.igem.org/Team:UNIPV-Pavia/Parts/Characterized#pTetAiiA'>p<sub>Tet</sub>-RBSx-AiiA-TT</a> assayed by <a href='https://2011.igem.org/Team:UNIPV-Pavia/Measurements#T9002'>BBa_T9002 biosensor</a> section. |
| - | | + | Unfortunately, their estimation revealed difficult.<br> |
| - | <p style="text-align:center;"><span style="font-style:italic;">Signalling is nothing without control...</span></p> | + | In the first experiments with the measurement system <a href="https://static.igem.org/mediawiki/2011/1/11/Pc_aiia.jpg">p<sub>Tet</sub>-RBSx-AiiA-TT</a> in LOW-COPY at pH=7 no degradation of HSL was observed. The collected data are shown in the figure below. HSL degradation is identical in the measurement system and in the negative control after 21 hours. |
| | + | <table width='100%'> |
| | + | <tr> |
| | + | <td width='50%'><div style="text-align:justify"> |
| | + | <div class="thumbinner" width='100%'> <a href="https://static.igem.org/mediawiki/2011/6/69/Aiia_LC_B0030.jpg"> <img alt="" src="https://static.igem.org/mediawiki/2011/6/69/Aiia_LC_B0030.jpg" class="thumbimage" width="100%"></a></div> |
| | + | </div></td> |
| | + | <td width='50%'><div style="text-align:justify"> |
| | + | <div class="thumbinner" width='100%'> <a href="https://static.igem.org/mediawiki/2011/6/63/Aiia_LC_B0031.jpg"> <img alt="" src="https://static.igem.org/mediawiki/2011/6/63/Aiia_LC_B0031.jpg" class="thumbimage" width="100%"></a></div> |
| | + | </div></td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td width='50%'><div style="text-align:justify"> |
| | + | <div class="thumbinner" width='100%'> <a href="https://static.igem.org/mediawiki/2011/f/f5/Aiia_LC_B0032.jpg"> <img alt="" src="https://static.igem.org/mediawiki/2011/f/f5/Aiia_LC_B0032.jpg" class="thumbimage" width="100%"></a></div> |
| | + | </div></td> |
| | + | <td width='50%'><div style="text-align:justify"> |
| | + | <div class="thumbinner" width='100%'> <a href="https://static.igem.org/mediawiki/2011/8/8d/Aiia_LC_B0034.jpg"> <img alt="" src="https://static.igem.org/mediawiki/2011/8/8d/Aiia_LC_B0034.jpg" class="thumbimage" width="100%"></a></div> |
| | + | </div></td> |
| | + | </tr> |
| | + | </table> |
| | | | |
| | | | |
| | + | Experiments on these parts gave us the opportunity to characterize only the activity of the enzyme in <em>E. COLI</em> TOP10 in high copy number plasmid, providing only some information about the order of magnitude of the model parameters, which has been designed to work in <em>E. COLI</em> MGZ1 in low copy number plasmid. |
| | + | <div align="right"><small><a href="#indice" title="">^top</a></small></div> |
| | + | |
| | + | <!--------------------------------> |
| | + | <!--------------------------------> |
| | + | <!--------------------------------> |
| | + | <!--------------RBS---------------> |
| | + | <!--------------------------------> |
| | + | <!--------------------------------> |
| | + | <!--------------------------------> |
| | + | |
| | + | <a name='rbs'></a> |
| | + | <h2>Characterization of the efficiency of RBSs from the community collection</h2> |
| | + | RBSs were used for the fine tuning of CTRL+E. Different experimental conditions were assayed. <br> |
| | + | <br> |
| | + | Estimated efficiencies in pSB4C5 plasmid with -RBSx-mRFP-TT coding sequence under the control of the specified promoter: <br> |
| | + | <br> |
| | + | <div align="center"> |
| | + | <table class='data' width='70%'> |
| | + | <tr> |
