Team:Arizona State/Notebook/PCRLog
From 2011.igem.org
|
|
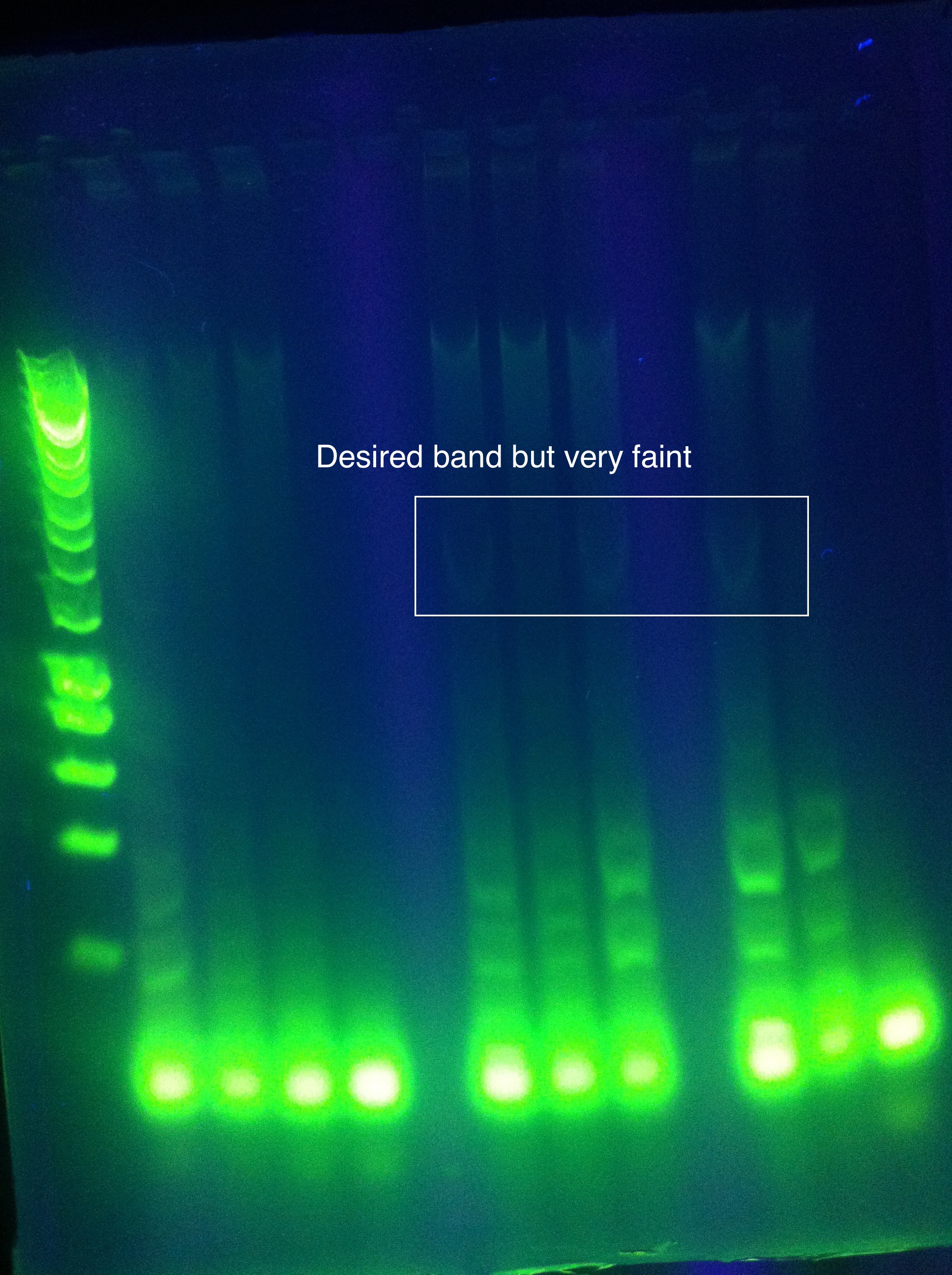
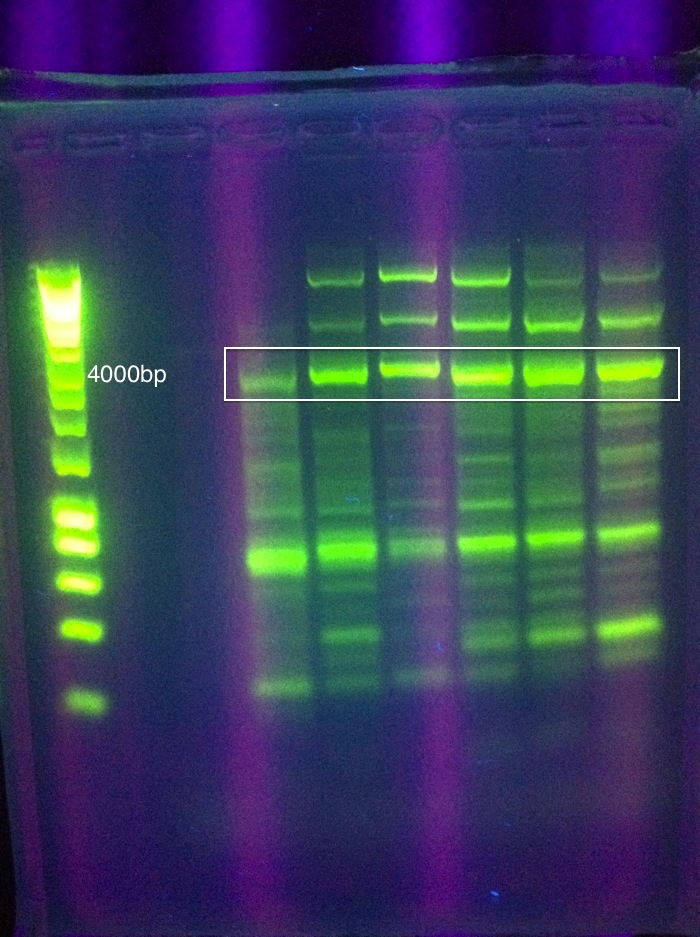
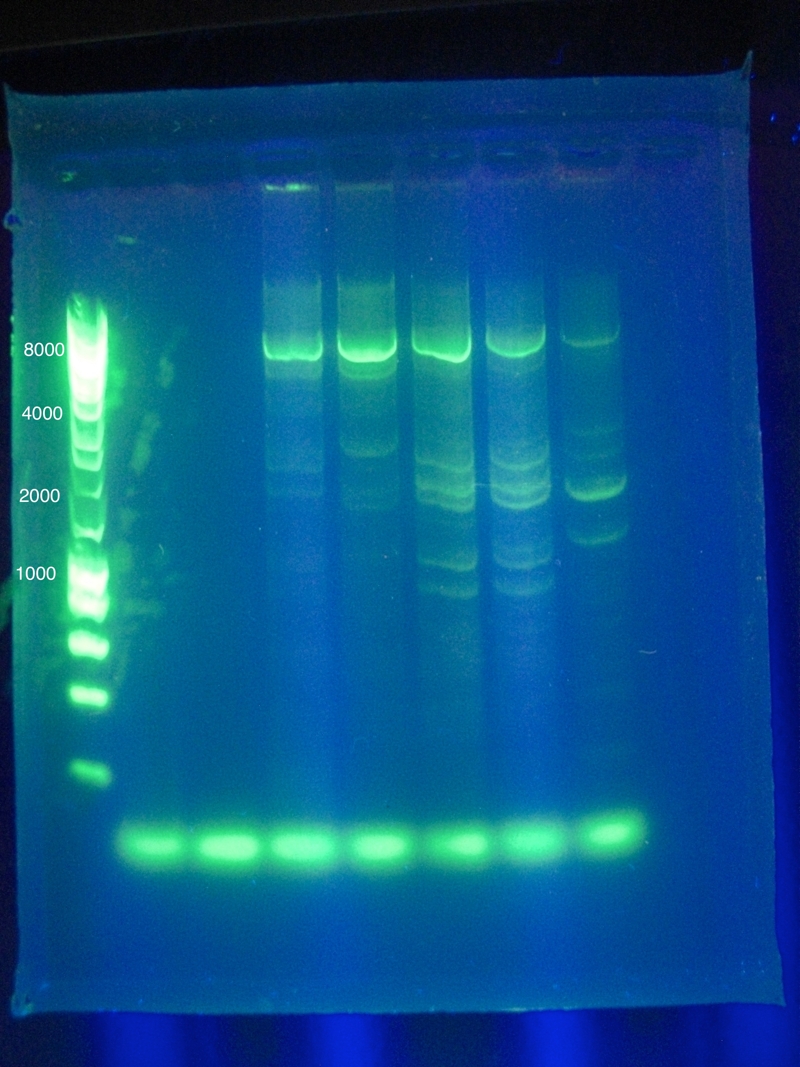
OverviewThis logbook is a record of the majority of our attempts to PCR amplify CRISPR-associated (Cas) ABCDE and Cas 3 of E. coli K12 MG1655. The record begins after our attempt to individually PCR amplify each of the 6 genes, which was unsuccessful. Here, three