Team:Edinburgh/Experiments
From 2011.igem.org
Experiments
Actual results of experiments are placed here, but more detailed discussions are also found on the relevant Registry pages.
Contents |
malS
We placed malS (an E. coli amylase gene) under the control of the lac promoter and grew the cells on starch agar. An iodine-based assay was carried out testing for starch degradation.
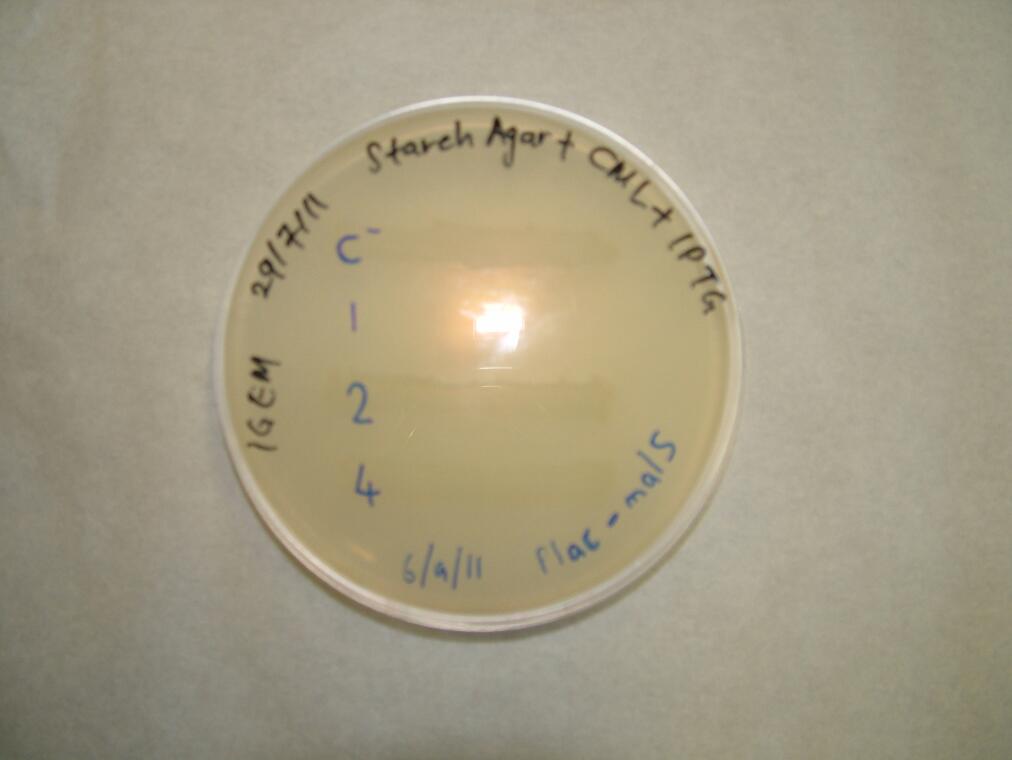
