Team:Washington/Magnetosomes/Magnet Toolkit
From 2011.igem.org
(→Construction of the Magnetosome Genome (in parts) in E.coli) |
(→What are magnetosomes? Where do they come from?) |
||
| (37 intermediate revisions not shown) | |||
| Line 2: | Line 2: | ||
__NOTOC__ | __NOTOC__ | ||
| - | + | <center><big><big><big><big>'''iGEM Toolkits: Magnetosomes'''</big></big></big></big></center><br><br> | |
===What are magnetosomes? Where do they come from?=== | ===What are magnetosomes? Where do they come from?=== | ||
| + | [[File:Washington iGEM2011 magnetotatic bacteria picture.jpg|thumb|right|350px|Magnetotactic Bacteria (left) and Magnetosome chains (right)]] | ||
| + | <br> Magnetotactic bacteria are prokaryotic organisms that possess the unique ability to align themselves along a magnetic field. This form of taxis is made possible by the formation of a magnetosome. Magnetosomes are small invaginations of the bacterial inner membrane that contain magnetite particles. | ||
| - | [ | + | These particles range in size between 20 and several hundred nanometers and are aligned in one or several chains along the long axis of the bacteria. These particles act together to form a magnetic dipole across the bacteria, allowing it to sense the earth’s magnetic field. Magnetotactic bacteria are microaerophilic; therefore, magnetosomes are thought to help aid the organism in its search for the optimal oxygen level from a search in three dimensional space (in all directions) to a one dimensional space along a single path. |
| + | <br><br><br> | ||
| + | <center>Video demonstration of magnetic property of AMB-1 ([http://youtu.be/evrZEe_q4V4 direct link]):</center> | ||
| + | <html><center><iframe width="420" height="315" src="http://www.youtube.com/embed/evrZEe_q4V4" frameborder="0" allowfullscreen></iframe></center></html> | ||
| + | <br> <br> | ||
| + | <i>Narration text from video:</i> Magnetotactic bacteria are named for their ability to respond to and move along magnetic fields. They were first discovered in 1975 by Richard Blakemore when he noticed bacteria collecting on the north most edge of a water droplet he had placed on a microscope slide. Magnetotactic bacteria use a chain of vesicle-bound magnetite particles (known as magnetosomes) as a biological compass to orient themselves along the earth's magnetic field lines. They then swim along these field lines with flagella. It is thought that this process evolved to allow magnetotactic bacteria to search for certain micro-environments more efficiently. Magnetotactic bacteria in the northern hemisphere usually swim northward while those found in the southern hemisphere swim southward. In both cases this would direct the bacterium downward and is thought to allow it to find bottom sediments in an aqueous environment. As shown here, when a strong magnetic field is applied to the bacteria they can be moved as the magnetite particles within them become polarized and attracted to the magnetic field source. | ||
| - | + | ===A Closer look at Magnetosome Formation === | |
| - | + | [[File:F6.medium.png|300px|thumb|right|Diagram of stepwise magnetosome construction within AMB-1]] | |
| + | <br> | ||
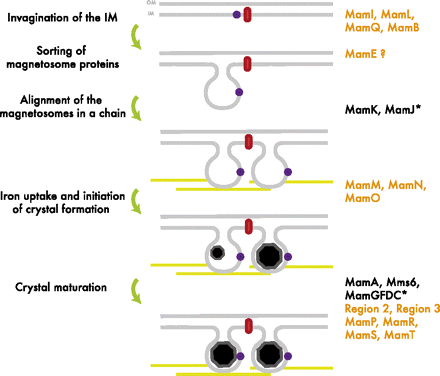