different sets of primers were used to attempt PCR amplification of the Cas genes in two sections. NotesSuggested annealing temperatures are based on NEB Tm Calculator, as called for in the NEB Phusion DNA Polymerase protocol. Desired band for CasABCDE at ~4300bp Desired band for Cas3 at ~2667bp Primer Round 1July 17, 2011CasABCDE: Touchdown PCR, Start Temp 70, -0.2 / cycle, Final Temp 64
Gel Results:
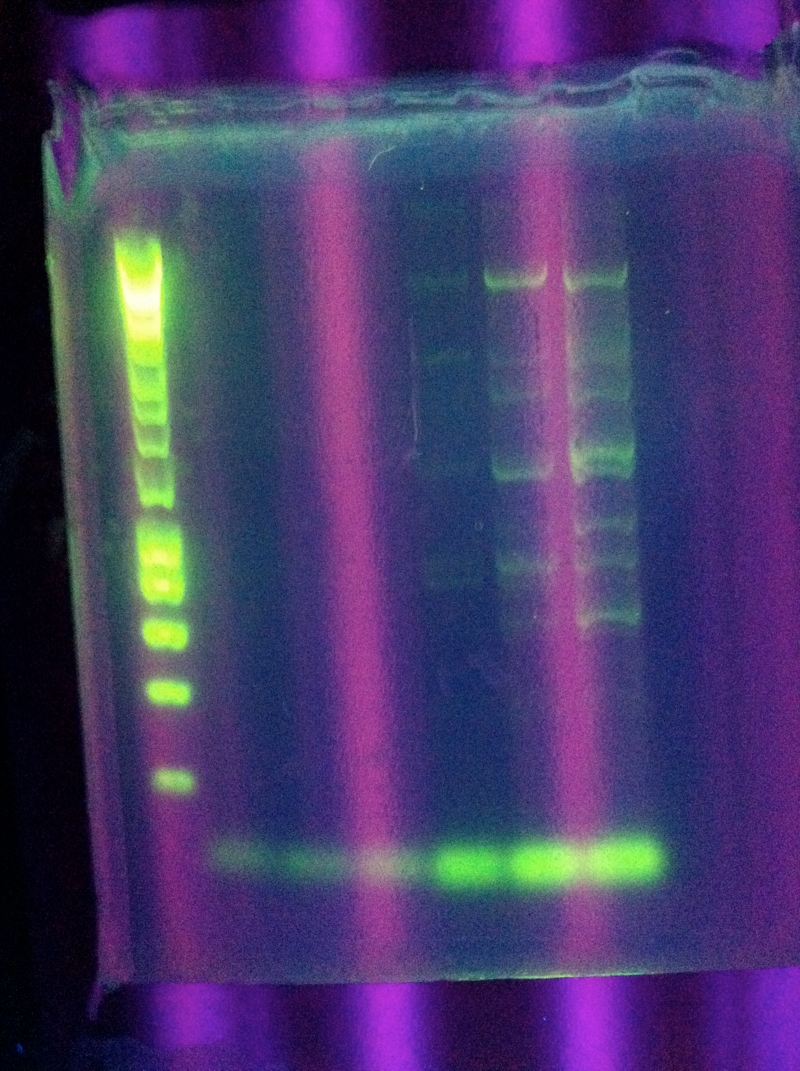