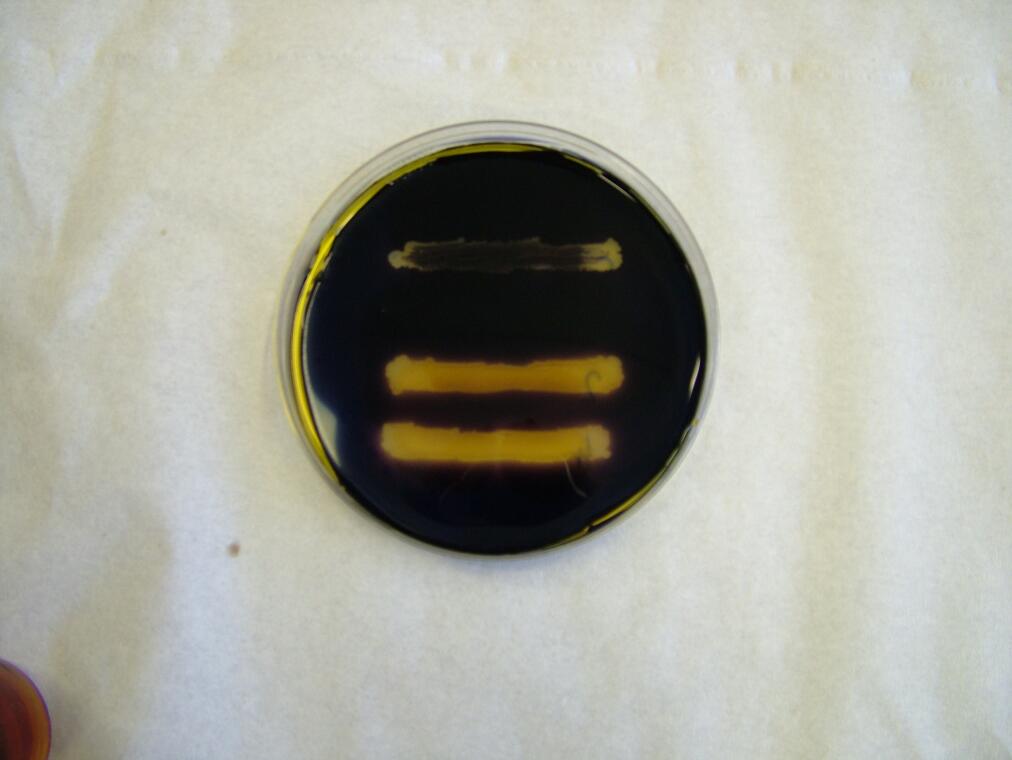
| 
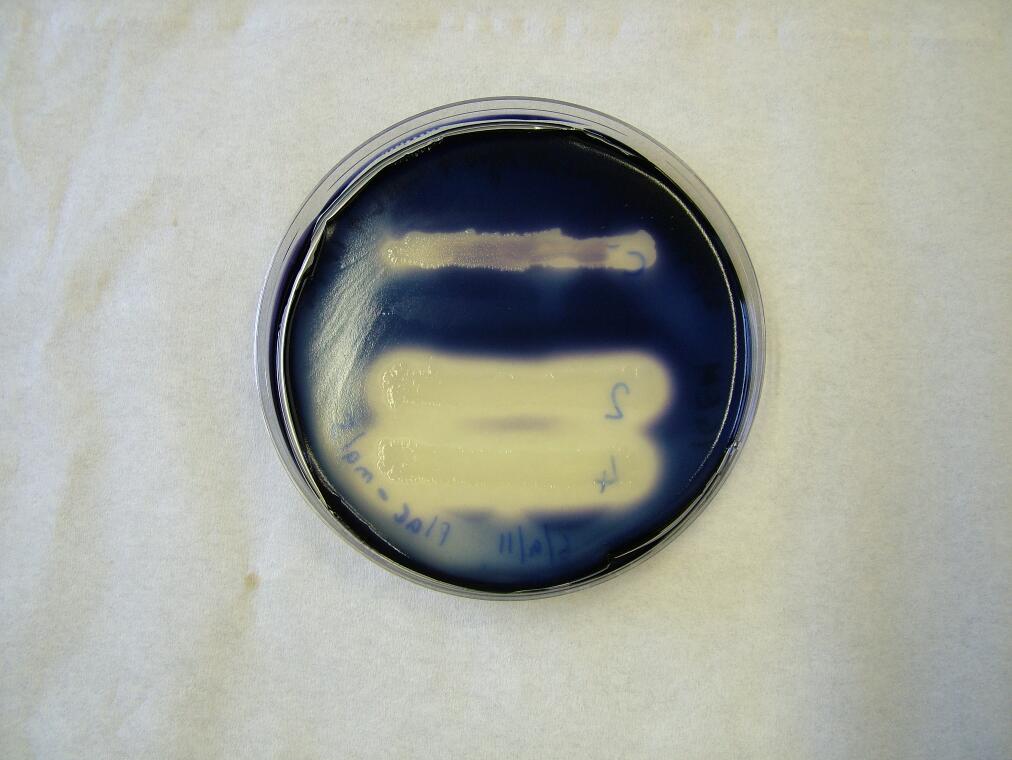
| 
| ||
| Before the assay. Colony 1 failed to grow. Control at top. | Immediately after iodine flooding. | 40 minutes after iodine flooding. |
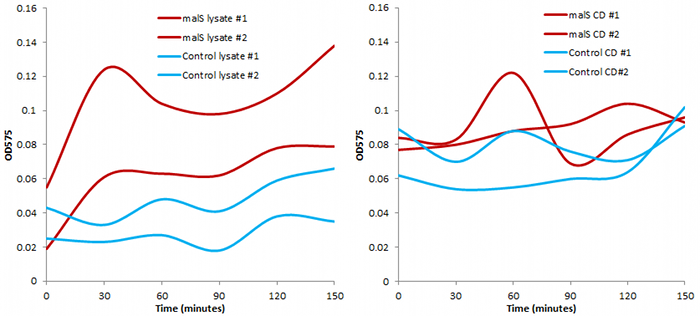
We also conducted an assay based on cell extract's ability to degrade starch, producing glucose that would react with 3,5-dinitrosalicylic acid.