| | + | <td class='row'><b>RBS</b></td> |
| | + | <td class='row'><b>eff<sub>p<sub>Lux</sub></sub></b></td> |
| | + | <td class='row'><b>eff<sub>p<sub>Tet</sub></sub></b></td> |
| | + | <td class='row'><b>eff<sub>J23101</sub></b></td> |
| | + | <td class='row'><b>Declared efficiency</b></td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td class='row'>B0030</td> |
| | + | <td class='row'>0.40</td> |
| | + | <td class='row'>1.6814</td> |
| | + | <td class='row'>2.45</td> |
| | + | <td class='row'>0,6</td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td class='row'>B0031</td> |
| | + | <td class='row'>0.01</td> |
| | + | <td class='row'>ND</td> |
| | + | <td class='row'>0.04</td> |
| | + | <td class='row'>0,07</td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td class='row'>B0032</td> |
| | + | <td class='row'>0.19</td> |
| | + | <td class='row'>0.4193</td> |
| | + | <td class='row'>0.40</td> |
| | + | <td class='row'>0,3</td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td class='row'>B0034</td> |
| | + | <td class='row'>1</td> |
| | + | <td class='row'>1</td> |
| | + | <td class='row'>1</td> |
| | + | <td class='row'>1</td> |
| | + | </tr> |
| | + | </table> |
| | + | </div> |
| | + | <br> |
| | + | <br> |
| | + | Estimated efficiencies in pSB4C5 plasmid with pTet-RBSx-GeneX-TT, with GeneX=mRFP, AiiA or LuxI: <br> |
| | + | <br> |
| | + | <div align="center"> |
| | + | <table class='data' width='70%'> |
| | + | <tr> |
| | + | <td class='row'><b>RBS</b></td> |
| | + | <td class='row'><b>eff<sub>mRFP</sub></b></td> |
| | + | <td class='row'><b>eff<sub>AiiA</sub></b></td> |
| | + | <td class='row'><b>eff<sub>LuxI</sub></b></td> |
| | + | <td class='row'><b>Declared efficiency</b></td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td class='row'>B0030</td> |
| | + | <td class='row'>1.72</td> |
| | + | <td class='row'>0.53</td> |
| | + | <td class='row'>0.45</td> |
| | + | <td class='row'>0,6</td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td class='row'>B0031</td> |
| | + | <td class='row'>0.03</td> |
| | + | <td class='row'>0.83</td> |
| | + | <td class='row'>0.028</td> |
| | + | <td class='row'>0,07</td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td class='row'>B0032</td> |
| | + | <td class='row'>0.37</td> |
| | + | <td class='row'>0.50</td> |
| | + | <td class='row'>N.D.</td> |
| | + | <td class='row'>0.3</td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td class='row'>B0034</td> |
| | + | <td class='row'>1</td> |
| | + | <td class='row'>1</td> |
| | + | <td class='row'>1</td> |
| | + | <td class='row'>1</td> |
| | + | </tr> |
| | + | </table> |
| | + | </div> |
| | + | <div align="right"><small><a href="#indice" title="">^top</a></small></div> |
| | + | |
| | + | <!--------------------------------> |
| | + | <!--------------------------------> |
| | + | <!--------------------------------> |
| | + | <!-----------growth---------------> |
| | + | <!--------------------------------> |
| | + | <!--------------------------------> |
| | + | <!--------------------------------> |
| | + | |
| | + | <a name='growth'></a> |
| | + | <h2>Identification of bacterial growth parameters</h2> |
| | | | |
| | + | |
| | + | <p align='justify'> The bacterial growth curve has been modelled as a logistic curve and is represented by the following equation: <br> |
| | + | <br> |
| | + | dN/dt=N*μ*(N<sub>max</sub>-N)/N<sub>max</sub><br> |
| | + | N(0)=n<sub>0</sub> <br> |
| | + | <br> |