| + | The formation of the magnetosome organelle is a highly regulated, step-wise process requiring a cascade of essential genes. The process is generally hypothesized as four stages: | ||
| + | # Membrane invagination | ||
| + | # Acquiring minerals for magnetite formation | ||
| + | # Iron-oxidation and reduction | ||
| + | # Magnetite nucleation and morphology regulation. | ||
| + | <br> Earlier gene products must be present for later gene products to be formed as shown in the diagram on the right[http://www.pnas.org/content/107/12/5593.full.pdf+html]: | ||
| - | + | <br><br><br> | |
| - | + | ===What did the UW iGEM team do with Magnetotactic Bacteria?=== | |
| - | + | <br> | |
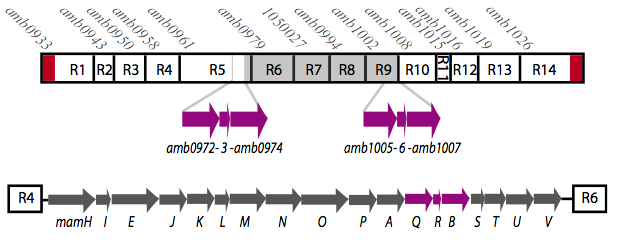
| + | It is thought that many of the essential genes associated with magnetosome formation are located within a well-conserved region known as the magnetosome island (MAI). The MAI consists of 14 gene clusters labeled R1-R14 (see diagram below).Our team focused on the genes of the mamAB gene cluster (R5), as they were previously shown to be the only cluster essential for magnetosome membrane biogenesis in AMB-1 (diagram show below).[http://www.pnas.org/content/107/12/5593/F1.expansion.html]. | ||
| - | + | The goal of our project was to extract all the essential genes from (R5) required for magnetosome formation and express them in ''E. coli''. We are doing this to learn more about magnetosome formation and the magnet synthesis mechanism, because many of the genes' functions are still unknown in the host species. Using the information we have gained, we have organized a '''Magnetosome Toolkit''' containing most of the essential genes for proper magnetosome formation. Ultimately, we would like to continue expanding the magnetosome toolkit to have enough parts to show complete magnetosome formation in ''E.coli''. | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | The goal of our project was to extract all the essential genes from (R5) required for magnetosome formation and express them in E.coli. | + | |
[[File:MamAB.png|center|500px|thumb|Fig. 3: The mamAB operon (R5) located in the magnetosome island (MAI).]] | [[File:MamAB.png|center|500px|thumb|Fig. 3: The mamAB operon (R5) located in the magnetosome island (MAI).]] | ||
| + | <br> | ||
| + | ==='''About the Magnetosome Toolkit'''=== | ||
| + | <br> | ||
| + | Using standard synthetic biology protocols and the vectors we created in our Gibson Assembly Toolkit, our team created the '''"Magnetosome Toolkit"''' which contains many of the genes required for magnetosome formation. Providing this toolkit to the Parts Registry will help allow future iGEM teams to manipulate and further understand magnetosome formation to eventually synthesize magnets in multiple organisms. | ||
| + | <br> <br/> | ||