Finally, we tested the ability of bacteria with overexpression of malS to grow on a starch agar with no other carbon source. Indeed it could, while normal E. coli cannot:
For more information, see the Registry page on BBa_K523006.
bglX and cex
We placed bglX (a cryptic E. coli β-glucosidase gene) under the control of the lac promoter and compared its activity to the exoglucanase cex.
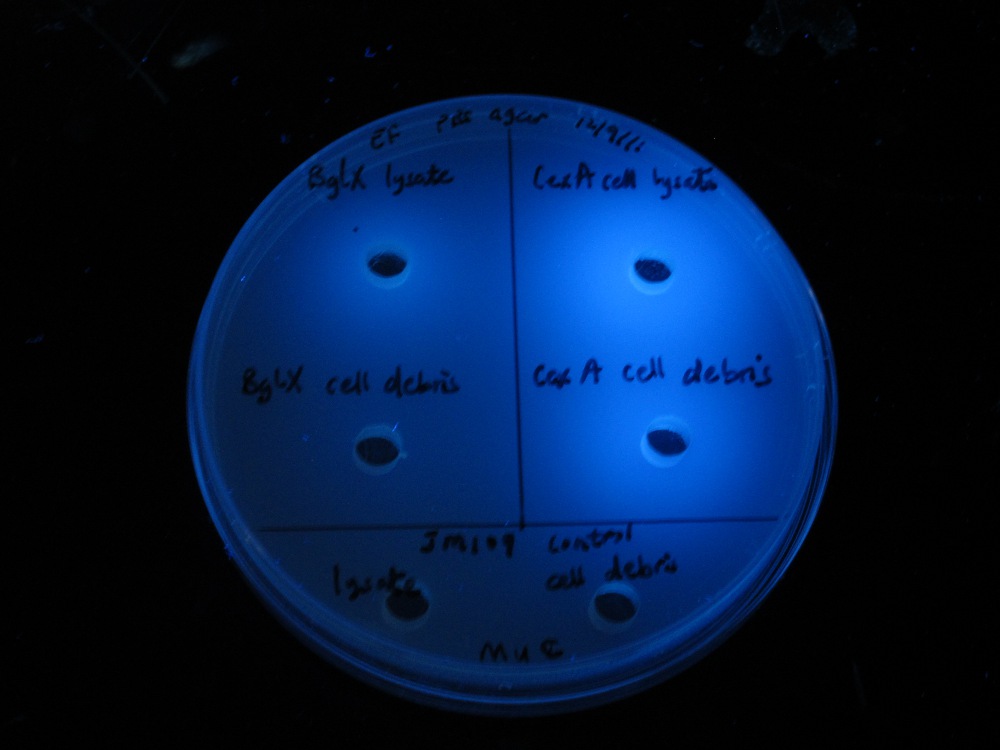
| 
| |
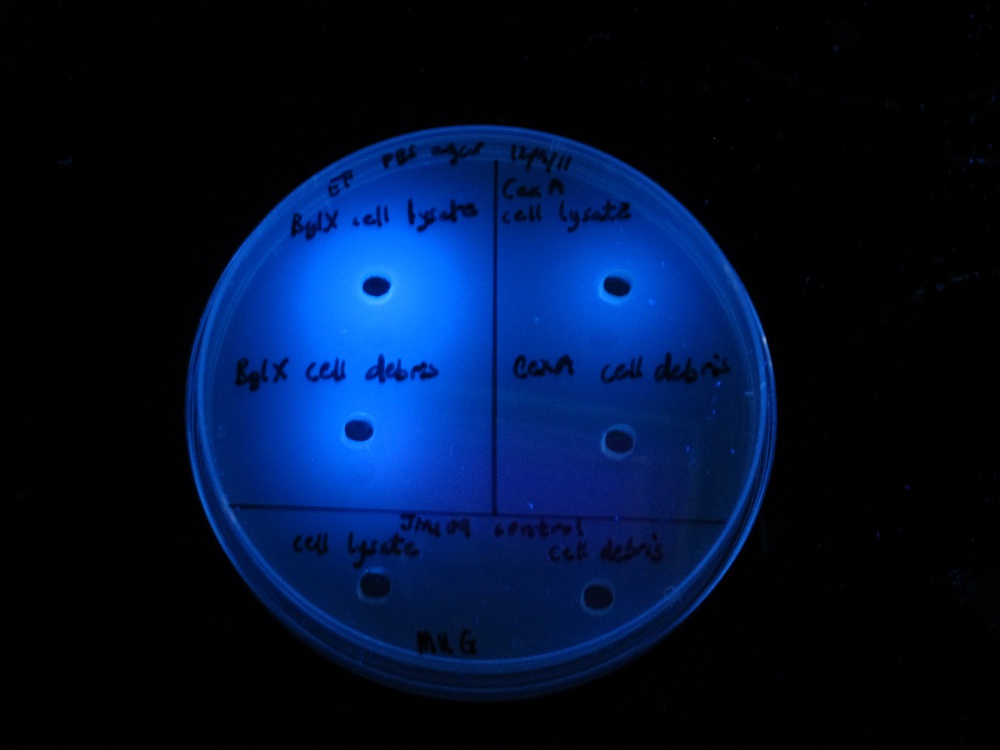
| MUG assay. bglX on left, cex on right. | MUC assay. bglX on left, cex on right. |