| | + | where μ represents the growth rate of the cells (<em>E. coli</em> MGZ1 in M9 supplemented medium) and N<sub>max</sub> represents the maximum number of cells in the well. For a detailed description of the parameters, see <a href='https://2011.igem.org/Team:UNIPV-Pavia/Project/Modelling#Table_of_parameters'>modelling section</a>. For details on parameters identification, see <a href='https://2011.igem.org/Team:UNIPV-Pavia/Project/Modelling#N'>identification section.</a> The growth curves in all the performed experiments are measured in O.D.<sub>600</sub>. Since the <em>N</em> species in the model is expressed in <em>cell number</em>, a conversion factor has been estimated. The conversion factor <b>K<sub>O.D.toC.F.U.</sub></b> has been estimated as follows. |
| | + | <ol> |
| | + | <ul> |
| | + | <li>Two cultures C1 and C2 (MGZ1 cells) were grown in 1ml M9 medium till saturation (ON liquid culture, 37°C, 220 rpm).</li> |
| | + | <li>Next morning, both C1 and C2 were diluted in M9 medium with a final volume of 1ml with the following dilution factors: |
| | + | <ul> |
| | + | <li>1:1</li> |
| | + | <li>1:10</li> |
| | + | <li>1:100</li> |
| | + | <li>1:1000</li> |
| | + | </ul> |
| | + | in fresh M9 medium. These cultures were grown for further 1 hour at 37°C, 220 rpm. </li> |
| | + | <li>After 1 hour, O.D.<sub>600</sub> was measured using TECAN microplate reader (don't forget to measure a M9 sample for blanking!)<br> |
| | + | <em>NB: from now on, cultures must be placed in ice to stop cell growth.</em></li> |
| | + | <li>At the same time, proper dilution of the cultures were plated on LB agar plates.<br> |
| | + | <em>NB: All the dilutions are performed moving 100 μl of culture in previously ice-chilled 900 μl fresch M9. 100 μl of the final dilution are plated (It still represents a 1:10 dilution!)</em></li> |
| | + | <li>Plates were grown overnight and next morning C.F.U. were counted.</li> |
| | + | <li>C.F.U. values were corrected by the dilution factor and a linear regression (N vs O.D.<sub>600</sub>) was performed in order to evaluate the conversion factor <b>K<sub>O.D.toC.F.U.</sub></b>. </li> |
| | + | <li><b>K<sub>O.D.toC.F.U.</sub></b> was used as conversion factor to multiply the O.D.<sub>600</sub> value of saturation in the growth curves (~0,5). </li> |
| | + | </ul> |
| | + | </ol> |
| | + | <p>The results are summarized in the table and in the figure below: </p> |
| | + | <center> |
| | + | <table class='data'> |
| | + | <tr> |
| | + | <td class='row'><b> Culture </b></td> |
| | + | <td class='row'><b> O.D.<sub>600</sub> TECAN </b></td> |
| | + | <td class='row'><b> O.D.<sub>600</sub> Spectrophotometer </b></td> |
| | + | <td class='row'><b> C.F.U. </b></td> |
| | + | <td class='row'><b> Dilution Factor (10^) </b></td> |
| | + | <td class='row'><b> N </b></td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td class='row'> C1 </td> |
| | + | <td class='row'> 0,367950002 </td> |
| | + | <td class='row'> 0,758004416 </td> |
| | + | <td class='row'> 990 </td> |
| | + | <td class='row'> 5 </td> |
| | + | <td class='row'> 990000000 </td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td class='row'> C1 </td> |
| | + | <td class='row'> 0,044649998 </td> |
| | + | <td class='row'> 0,091982322 </td> |
| | + | <td class='row'> 141 </td> |
| | + | <td class='row'> 6 </td> |
| | + | <td class='row'> 1410000000 </td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td class='row'> C1 </td> |