July 18, 2011CasABCDE: Touchdown PCR, Start Temp 65, -0.2 / cycle, Final Temp 59 Gel Results: CasABCDE: No bands July 19, 2011CasABCDE: Touchdown PCR, Start Temp 70, -0.2 / cycle, Final Temp 63
July 20, 2011CasABCDE: Touchdown PCR, Start Temp 71, -0.2 / cycle, Final Temp 64
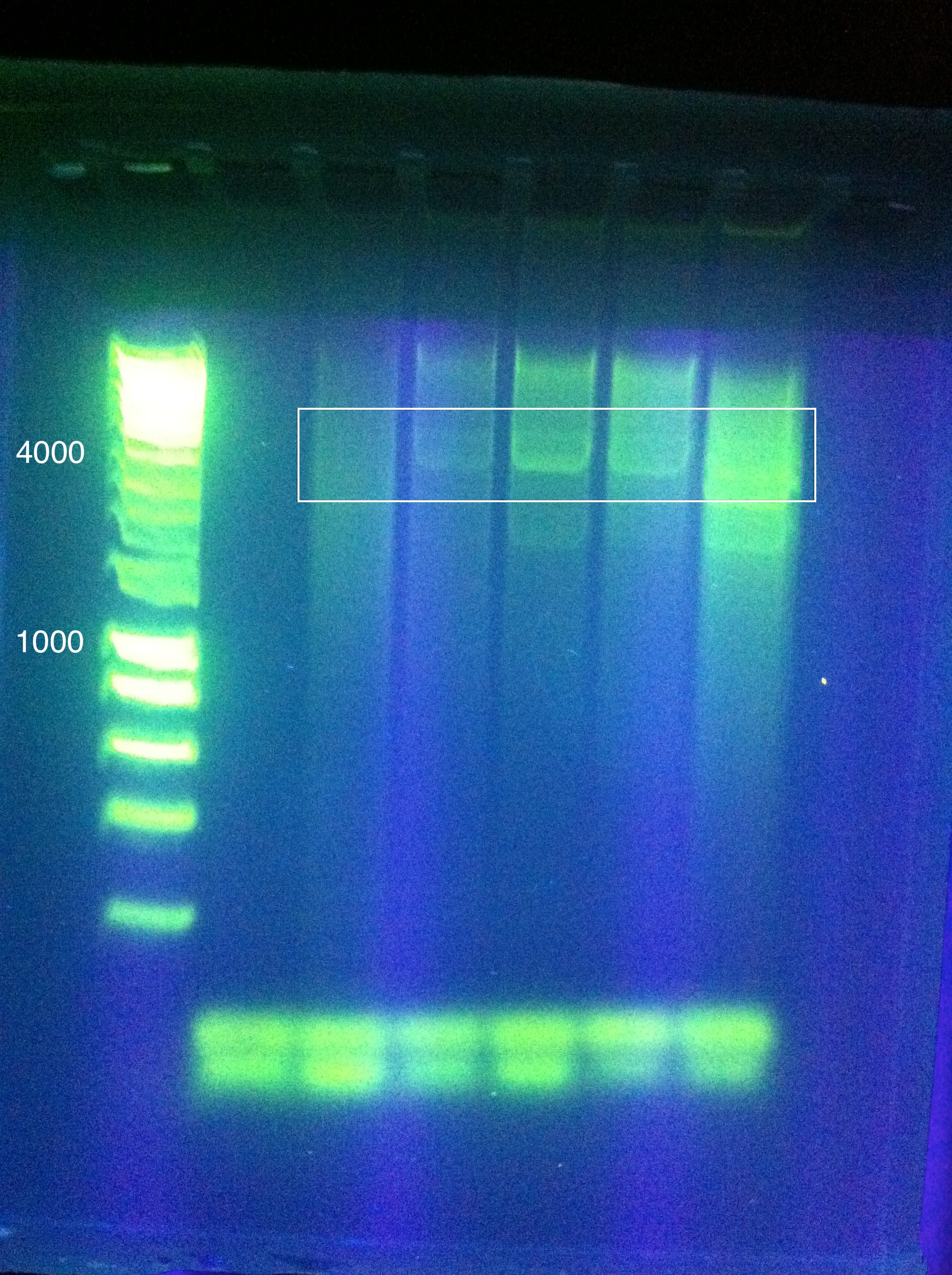
Gel Results:
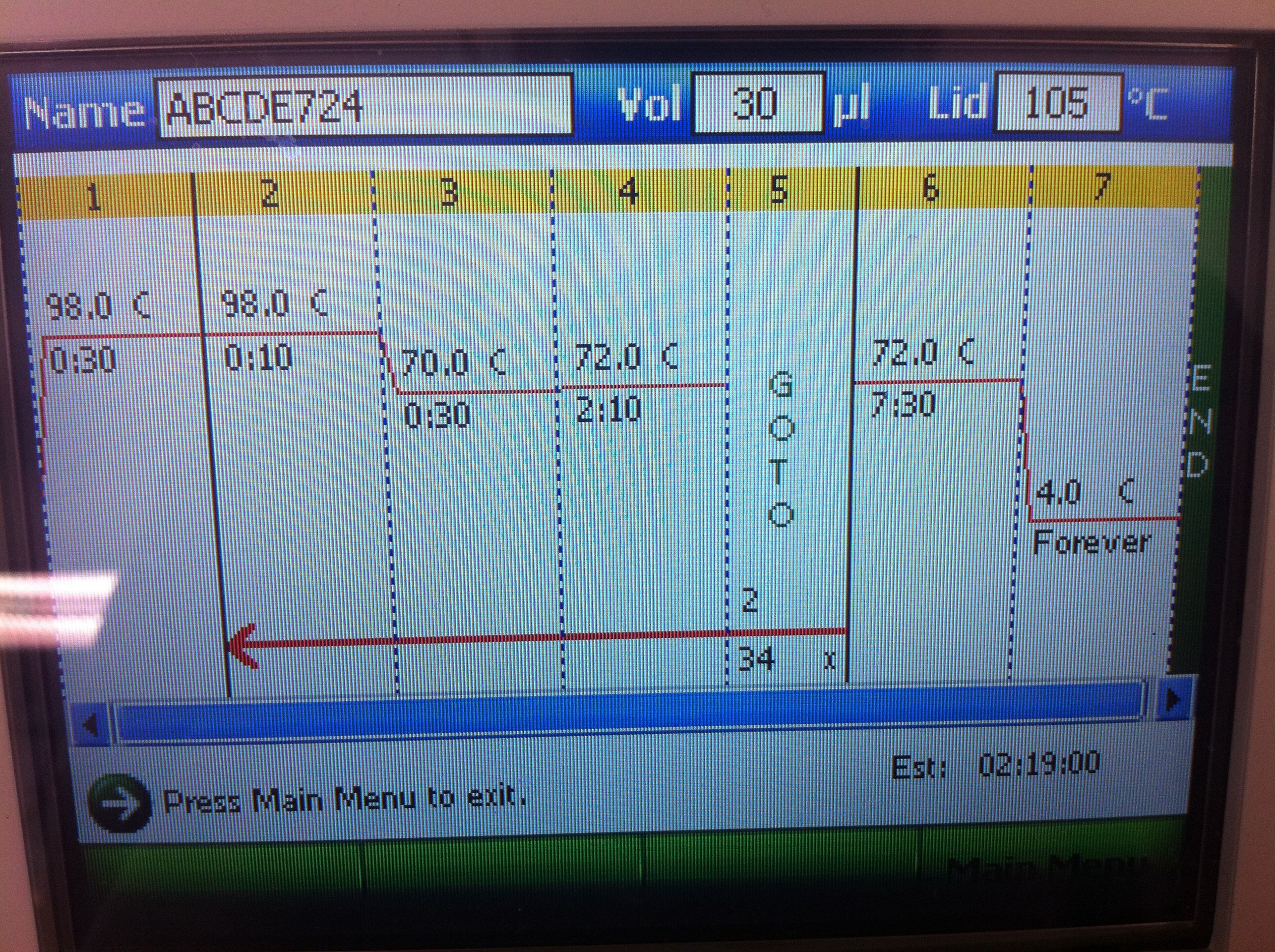
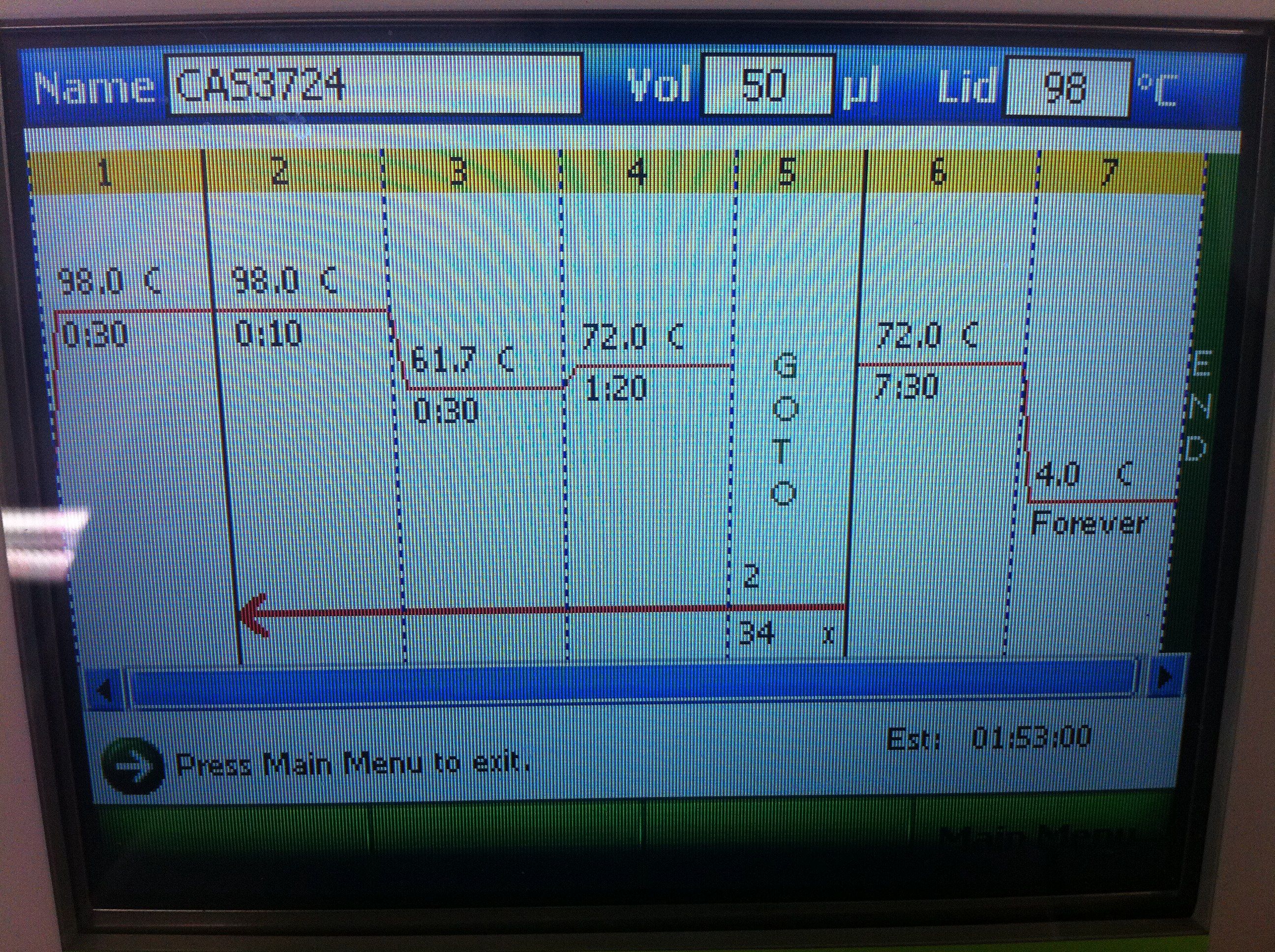