bglX was capable of degrading the substrate MUG, which has a β(1→4) bond, similar to that of cellobiose.
We tested the ability of E. coli with this bglX part to grow on minimal media with cellobiose as the only carbon source. It could not. By contrast, it did grow if glucose was the carbon source, showing that it is fundamentally capable of growing on minimal media:
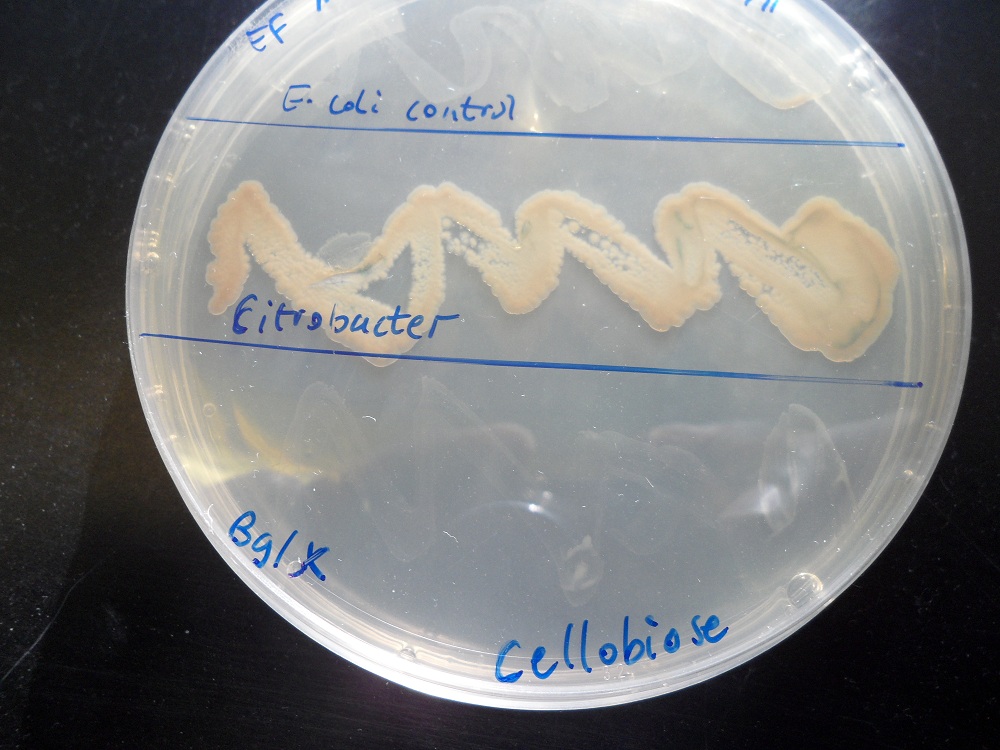
| 
| |
| Cellobiose as the only carbon source. K523014 (bottom) fails to grow. | Glucose as the only carbon source. K523014 can grow. |
For more information, see the Registry page on BBa_K523014.
