Team:Paris Liliane Bettencourt/Notebook/2011/08/07/
From 2011.igem.org
(Difference between revisions)
(Created page with "== Cyrille == Treatement using inkscape of the gels done yesterday. In this picture, the progressive explanation of the identification of the bands. File:CP0807_analysis.jpg") |
|||
| (13 intermediate revisions not shown) | |||
| Line 1: | Line 1: | ||
| + | {{:Team:Paris_Bettencourt/tpl_test}} | ||
| + | |||
== Cyrille == | == Cyrille == | ||
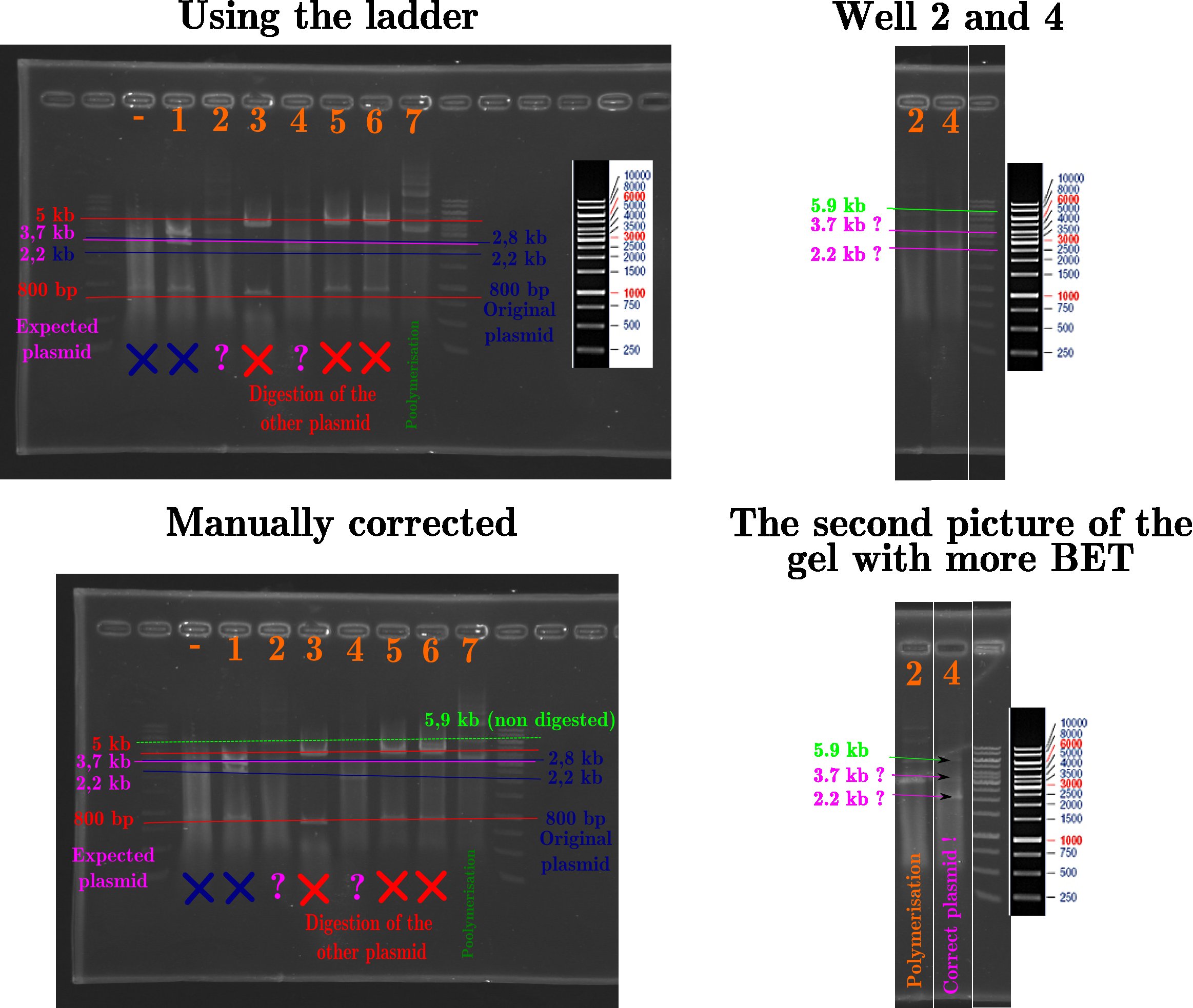
Treatement using inkscape of the gels done yesterday. In this picture, the progressive explanation of the identification of the bands. | Treatement using inkscape of the gels done yesterday. In this picture, the progressive explanation of the identification of the bands. | ||
| - | [[File:CP0807_analysis.jpg]] | + | [[File:CP0807_analysis.jpg|400px|thumb|center|Analysis]] |
| + | |||
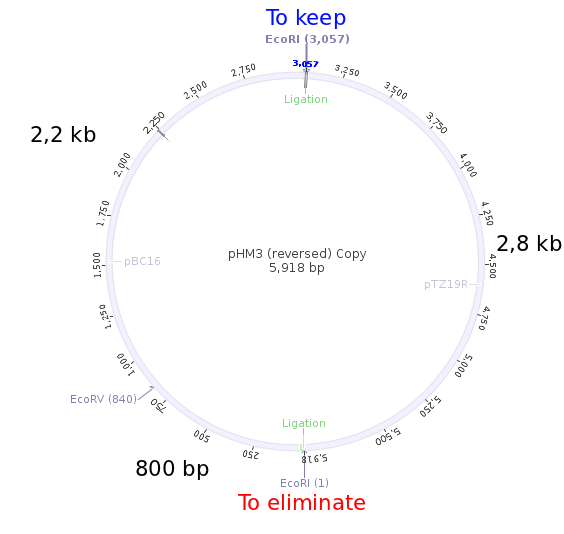
| + | So the miniprep 4 seems to contain the plasmid with the EcoRI at the good place! To understand the process, here is the plasmid map. | ||
| + | |||
| + | [[File:CP0807_plasmidmap.jpg|400px|thumb|center|pHM3 map]] | ||
| + | |||
| + | == Kevin == | ||
| + | |||
| + | === Miniprep from glycerol === | ||
| + | To re-test pDAG464 and pDAG479 plasmids, culture of them overnight has been done and miniprep. Quantification by TECAN : | ||
| + | *pDAG464 : 122,9 ng/uL | ||
| + | *pDAG479 : 204,9ng/uL | ||
| + | |||
| + | === Gel verification of Dave Lane plasmids (suite)=== | ||
| + | In a 1% agar gel, we have test pDAG464 and pDAG479 plasmids digested by XbaI and PstI enzyme. We've look if lines that we should see are the good one. | ||
