Violacein
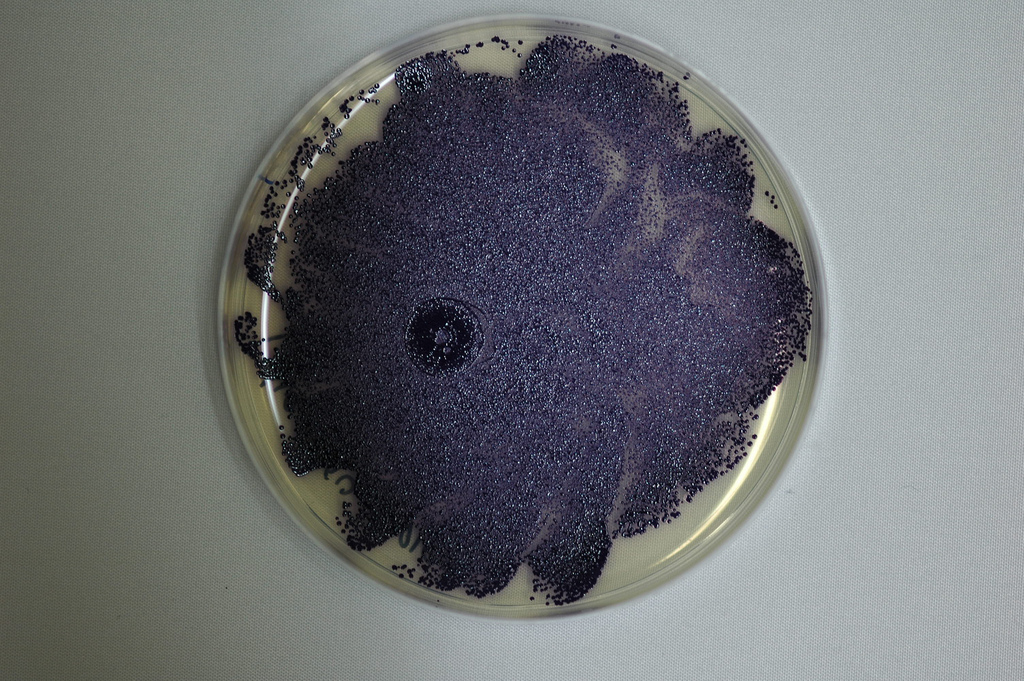
E. coli engineered by the
2009 Cambridge iGEM team to produce violacein. We are expressing the violacein pathway in yeast.
Courtesy of the
Boeke Lab, our violacein biosynthesis genes VioA, VioB, VioC, VioD and VioE have been codon-optimized for yeast and designed to ligate with yeast specific promoters and UTR terminators. To assemble full expression cassettes for each gene, the genes have been fitted with flanking restriction sites as per the
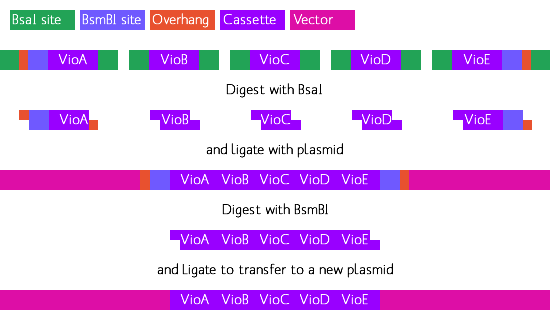
Golden Gate or "off-cutter" assembly method. This allows for
single-reaction, gapless, ordered assembly of multiple parts simultaneously.
Violacein Open Reading Frame (ORF) construct:
- XmaI site/BsaI site/Violacein ORF/inverted BsaI site/SphI site.
Image courtesy of the Boeke lab
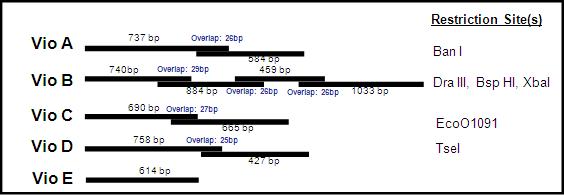
Due to the size of the above constructs, we received them in pieces, or 'building blocks'. VioA, C, D came in two building blocks. VioE was small enough to come in one piece, while VioB--around 3kb in length--came in 4 building blocks. Each building block overlaps its counterpart, and within the overlap region is a unique restriction enzyme site. By digesting both blocks with the right restriction enzyme, they can be ligated together into to make a full construct such as the one above.
The flanking XmaI and SphI sites were added for the easy insertion of the ORF into a vector double digested with XmaI and SphI. In addition to the Vio ORF's, the 12 promoters and terminators have flanking BsaI sites as above. The flanking BsaI sites on the ORF are designed for addition of the promoter and the terminator in the right spot, thus creating a viable Violacein gene expression cassette designed for yeast.
BsaI is an off-site restriction enzyme, which allows the design of a synthetic sticky overhang
of your choice. For further details, we have added a Request for Comment (RFC) dedicated to a form of BsaI gene assembly called "Golden Gate Assembly."
Violacein loxPsym Integration

The design of the violacein 'expression circles'. A promoter, violacein ORF, 3'UTR and loxPsym linker can only attach in the correct order due to the sequence of the overhangs. URA3 is added to allow for screening of synthetic DNA uptake by yeast.
We are also taking advantage of our collection of Golden Gate Assembly-compatible yeast promoters, 3'UTRs, and violacein ORFs by efficiently creating a combinatorial library which will then be integrated into the yeast genome by Cre-Lox recombination. Our promoters, ORFs, and 3'UTRs have custom overhangs such that following digestion with BsaI, they should only ligate in the correct sequence, along with a loxPsym linker. This means that we can combine our 12 promoters, 5 ORFs, and 12 3'UTRs together with the loxPsym linker in one tube to form 720 (12x5x12) possible expression casettes that have violacein ORFs paired with promoters and 3'UTRs of varying strengths. These expression cassettes will then be transformed into a yeast strain containing synIXR, a synthetic right arm of chromosome IX that contains 43 loxP sites where, in the presence of estradiol, the Cre-EBD fusion protein will catalyze random shuffling of DNA between loxP sites. The end result should be yeast with varying levels of violacein expression.
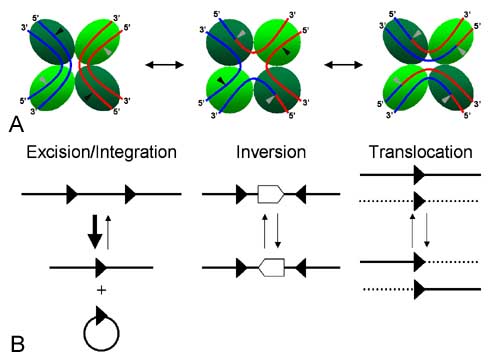
Cre-mediated loxP recombination events. Since loxPsym is symmetrical, all three types of events should take place with equal probability. Thus, some fraction of yeast transformed with the violacein 'expression circles' should integrate all five violacein genes and express the pathway
(Source)


 "
"