INP-EYFP
We made a fusion (BBa_K523013) of Ice Nucleation Protein and Enhanced Yellow Fluorescent Protein, and placed it under the control of the Lac promoter.
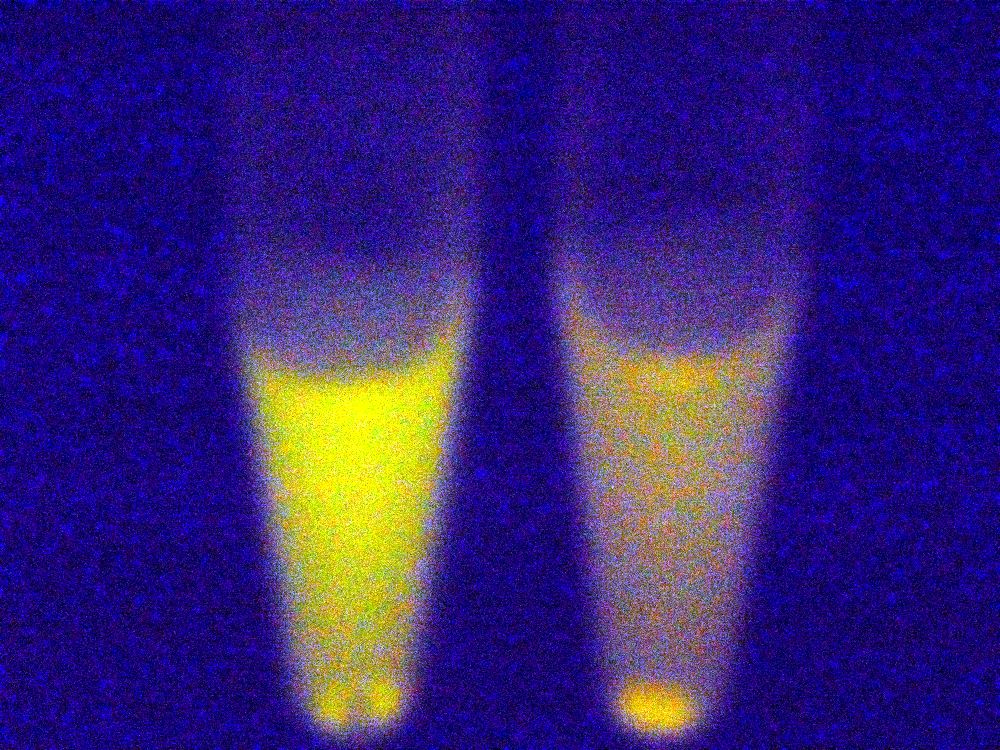
Under blue light, the cells fluoresced yellow, as shown below (left image). Unfortunately, our imaging technology was not capable of showing whether the fluorescence was localised to the outer membrane, so we centrifuged the cells so that the membrane fraction was localised to the bottom of the tube and compared the result to a colony expressing EYFP without INP. This showed that the INP-EYFP was indeed localised to the cell membrane.
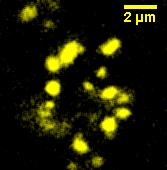
| 
| 
| ||
| Cells fluorescing yellow. | Centrifuged cells. Control on left. INP-EYFP on right. | The same image passed through GIMP's auto white balance filter. |
For more information, see the Registry page on BBa_K523013.
cxnA
In line with our beads on a string proposal, We created a fusion of cex and cenA. This showed the activity of both enzymes:
- Cex activity was tested on MUC plates.
- CenA activity was tested on carboxymethylcellulose (CMC) plates.
The latter test produces a zone of clearing if CMC is degraded by the endoglucanase domain. We had three successful colonies:
For more information, see the Registry page on BBa_K523025.
BioSandwich
The BioSandwich system is outlined on its own page. BioSandwich creates parts with homologous ends, but there are a number of ways in which the final assembly could be carried out. Throughout the project we mostly used Gibson Assembly.
Parts made with BioSandwich
Parts that we believe are correct (subject to final verification sequencing) include:
- BBa_K523013: PlacLacZ-INP-YFP
- BBa_K523019: PlacLacZ-INP-cex [update: sequencing shows this part has errors]
- BBa_K523020: PlacLacZ-INP-bglX [update: sequencing shows this part has errors]
- BBa_K523022: PlacLacZ-crtEIB
Extensive tests - what can go wrong?