| + | As previously noted, magnetosome formation within the host-organism, ''Magnetospirillium magneticum'', strain AMB-1, is a highly regulated step-wise process. As shown in diagram of stepwise magnetosome construction above, some genes encode proteins that form an invagination of the inner membrane, other genes which help align the magnetosomes into their characteristics chains, and others which regulate the biomineralization of magnetic particles. Our team chose to focus on genes specifically related to magnetosome scaffolding/alignment since they are the essential foundation for magnetosome development. In addition, the creation of a scaffold to which other genes localize is highly applicable to systems in synthetic biology. (for more information, please see our [https://2011.igem.org/Team:Washington/Magnetosomes/Future Future Directions] page) | ||
| + | <br> | ||
| - | + | The genes we focused on are <i>mamK</i> and <i>mamI</i> since they have known functions related to localization of the magnetosome. Specifically, MamK is a bacterial actin-like cytoskeleton protein required for proper alignment of the magnetosomes in a chain. MamK is also shown to localize the MamI, which when lost inhibits vesicle formation. | |
| - | + | (for other gene functions, please see the iGEM Toolkits parts submitted page) | |
| - | + | <br> <br> | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | (for other gene functions, see the | + | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
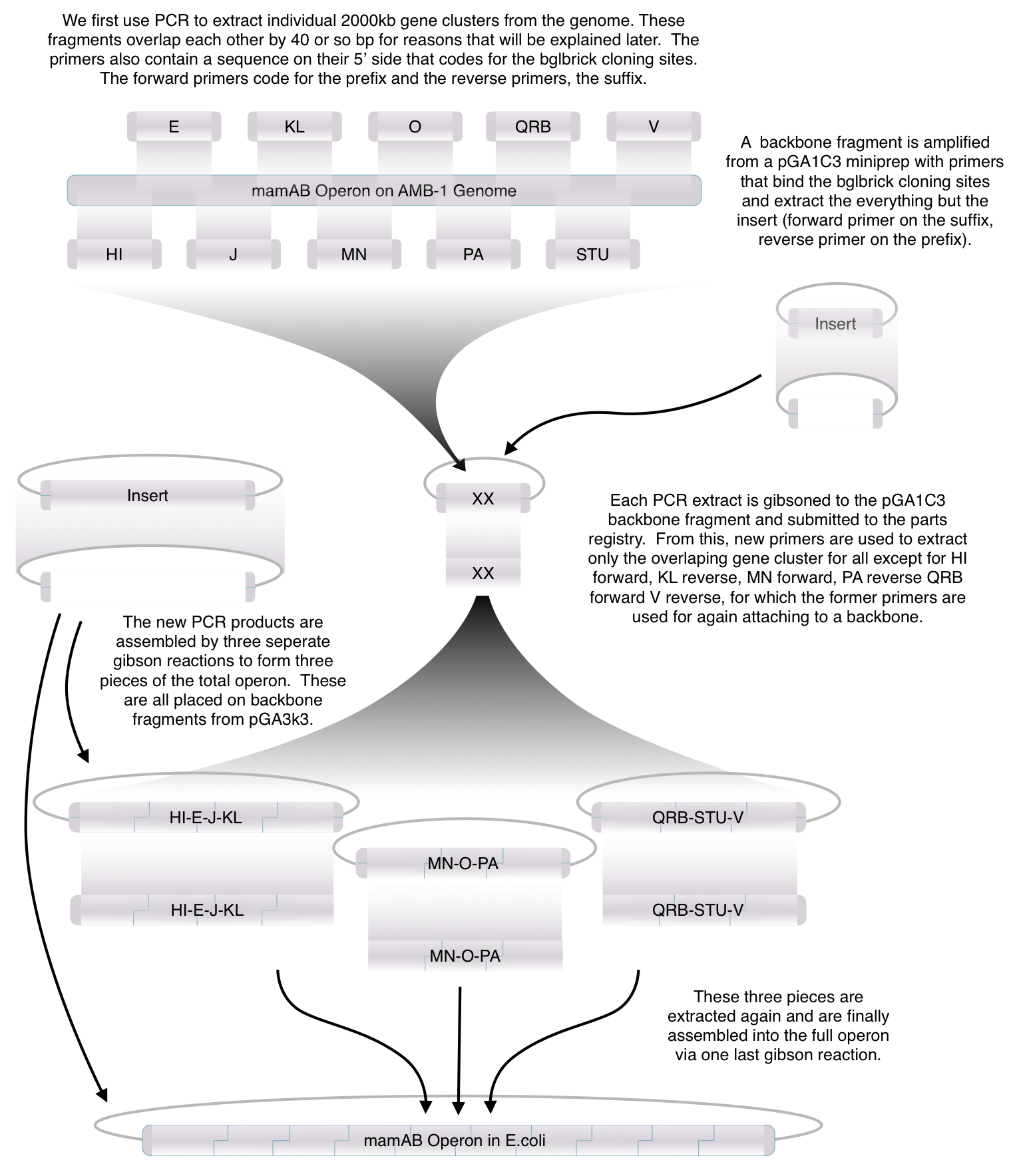
| + | ==='''Toolkit Contruction'''=== | ||
| + | [[File:Washington Methode image.jpg|700px|center]] | ||
| - | [ | + | <br/>Please see our [https://2011.igem.org/Team:Washington/Magnetosomes/Magnet_Results Result Summary] page for to see how far we were able to get this summer! |
| - | + | <br/> | |
| - | = | + | =References:= |
| - | |||
| - | ----- | + | # Matsunaga, T., Okamura, Y., Fukuda, Y., Wahyudi, A.T., Murase, Y., Takeyama, H. (2005). Complete genome sequence of the facultative anaerobic Magnetotactic bacterium Magnetospirillum sp. strain AMB-1. ''DNA research''; 12: 157-166. Doi:10.1093/dnares/dsi002. |
| + | # Murat, D., Quinlan, A., Vali, H., Komeili, A. (2010). Comprehensive genetic dissection of the magnetosome gene island reveals the step-wise assembly of a prokaryotic organelle. ''PNAS''; 107 (12): 5593-5598. Doi:10.1073/pnas.0914439107. | ||
| + | # Murat, D., Quinlan, A., Vali, H., Komeili, A. (2010). Supporting Information. ''PNAS''; 107 (12): 5593-5598. Doi:10.1073/pnas.0914439107. | ||
| + | # Quinlan, A., Murat, D., Vali, H., Komeili, A. (2011).The HtrA/DegP family protease MamE is a bifunctional protein with roles in magnetosome protein localization and magnetite biomineralization. ''Molecular Microbiology''; 80 (4): 855-1131. Doi:10.1111/j.1365-2958.2011.07631.x. | ||
| + | # Richter, M., Kube, M., Bazylinski, D.A., Lombardot, T.,Glockner, F.O., Reinhardt, R., Shuler, D. (2007). Comparative genome analysis of four Magnetotactic bacteria reveals a complex set of group-specific genes implicated in magnetosome biomineralization and function. ''Journal of Bacteriology''; 189(13): 4899-4910. Doi:10.1128/JB.00119-07. | ||
| + | # Rioux, J.B., Philippe, N., Pereia, S., Pignol, D., Wu, L.F., Ginet, N. (2010). A second actin-like mamK protein in Magnetospirillum magneticum AMB-1 encoded outside the genomic magnetosome island. ''PLoS ONE''; 5(2): e9151. Doi:10.1371/journal.pone.0009151. | ||
Latest revision as of 18:45, 27 October 2011
What are magnetosomes? Where do they come from?