July 24, 2011PCR of CasABCDE, Cas3 with new settings; Preparation for extraction, "PCRception" (PCRing the PCR results); Included an elongation step in each cycle, which was not done for the previous runs; Settings stored in PCR machine as ABCDE724, CAS3724 CasABCDE Settings and Results:
Primer Round 2July 26, 2011CasABCDE: 68 degree annealing temp, 2 minute elongation
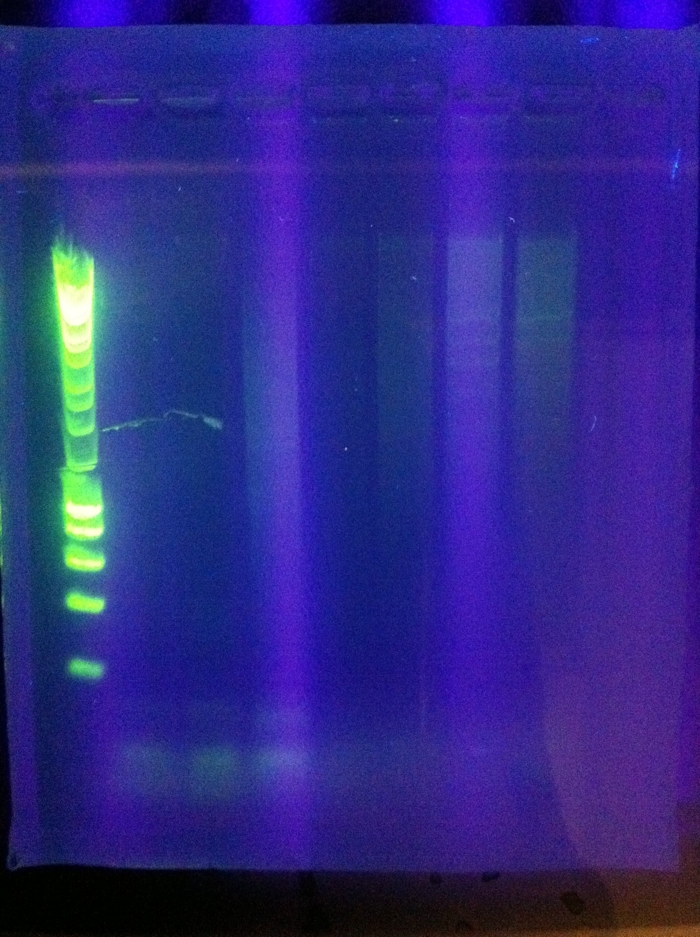
Gel Results: CasABCDE: mostly blank
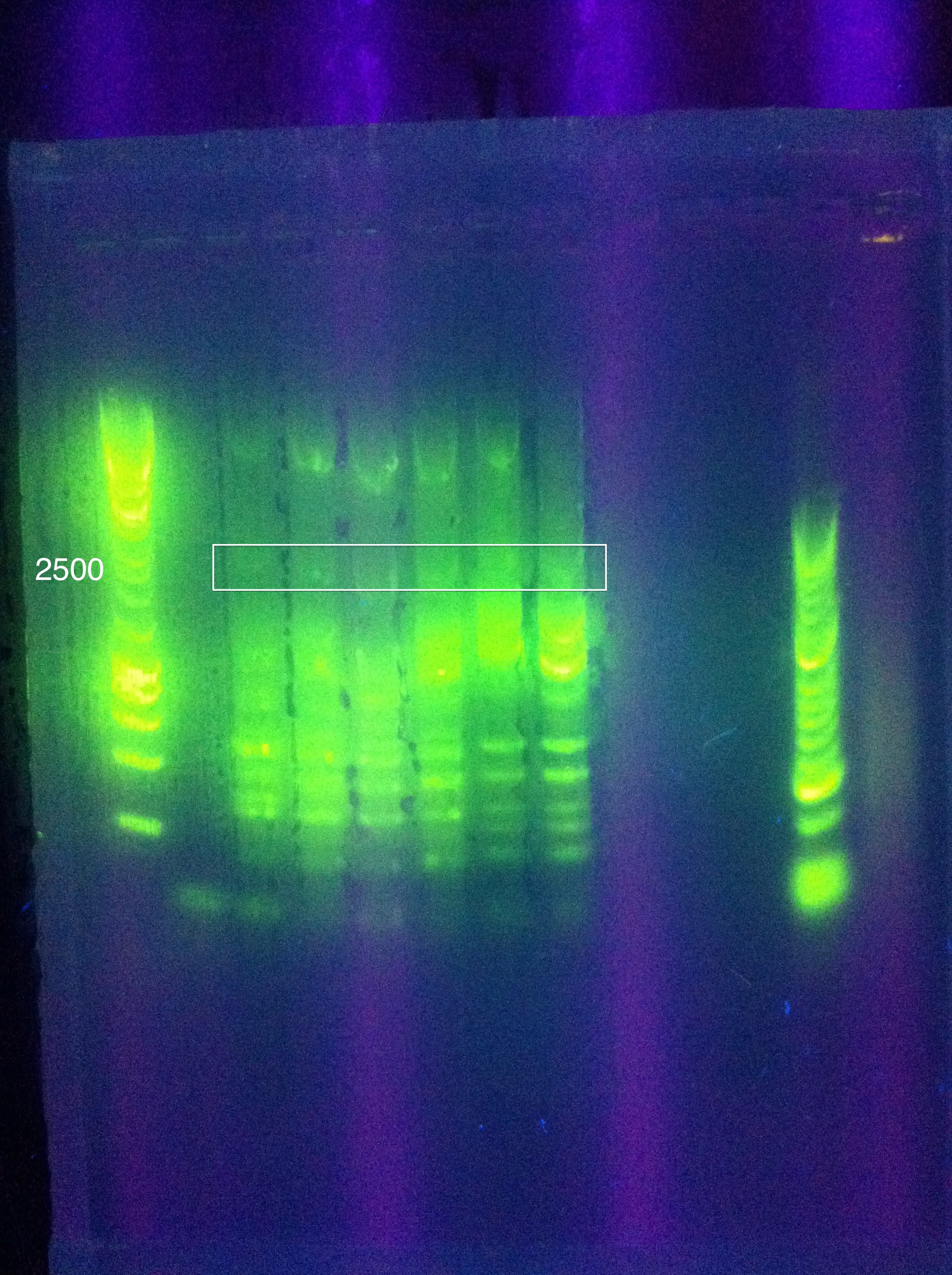