We also tested a number of final assembly methods, by attempting to join a promoter and LacZ construct to a Yellow Fluorescent coding sequence, making BBa_K523023. This part will produce a polycistronic RNA transcript, coding for proteins that turn the bacteria blue (on Xgal) but also make it fluoresce yellow.
BioSandwich inserts short spacers between each part to be assembled. Note that after spacer2 we expect to see some extra bases before we reach the RFC10 suffix, because of the choice of vector. We can call these extra bases the "stuffing". So, the expected sequence is:
- Vector -- spacerT7 -- Plac,LacZ -- spacer1 -- EYFP -- spacer2 -- stuffing
Successful assemblies are expected to generate blue colonies that fluoresce yellow under a blue light. We tried several assembly methods, all of which generated many colonies with the correct phenotype, but also many colonies with an incorrect phenotype...
From each plate we sequenced a colony with the correct phenotype (if there were any) and some colonies with incorrect phenotypes, in an attempt to understand what the failure modes were.
BioSandwich with Gibson Assembly
Like, OEPCR, this method produces many correct colonies and many incorrect colonies.
The Gibson method was described by Gibson et al (2009). A gentle introduction was written by the 2010 Cambridge iGEM team, and is found here.
We sequenced a colony with the correct phenotype, as well as others with incorrect phenotypes:
- Colony "gby" [blue, yellow] - expected scar is missing at approx base 680 of the forward chromatogram. Also, the spacer ends with ATG and the part starts with ATG, but only one ATG is seen. In total 9 bases are missing. There is homology of "tatgg" at both ends of the deleted fragment. There is a point mutation at approx base 500 of the forward chromatogram.
- Colony "gb" [blue] - there is a deletion of about 80 bases at approx base 700 of the chromatogram. This has been caused by homology of a "gggcgagg" found at both ends of the deleted sequence. The 9 bases mentioned for gby are also missing.
- Colony "gwy" [yellow] - seems correct except for a point mutation in the reverse chromatogram. The mutation is synonymous (ctg -> ttg). There's no clear reason why the colony wasn't blue.
Lessons learned: bits of homology, even quite small, can cause problems with this method.
BioSandwich with Overlap Extension PCR
Overlap Extension PCR can be performed using spoligos as primers. The extension time should be calculated from the entire product size. The final product will not be circular, so can be run on a gel to test its length. Later, it will need to have its ends ligated.
This method produces many correct colonies, although it also produces many incorrect colonies.
[For our tests, different primers were used and spacer2 was not needed.]
- Colony "bby" [blue, yellow] - correct sequence.
- Colony "bb" [blue] - has a massive deletion (several hundred bases) in the EYFP gene after about 60 bases. There is partial homology between the start and end of the deletion: "tggtcgagctggac" vs "tggacgagctgtac".
- Colony "bw" [plain] - has spacerT7 joined to spacer1, and the same massive deletion as bb.
Again homology seems to be an issue. Parts joining randomly (in bw) is worrying.
BioSandwich with Ligation Independent Cloning
In our experience, this method has been only weakly successful. We have seen one correct colony in several attempts.
To attempt the LIC method, simply mix the parts and vector (4 uL of each). Heat a beaker of water to 95 C, and float the reaction tube in it, and allow everything to cool. Afterwards, it may now be necessary to centrifuge the tube briefly to move the liquid back to the bottom.
- Colony "b1" [blue, yellow] - sequence read is of low quality but things seem correct.
- Colony "y2" [yellow] - appears to simply be BBa_K216011, the template for the initial PCR
- Colony "n4" [plain] - sequence is unreadable.
The y2 colony is probably expressing a template for PCR which made it all the way through the various steps that came afterwards.
BioSandwich with Blunt-End Ligation Independent Cloning
This is expected to fail as Ligation Independent Cloning requires long overhangs at the end of each part, not blunt ends. No correct colonies were seen.
- Colony "bb1" [blue] - seems to be BBa_J33207 followed by part of E. coli ferrous iron transport gene A.
- Colony "bw2" [plain] - has spacerT7 followed by the stuffing.