| | + | <td class='row'> 0,004799999 </td> |
| | + | <td class='row'> 0,009888356 </td> |
| | + | <td class='row'> 136 </td> |
| | + | <td class='row'> 5 </td> |
| | + | <td class='row'> 136000000 </td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td class='row'> C1 </td> |
| | + | <td class='row'> 0,000549998 </td> |
| | + | <td class='row'> 0,001133037 </td> |
| | + | <td class='row'> 20 </td> |
| | + | <td class='row'> 6 </td> |
| | + | <td class='row'> 200000000 </td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td class='row'> C2 </td> |
| | + | <td class='row'> 0,54840003 </td> |
| | + | <td class='row'> 1,129744917 </td> |
| | + | <td class='row'> 165 </td> |
| | + | <td class='row'> 4 </td> |
| | + | <td class='row'> 16500000 </td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td class='row'> C2 </td> |
| | + | <td class='row'> 0,058200002 </td> |
| | + | <td class='row'> 0,119896339 </td> |
| | + | <td class='row'> 23 </td> |
| | + | <td class='row'> 5 </td> |
| | + | <td class='row'> 23000000 </td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td class='row'> C2 </td> |
| | + | <td class='row'> 0,008700002 </td> |
| | + | <td class='row'> 0,017922652 </td> |
| | + | <td class='row'> 251 </td> |
| | + | <td class='row'> 3 </td> |
| | + | <td class='row'> 2510000 </td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td class='row'> C2 </td> |
| | + | <td class='row'> 0,000100002 </td> |
| | + | <td class='row'> 0,000206011 </td> |
| | + | <td class='row'> 24 </td> |
| | + | <td class='row'> 4 </td> |
| | + | <td class='row'> 2400000 </td> |
| | + | </tr> |
| | + | </table> |
| | + | </center> |
| | + | <br> |
| | + | </p> |
| | + | <center> |
| | + | <img alt="" src="https://static.igem.org/mediawiki/2011/3/35/UNIPV_ODvsCFU.png" width="90%"> |
| | + | </center> |
| | + | <p>The estimation of μ parameter was performed by determining the slope of the logarithmic curve of O.D.<sub>600</sub> in exponential phase. Exponential phase was determined by visual inspection as the linear phase of the logarithmic curve of O.D.<sub>600</sub>. <br> |
| | + | <br> |
| | + | The estimated parameters are summarized in the table below: <br> |
| | + | </p> |
| | + | <center> |
| | + | <table width='50%' class='data'> |
| | + | <tr> |
| | + | <td class='row'><b>N<sub>max</sub> [cell number]</b></td> |
| | + | <td class='row'><b>μ [min<sup>-1</sup>]</b></td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td class='row'>1*10<sup>9</sup></td> |
| | + | <td class='row'>0.004925</td> |
| | + | </tr> |
| | + | </table> |
| | + | </center> |
| | + | <p> </p> |
| | + | <p>The reported value of μ corresponds to a doubling time of 142 minutes. </p> |
| | + | <div align="right"><small><a href="#indice" title="">^top</a></small></div> |
| | + | |
| | + | <!--------------------------------> |
| | + | <!--------------------------------> |
| | + | <!--------------------------------> |
| | + | <!--------------HSL---------------> |
| | + | <!--------------------------------> |
| | + | <!--------------------------------> |
| | + | <!--------------------------------> |
| | + | |
| | + | <a name='HSL'></a> |
| | + | <h2>Estimation of the spontaneous degradation of HSL in M9 medium and in cultures at different pH values</h2> |
| | + | <p align='justify'> In order to estimate the spontaneous degradation rate of HSL in M9 medium and in a culture of MGZ1 cells as a function of pH, two simple tests have been performed.<br> |