July 27, 2011CasABCDE (new): 64->69, three rows of 3 tubes
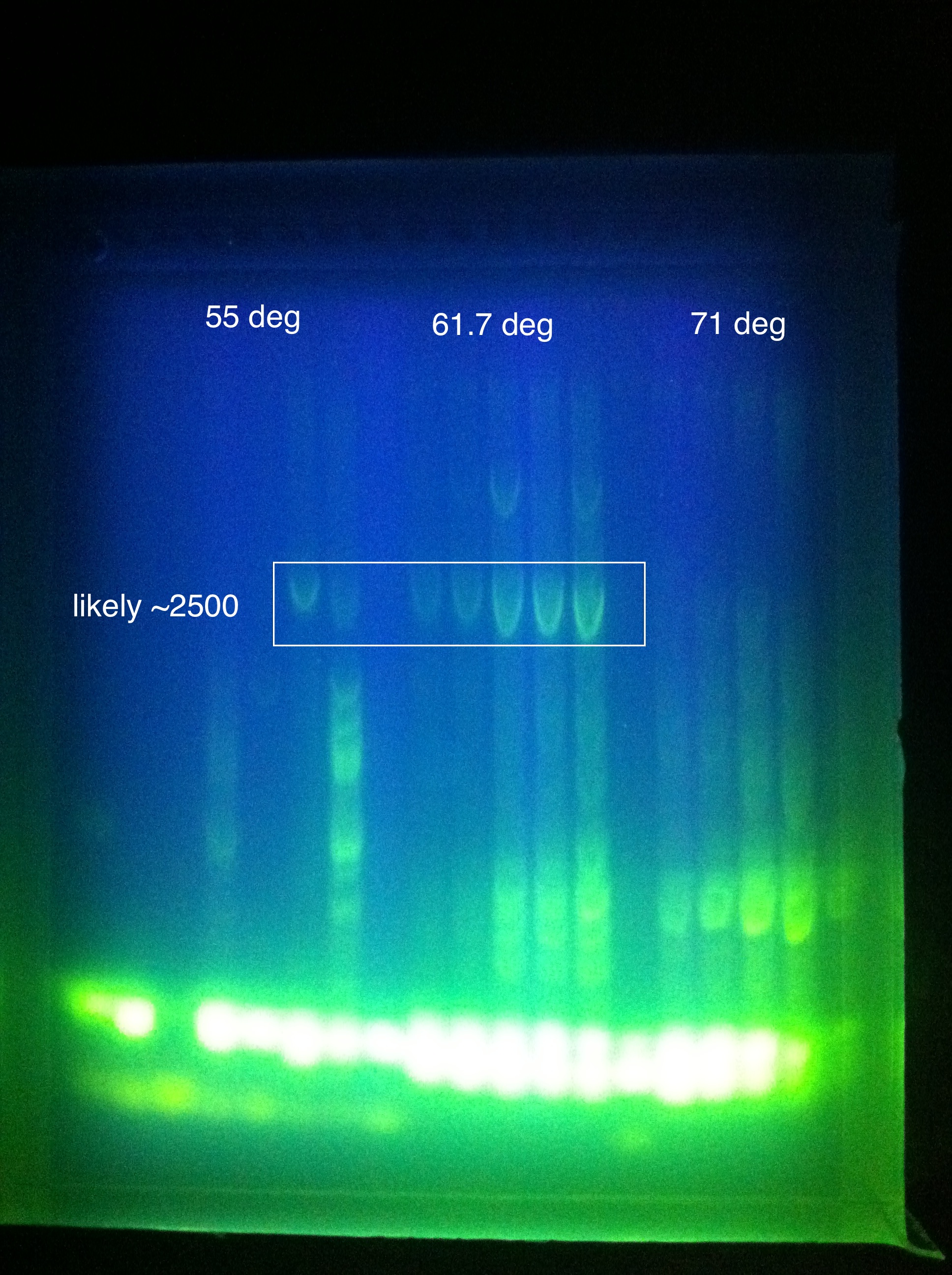
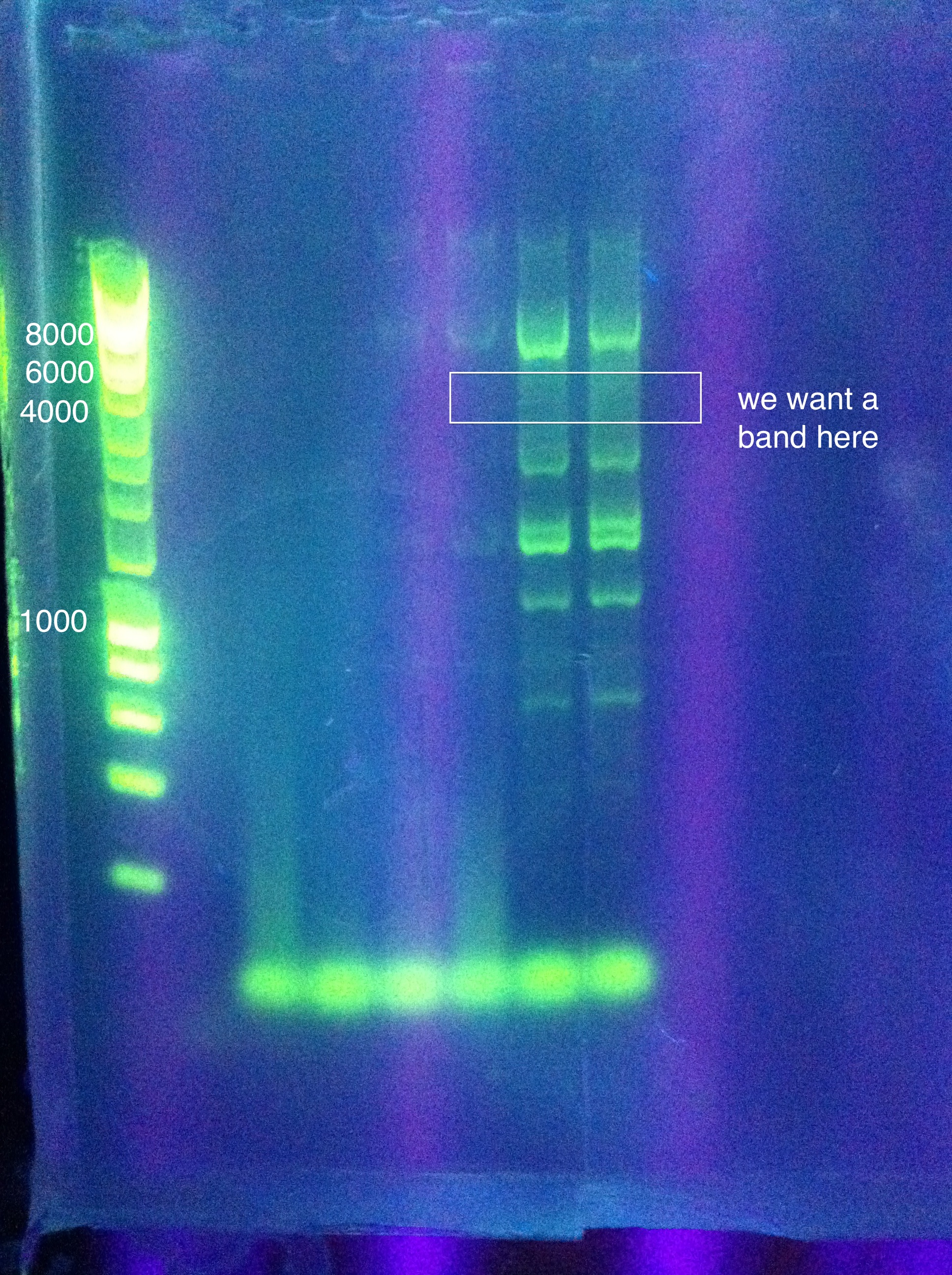
Gel Results:
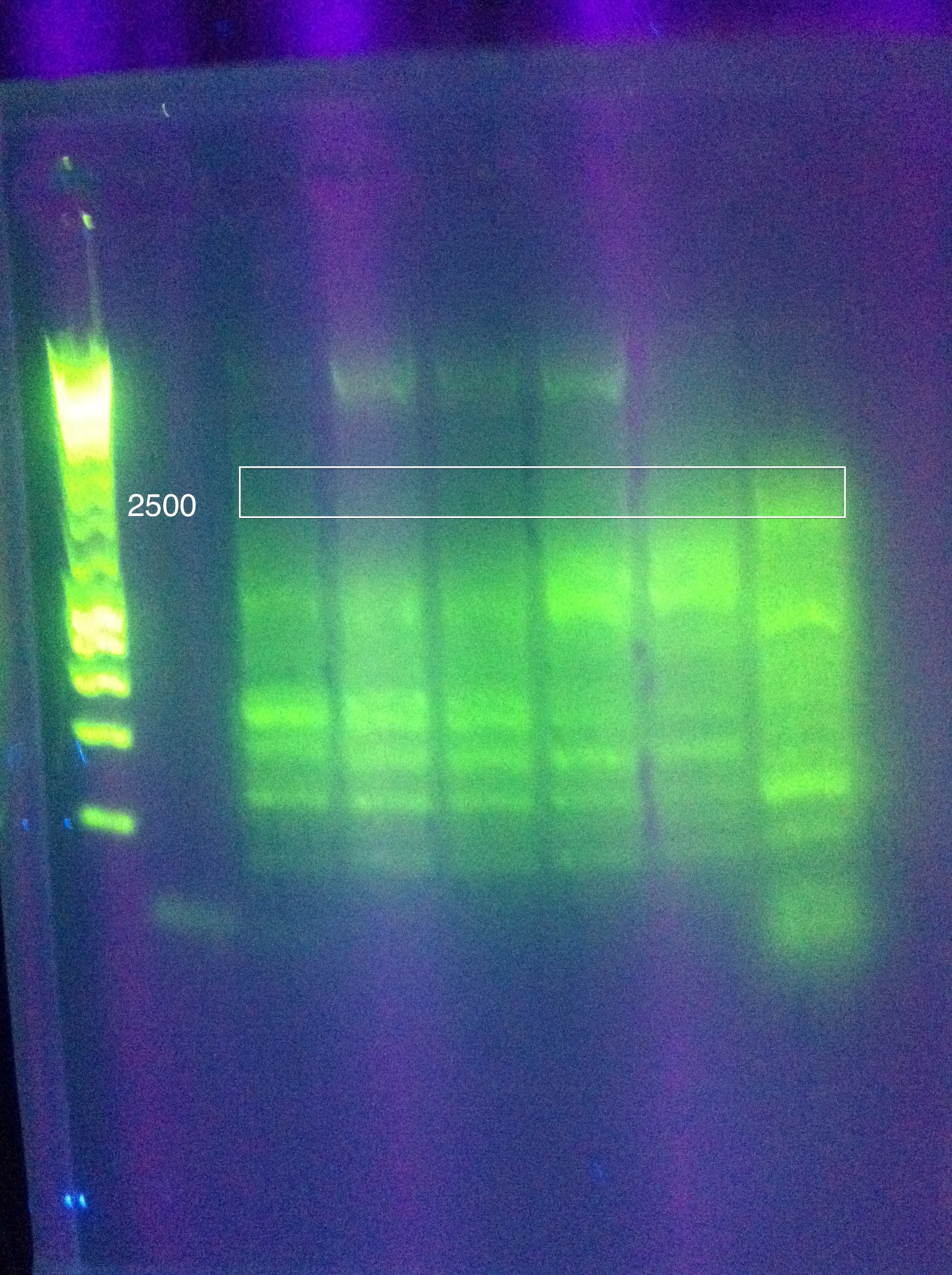
July 28, 2011CasABCDE (old): TDWN728B, 71 to 68, -.1/cycle, 30 cycles, 2:10 elongation
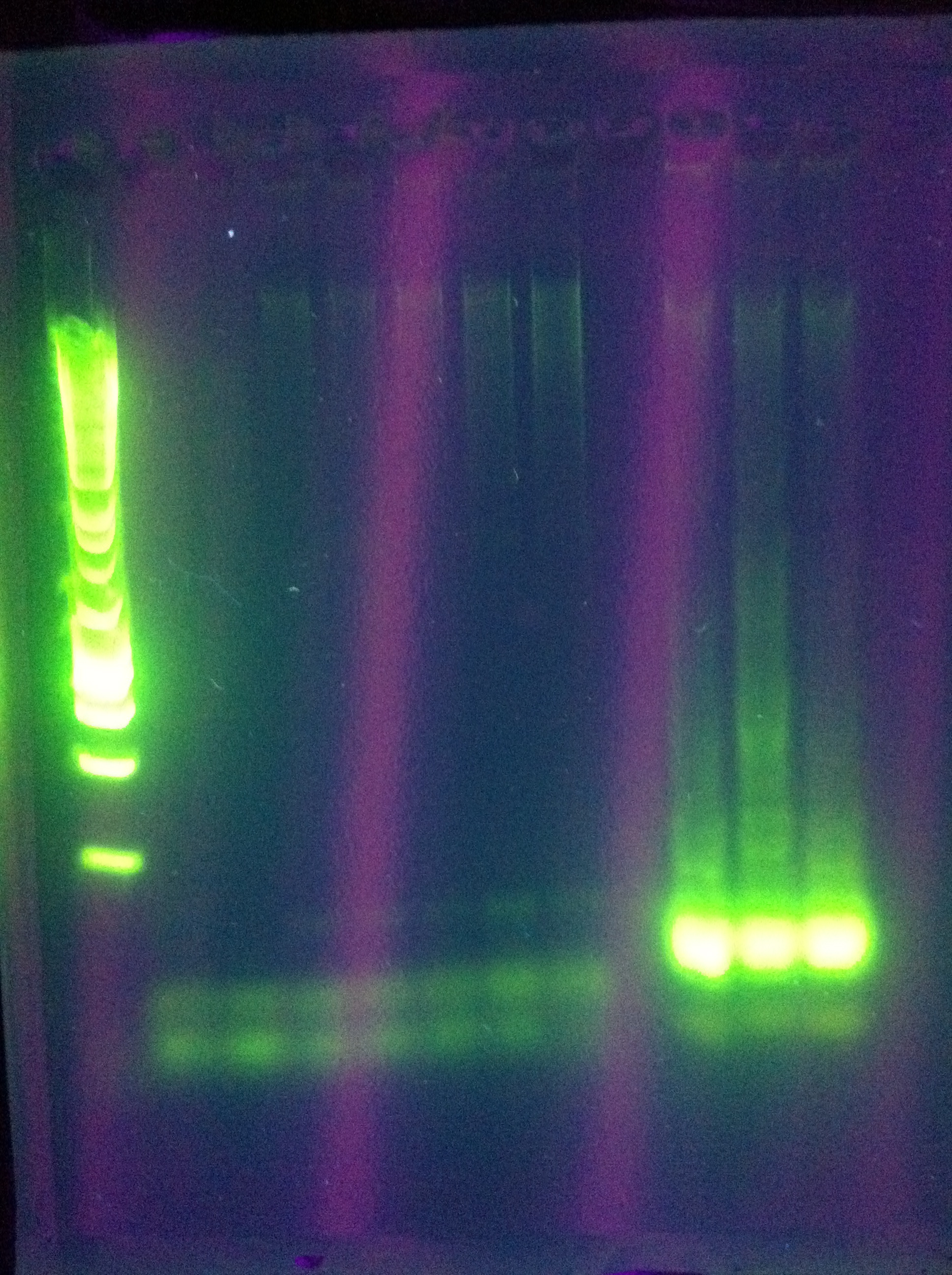
Gel Results:
July 29, 2011CasABCDE: 68 Touchdown from 72 to 65, -.2/cycle, 35 cycles, 2:10 elongation