| + | |||
| + | [[File:gelverif2dlplasmid.jpg|450px|thumb|center|Gel of verification Dave Lane plasmids digestion]] | ||
| + | |||
| + | *Column 1 : expected 450/833/950/1545/2110 pb | ||
| + | *Column 2 : expected 402/833/950/1545/2110 pb | ||
| + | |||
| + | I don't understand we don't have what we expected. I will talk about it tomorrow with the team. | ||
| + | ==Danyel & Camille== | ||
| + | ===T7 amber=== | ||
| + | The digested PCR products were transformed in MH1 cells and plated, then incubated at 37ºC overnight. | ||
Latest revision as of 16:08, 27 August 2011

Contents |
Cyrille
Treatement using inkscape of the gels done yesterday. In this picture, the progressive explanation of the identification of the bands.
So the miniprep 4 seems to contain the plasmid with the EcoRI at the good place! To understand the process, here is the plasmid map.
Kevin
Miniprep from glycerol
To re-test pDAG464 and pDAG479 plasmids, culture of them overnight has been done and miniprep. Quantification by TECAN :
- pDAG464 : 122,9 ng/uL
- pDAG479 : 204,9ng/uL
Gel verification of Dave Lane plasmids (suite)
In a 1% agar gel, we have test pDAG464 and pDAG479 plasmids digested by XbaI and PstI enzyme. We've look if lines that we should see are the good one.
- Column 1 : expected 450/833/950/1545/2110 pb
- Column 2 : expected 402/833/950/1545/2110 pb
I don't understand we don't have what we expected. I will talk about it tomorrow with the team.
Danyel & Camille
T7 amber
The digested PCR products were transformed in MH1 cells and plated, then incubated at 37ºC overnight.
 "
"