| | + | Two different M9 media were prepared, one with the nominal pH (7.0) and one with pH=6.0.<br> |
| | + | These media, now named respectively M9<sub>pH 7</sub> and M9<sub>pH 6</sub>, were added with a known concentration of HSL (100 nM) and then incubated at 37°C, 220 rpm (NB: the media were not infected with any culture but the standard growth conditions were reproduced). The amount of HSL present in the medium was assayed through the BBa_T9002 biosensor at 4 time points: <br> |
| | + | <ul> |
| | + | <li>t=0 h;</li> |
| | + | <li>t=1 h;</li> |
| | + | <li>t=2 h;</li> |
| | + | <li>t=4 h;</li> |
| | + | <li>t=8 h;</li> |
| | + | </ul> |
| | + | <p>The obtained time series of HSL amounts were processed to evaluate the time constant governing the dynamic of HSL degradation, supposing an exponential decay. The results are reported in the table below: </p> |
| | + | <center> |
| | + | <table class='data'> |
| | + | <tr> |
| | + | <td class='row'></td> |
| | + | <td class='row'><b>t<sub>1/2</sub><sup>*</sup> [h]</b></td> |
| | + | <td class='row'><b>γ<sub>HSL</sub><sup>**</sup> [h<sup>-1</sup>]</b></td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td class='row'><b>M9<sub>pH 6</sub></b></td> |
| | + | <td class='row'>32 </td> |
| | + | <td class='row'>0.022 </td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td class='row'><b>M9<sub>pH 7</sub></b></td> |
| | + | <td class='row'>8 </td> |
| | + | <td class='row'>0.087 </td> |
| | + | </tr> |
| | + | </table> |
| | + | </center> |
| | + | <p> The described experiment was repeated with a MGZ1 culture in order to evaluate the effect of culture on HSL stability. The estimated values are reported in the table below:</p> |
| | + | <center> |
| | + | <table class='data'> |
| | + | <tr> |
| | + | <td class='row'></td> |
| | + | <td class='row'><b>t<sub>1/2</sub><sup>*</sup> [h]</b></td> |
| | + | <td class='row'><b>γ<sub>HSL</sub><sup>**</sup> [h<sup>-1</sup>]</b></td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td class='row'><b>Culture<sub>pH 6</sub></b></td> |
| | + | <td class='row'>∞ </td> |
| | + | <td class='row'>0 </td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td class='row'><b>Culture<sub>pH 7</sub></b></td> |
| | + | <td class='row'>19 </td> |
| | + | <td class='row'>0.037 </td> |
| | + | </tr> |
| | + | </table> |
| | + | </center> |
| | + | <p>It is evident from the reported data that the spontaneous degradation of HSL is negligible when CTRL+E is implemented in MGZ1 cells grown in M9 medium with pH=6.0 and pH=7.0.<br> |
| | + | Thus in the simulations we set γ<sub>HSL</sub>=0; </p> |
| | + | </p> |
| | + | <div align="right"><small><a href="#indice" title="">^top</a></small></div> |
| | + | |
| | + | <!--------------------------------> |
| | + | <!--------------------------------> |
| | + | <!--------------------------------> |
| | + | <!-------------t9002--------------> |
| | + | <!--------------------------------> |
| | + | <!--------------------------------> |
| | + | <!--------------------------------> |
| | + | |
| | + | <a name='t9002'></a> |
| | + | <h2>Characterization of BBa_T9002 biosensor</h2> |
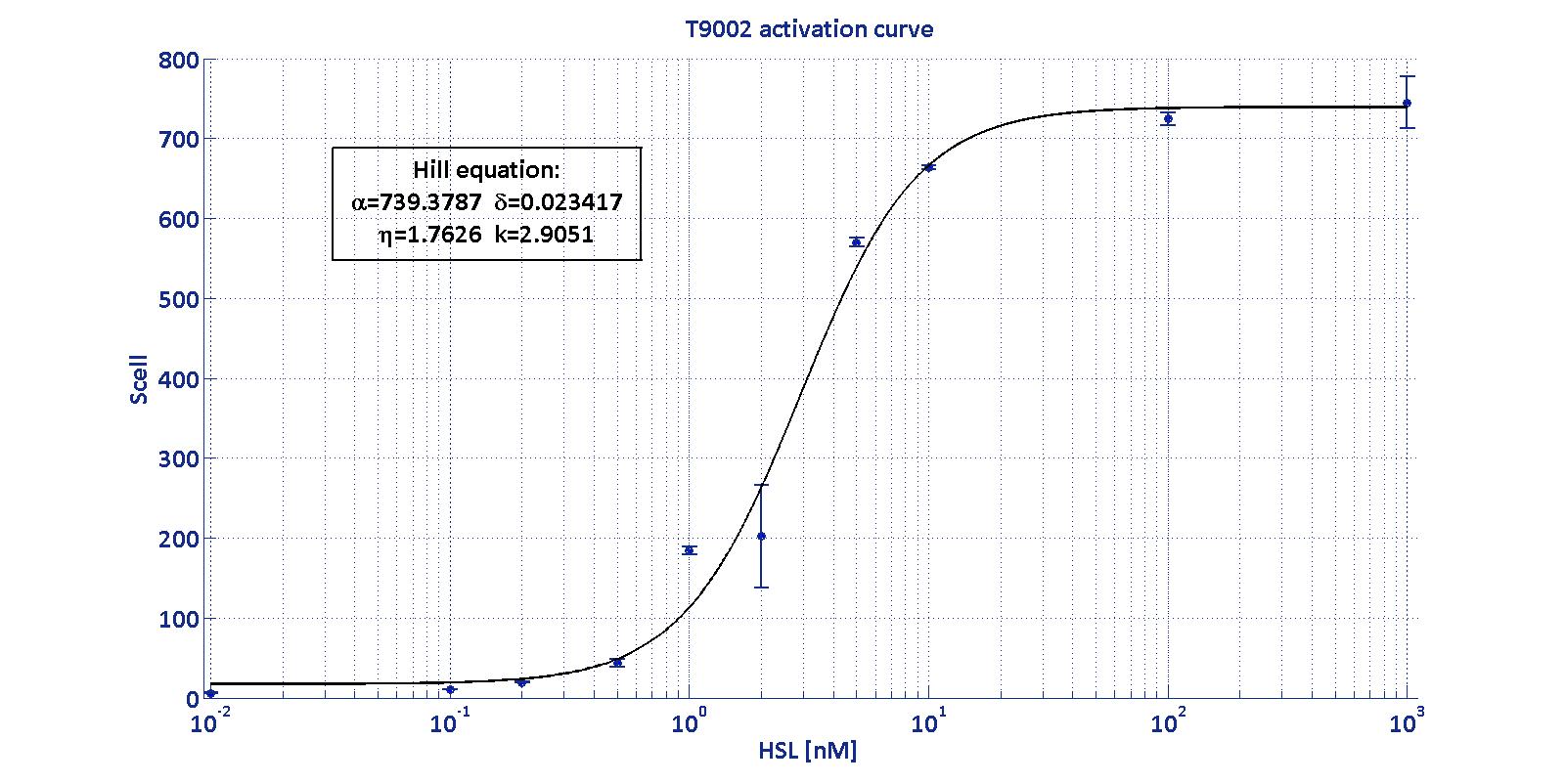