Gel Results: August 9, 2011Cas3: Ran Cas3724 protocol Gel Results: Primer Round 3 (CasABCDE only)August 4, 2011Settings: 98 initial denaturation for 30 seconds, Cycle (10sec at 98deg, anneal 30 sec at 63 deg, elongate 130sec at 72deg), 35 Cycles, Extension for 450 seconds
August 8, 2011Settings:ABCDE808, which is same as 8/4 run but only 29 cycles in the hope that the bright band from the previous try would be the only one visible.
August 8, 2011Settings: Retry same protocol in thermocycler, however increase back to 35 cycles.
August 10, 2011 (Evening)Settings: Ran another PCR for ABCDE, same settings as before but one degree higher for annealing temp. Also added DMSO.
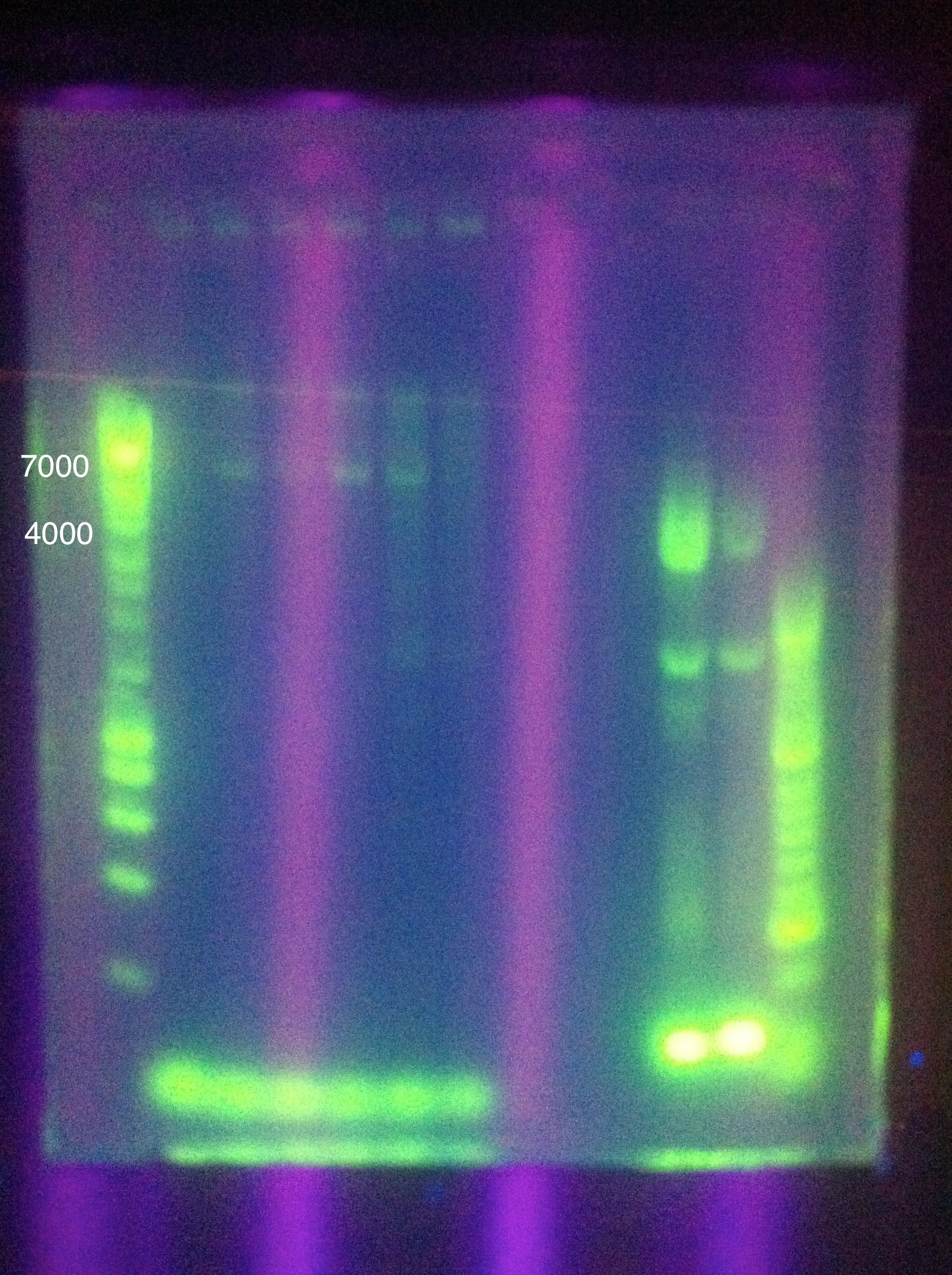
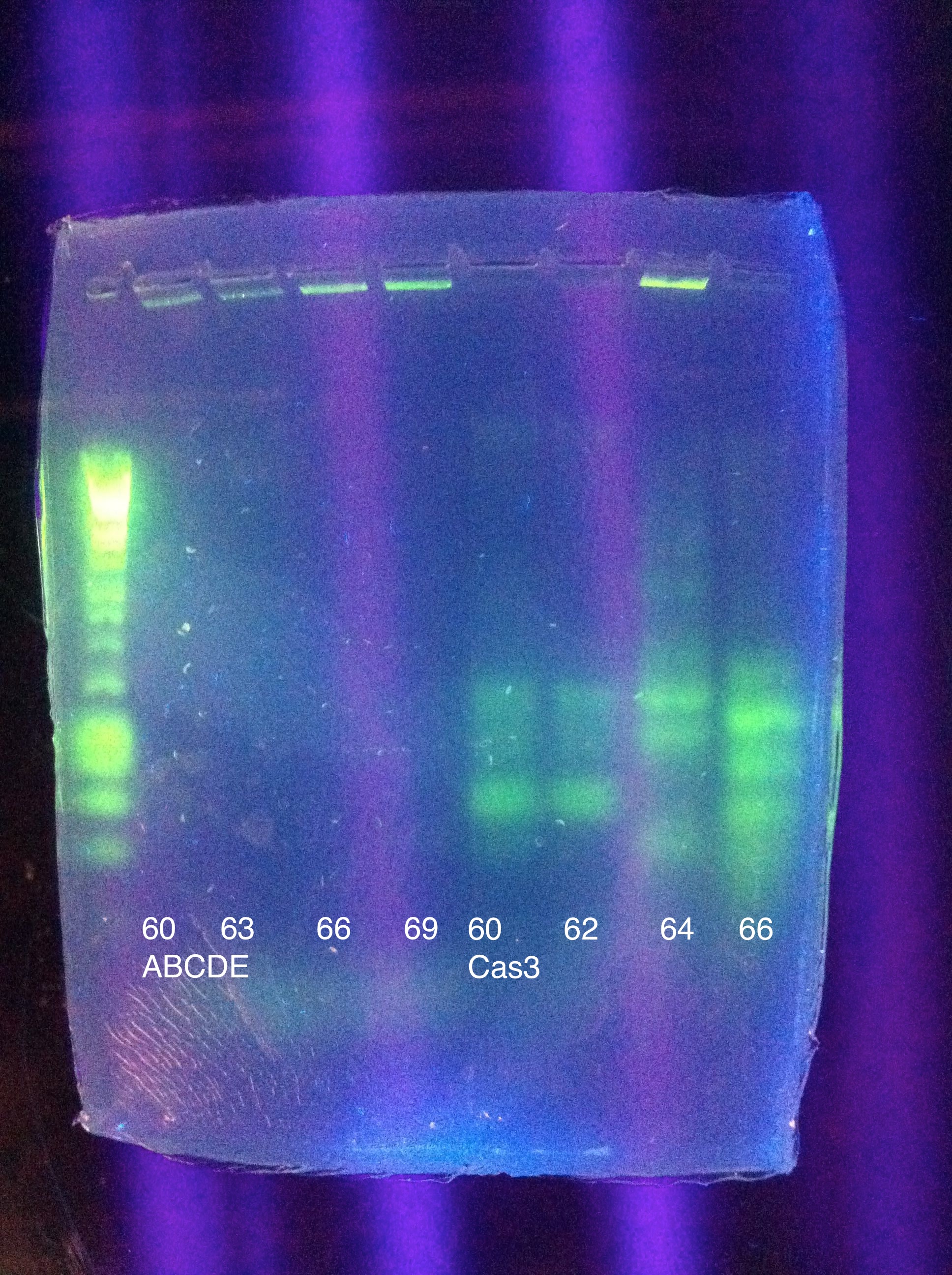
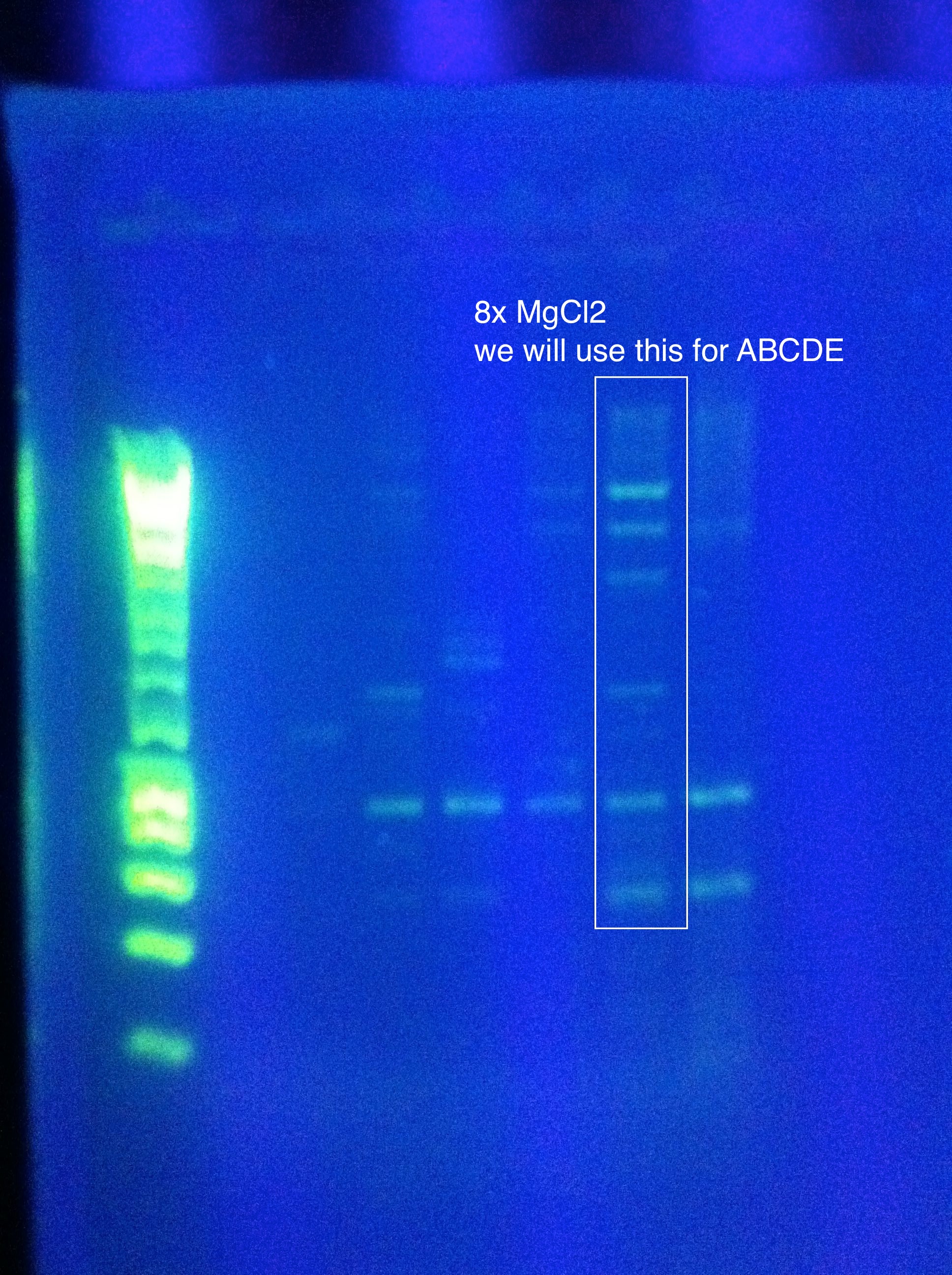
August 11, 2011Temperature Gradient PCR CasABCDE: MgCl2 --> 2uL, 4uL, 5uL, Temp --> 60, 63, 66, 69
Moving forward to nested PCR for ABCDE, we will see how this goes. Cas3 we'll have to keep trying…perhaps it may be worth it to try DMSO, longer elongation time, something. Maybe Taq. August 12, 2011
Use 1uL per tube (note, we don't know the concentration of the strand we want as template, so this is our best guess as to what would get us between 1pg and 10ng as recommended by the Phusion protocol for non-genomic DNA) |
 "
"