Team:Edinburgh/Data
From 2011.igem.org
Data Overview
The iGEM rules require us to have simple illustrations of how our devices work and where the Parts function in the system; and links to the Registry for the parts/constructs for which we have produced data.
See the sample Data page.
Contents |
Overview
This page provides an overview of the purely biological aspects of the project.
Cell Surface Display System
(Further details are at the dedicated Cell Surface Display page.)
This system aims at achieving synergy between the enzymes by displaying them at high copy number on the cell's outer membrane. Ice Nucleation Protein is used as a carrier for display of the enzymes; it carries them to the outer membrane.
Schematic diagram
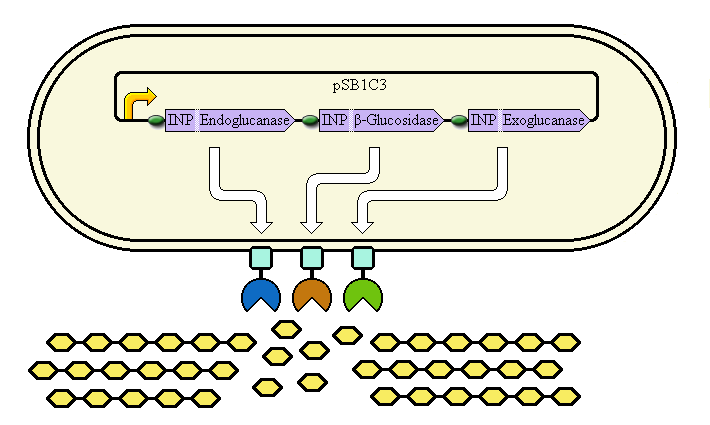
The completed system should contain:
A promoter (<partinfo>BBa_K523000</partinfo>) controlling:
an INP—Endoglucanase fusion (<partinfo>BBa_K523008</partinfo> + <partinfo>BBa_K523011</partinfo>)
an INP—β-glucosidase fusion (<partinfo>BBa_K523008</partinfo> + <partinfo>BBa_K523010</partinfo>)
an INP—Exoglucanase fusion (<partinfo>BBa_K523008</partinfo> + <partinfo>BBa_K523009</partinfo>)
Ribosome Binding Sites are indicated as green ovals.
Cellulose degradation is shown at the bottom. In reality, tens of thousands of enzymes will cover the outer membrane in random places.
A test system to prove that <partinfo>BBa_K523008</partinfo> can be used to carry proteins to the outer membrane uses a fusion of INP to Yellow Fluorescent Protein (YFP) or the E. coli amylase MalS (<partinfo>BBa_K523003</partinfo>) instead.
Phage Display System
(Further details are at the dedicated Phage Display page.)
This system aims at achieving synergy between the enzymes by displaying them on an M13 phage. The major coat protein pVIII is used as a carrier for display of the enzymes; it incorporates them into the phage.
Schematic diagram
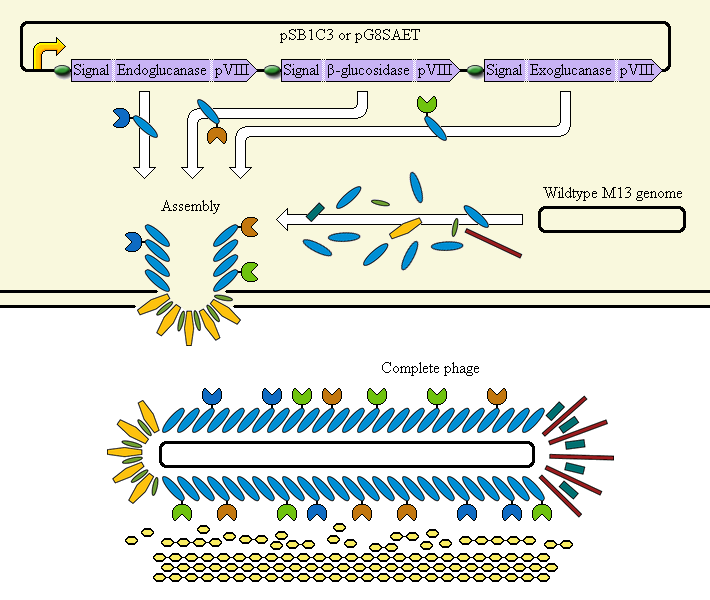
The completed system should contain:
A promoter (<partinfo>BBa_K523000</partinfo>) controlling:
an Endoglucanase—pVIII fusion
a β-glucosidase—pVIII fusion
an Exoglucanase—pVIII fusion
Ribosome Binding Sites are indicated as green ovals. "Signal" means a periplasmic signal sequence, directing the protein to the periplasm to be assembled into the phage.
A test system uses a fusion of pVIII to E. coli amylase MalS instead.
 "
"