| | + | As described in the <a href="https://2011.igem.org/Team:UNIPV-Pavia/Project/Modelling#introduction_to_T9002">modelling section</a>, BioBrick <a href="http://partsregistry.org/Part:BBa_T9002">BBa_T9002</a> is an HSL biosensor, which provides a non linear relationship between HSL input and S<sub>cell</sub> output. More precisely, the characteristic sigmoidal curve requires synthetic parameters for its accurate identification. These are the minimum and maximum values, the swtich point (i.e., the curve inflection point), and the upper and lower boundaries of linearity. This biosensor revealed greatly reliable, providing measurement repeatability and minimal experimental noise. Referring to its activation formula, the calibration curve is shown below.<br> |
| | + | <br> |
| | + | <div style='text-align:center'> |
| | + | <div class="thumbinner" style="width:100%;"> <img alt="" src="https://static.igem.org/mediawiki/2011/5/50/Activation_T9002.jpg" class="thumbimage" width="47%" height="50%"></a></div> |
| | + | </div> |
| | + | <br> |
| | + | <div style='text-align:center'> |
| | + | <div class="thumbinner" style="width:100%;"> <img alt="" src="https://static.igem.org/mediawiki/2011/4/4e/T9002_activation.jpg" class="thumbimage" width="100%"></a></div> |
| | + | </div> |
| | + | <center> |
| | + | <table width='50%' class='data'> |
| | + | <tr> |
| | + | <td class='row'><b>Minimum [S<sub>cell</sub>]</b></td> |
| | + | <td class='row'><b>Maximum [S<sub>cell</sub>]</b></td> |
| | + | <td class='row'><b>Switch point [nM]</b></td> |
| | + | <td class='row'><b>Lower boundary of linearity [nM]</b></td> |
| | + | <td class='row'><b>Upper boundary of linearity [nM]</b></td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td class='row'>17.31</td> |
| | + | <td class='row'>739.4</td> |
| | + | <td class='row'>1.39</td> |
| | + | <td class='row'>0.38</td> |
| | + | <td class='row'>5.07</td> |
| | + | </tr> |
| | + | </table> |
| | + | </center> |
| | + | <br> |
| | + | <br> |
| | + | In order to determine the threshold sensitivity of T9002 biosensor, experiments were performed with several HSL inductions minimally interspaced in the region of low detectability. Hypothesizing that the inducer is 1:20 diluted (as for all of our tests), the minimum detectable HSL concentration is 3 nM. |
| | + | <div align="right"><small><a href="#indice" title="">^top</a></small></div> |
| | + | <br> |
| | <br> | | <br> |
| - |
| |
| - |
| |
| - |
| |
| - | <p style="text-align:justify;">
| |
| - | ho separato main<br>anche top è stato inserito ..
| |
| | <br> | | <br> |
| | | | |
| - | One of the pivotal objectives of synthetic biology is to build complex gene networks with a predictable behavior by combining well-characterized basic modules. </p>
| + | </html> |
| - | <p style="text-align:justify;">As a proof of concept of this fundamental, our project aims at designing and implementing in <em>E. coli</em> a quorum sensing-based control system, able to regulate the concentration of a signaling molecule (3OC6-HSL) via a negative feedback loop. </p>
| + | |
| - | <p style="text-align:justify;">In order to obtain the desired output, fine-tuning of the circuit is necessary; therefore, a mathematical model of the control system will be derived and identified by data coming from <em>ad hoc</em> experiments performed on basic modules. Model simulations will be used to meet the design specifications of the biological controller, demonstrating that full characterization of basic parts is a major goal to predict the behavior of more complex circuits.</p>
| + | |
| | | | |
| | | | |
| - |
| |
| - |
| |
| - | </html>
| |
| | | | |
| | {{end}} | | {{end}} |

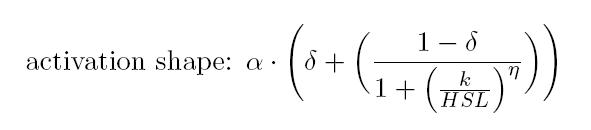
 "
"










