Team:Bielefeld-Germany/Results/S-Layer
From 2011.igem.org


Contents |
S-layer in general
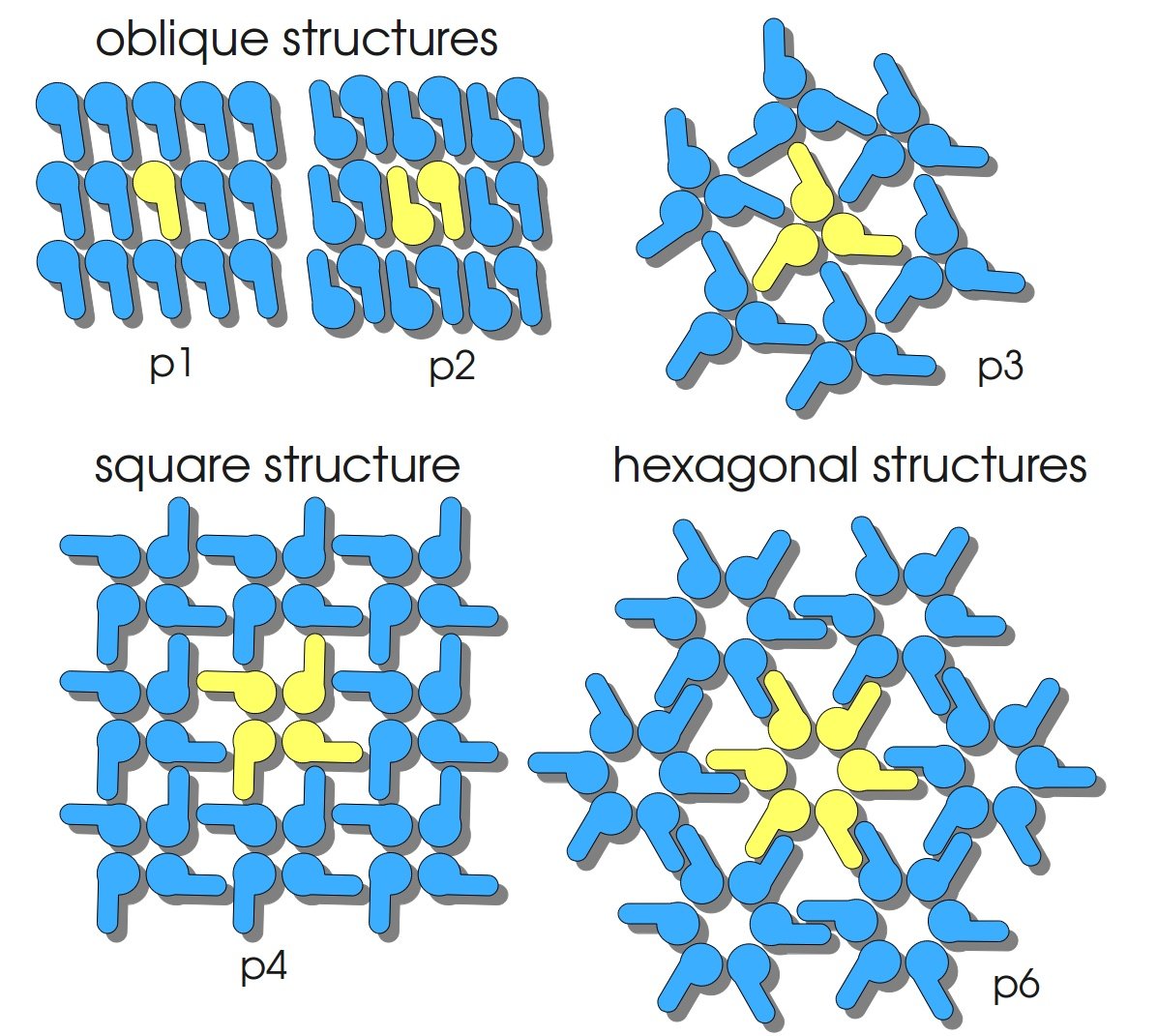
S-layer proteins have the ability to reassemble into two-dimensional crystals equal to the structures found on intact bacterial cells. Structures are periodic with very well defined distances between every protein unit. They posses porres of identical size and morphology and show equal physicochemical properties on each molecular unit. In solution, on solid supports and on phase-boundaries S-layer proteins can form self-assembly products or recrystallize into monomolecular layers. The protein subunits of S-layers are arranged in lattices with different symmetry: oblique (p1, p2), square (p4) or hexagonal (p3, p6) with a center-to-center spacing of the subunits of 3 – 35 nm (compare figure on the right). S-layers are highly porous with a porosity of 30 – 70 % (Sleytr et al., 2007).
The S-layer protein PS2 of Corynbacterium glutamicum
The S-layer protein PS2 of Corynbacterium halotolerans
The S-layer protein SgsE of Geobacillus stearothermophilus NRS 2004/3a
The S-layer protein SbpA of Lysinibacillus sphaericus CCM 2177
Summary of results
Four different S-layer BioBricks with different lattice structures were created and sent to the partsregistry. The behaviour of these genes when expressed in E. coli were characterized and purification strategies for the expressed proteins were developed. Two purified fluorescent S-layer fusion proteins from different organisms were immobilized on beads, leading to a highly significant fluorescence enhancement of these beads (p < 10-14). Furthermore regarding the other two S-layers (CspB from Corynebacterium glutamicum and Corynebacterium halotolerans) we discovered that while expression with a lipid anchor resulted in an integration into the cellmembrane, the expression with a TAT-sequence resulted in a segregation into the medium. We also detected, that those S-layers seem to stabilize the biologically active conformation of mRFP. Furthermore we expressed and purified a fluorescent CspB fusion protein from C. halotolerans which has never been expressed in E. coli until now.
 "
"