- Colony "bw3" [plain] - has RFC10 prefix followed by RFC10 suffix, as if XbaI and SpeI sites have joined in a normal vector.
BioSandwich with CPEC: Circular Polymerase Extension Cloning
- Colony "cby" [blue, yellow] - there is a single-base deletion near the start of the PlacLacZ part, with no obvious cause. It is too early to affect expression. The last few bases of spacer1 are missing, as well as the first few bases of the EYFP part (9 bases total). This is explained by homology of "tatgg". Otherwise seems correct.
- Colony "cb" [blue] - Normal until the end of spacer1, which is followed immediately by spacer2 and the stuffing.
- Colony "cw" [plain] - spacerT7 is followed immediately by a second copy of spacerT7, after which comes spacer2 and a second copy of spacer2. Then the stuffing.
This gave some of the weirder results.
Semi-quantitative results
Our lab supervisor Chris French recorded the results of some early BioSandwich assembly attempts. Since correct assemblies of the construct described above (PlacLacZ-EYFP) should produce blue colonies that fluoresce yellow under blue light, we can get a rough idea of how many colonies are correct by counting phenotypes.
- BioSandwich with OEPCR
- Assembly produced around 100 colonies on a 100 microlitre plate, about half of these blue, and most of the blue also yellow.
- BioSandwich with Gibson
- Assembly produced about 100-200 colonies on a 100 microlitre plate, about half blue and half white, and the blue ones about 75% yellow.
- BioSandwich with CPEC
- Assembly produced 60 colonies, of which 26 were blue, of which half were also yellow.
Various assemblies
The core team made a number of constructs with PlacLacZ plus other genes. While these other genes usually did not give an easily spotted phenotype, we can at least see how many of them were blue on Xgal.
As an explanation, "100 plate" and "900 plate" refer to the concentration of transformed cells plated.
- BioSandwich with OEPCR
- pSB1C3-PlacLacZ-INP-YFP (should be blue and yellow)
- 100 plate> 1 white+yellow
- 900 plate> 1 blue, 7 white+yellow, 1 white
- pSB1C3-PlacLacZ-INP-MalS (should be blue)
- 100 plate> 188 white, 7 blue
- 900 plate> 165 blue, 376 white
- BioSandwich with Gibson
- pSB1C3-PlacLacZ-INP-cex (should be blue)
- 100 plate> 46 white, 3 blue
- 900 plate> 929 white, 48 blue
- pSB1C3-PlacLacZ-INP-cex (should be blue)
- 100 plate> 38 white, 2 blue, 1 red
- 900 plate> 633 white, 61 blue, 2 red
- pSB1C3-PlacLacZ-INP-bglX (should be blue)
- 100 plate> 57 white
- 900 plate> 575 white, 1 blue
- pSB1C3-PlacLacZ-INP-bglX (should be blue)
- 100 plate> 54 white, 16 blue
- 900 plate> 423 white, 156 blue
- pSB1C3-PlacLacZ-crtEIB (should be blue, or maybe red)
- 100 plate> 2 white
- 900 plate> 1 blue, 80 white
- pSB1C3-PlacLacZ-INP-malS (should be blue)
- 100 plate> 16 blue, 74 white
- BioSandwich with CPEC
- pSB1C3-PlacLacZ-crtEIB --- 7 cycles (should be blue, or maybe red)
- 100 plate> 31 white, 19 blue
- 900 plate> 257 white, 149 blue
- pSB1C3-PlacLacZ-crtEIB --- 3 cycles (should be blue, or maybe red)
- 100 plate> 1 white
- 900 plate> 15 white, 4 blue
- pSB1C3-PlacLacZ-INP-bglX ---7 cycles (should be blue)
- 100 plate> 21 white
- 900 plate> 363 white, 1 red, 1 blue
- pSB1C3-PlacLacZ-INP-bglX ---3 cycles (should be blue)
- 100 plate> 75 white, 4 blue
- 900 plate> 494 white, 36 blue, 2 red.
- pSB1C3-PlacLacZ-INP-YFP (should be blue and yellow)
- 100 plate> 15 white
- 900 plate> 74 white, 1 blue+yellow
- pSB1C3-PlacLacZ-INP-MalS (should be blue)
- 100 plate> 6 white, 4 blue
- 900 plate> 32 blue, 127 white
It seems our supervisor with his 15 years postdoctoral experience had slightly better results than we did. Hmm...
 "
"