Magnetotactic bacteria are prokaryotic organisms that possess the unique ability to align themselves along a magnetic field. This form of taxis is made possible by the formation of a magnetosome. Magnetosomes are small invaginations of the bacterial inner membrane that contain magnetite particles.
These particles range in size between 20 and several hundred nanometers and are aligned in one or several chains along the long axis of the bacteria. These particles act together to form a magnetic dipole across the bacteria, allowing it to sense the earth’s magnetic field. Magnetotactic bacteria are microaerophilic; therefore, magnetosomes are thought to help aid the organism in its search for the optimal oxygen level from a search in three dimensional space (in all directions) to a one dimensional space along a single path.
Narration text from video: Magnetotactic bacteria are named for their ability to respond to and move along magnetic fields. They were first discovered in 1975 by Richard Blakemore when he noticed bacteria collecting on the north most edge of a water droplet he had placed on a microscope slide. Magnetotactic bacteria use a chain of vesicle-bound magnetite particles (known as magnetosomes) as a biological compass to orient themselves along the earth's magnetic field lines. They then swim along these field lines with flagella. It is thought that this process evolved to allow magnetotactic bacteria to search for certain micro-environments more efficiently. Magnetotactic bacteria in the northern hemisphere usually swim northward while those found in the southern hemisphere swim southward. In both cases this would direct the bacterium downward and is thought to allow it to find bottom sediments in an aqueous environment. As shown here, when a strong magnetic field is applied to the bacteria they can be moved as the magnetite particles within them become polarized and attracted to the magnetic field source.
A Closer look at Magnetosome Formation
The formation of the magnetosome organelle is a highly regulated, step-wise process requiring a cascade of essential genes. The process is generally hypothesized as four stages:
- Membrane invagination
- Acquiring minerals for magnetite formation
- Iron-oxidation and reduction
- Magnetite nucleation and morphology regulation.
Earlier gene products must be present for later gene products to be formed as shown in the diagram on the right[http://www.pnas.org/content/107/12/5593.full.pdf+html]:
What did the UW iGEM team do with Magnetotactic Bacteria?
It is thought that many of the essential genes associated with magnetosome formation are located within a well-conserved region known as the magnetosome island (MAI). The MAI consists of 14 gene clusters labeled R1-R14 (see diagram below).Our team focused on the genes of the mamAB gene cluster (R5), as they were previously shown to be the only cluster essential for magnetosome membrane biogenesis in AMB-1 (diagram show below).[http://www.pnas.org/content/107/12/5593/F1.expansion.html].
The goal of our project was to extract all the essential genes from (R5) required for magnetosome formation and express them in E. coli. We are doing this to learn more about magnetosome formation and the magnet synthesis mechanism, because many of the genes' functions are still unknown in the host species. Using the information we have gained, we have organized a Magnetosome Toolkit containing most of the essential genes for proper magnetosome formation. Ultimately, we would like to continue expanding the magnetosome toolkit to have enough parts to show complete magnetosome formation in E.coli.
About the Magnetosome Toolkit
Using standard synthetic biology protocols and the vectors we created in our Gibson Assembly Toolkit, our team created the "Magnetosome Toolkit" which contains many of the genes required for magnetosome formation. Providing this toolkit to the Parts Registry will help allow future iGEM teams to manipulate and further understand magnetosome formation to eventually synthesize magnets in multiple organisms.
As previously noted, magnetosome formation within the host-organism, Magnetospirillium magneticum, strain AMB-1, is a highly regulated step-wise process. As shown in diagram of stepwise magnetosome construction above, some genes encode proteins that form an invagination of the inner membrane, other genes which help align the magnetosomes into their characteristics chains, and others which regulate the biomineralization of magnetic particles. Our team chose to focus on genes specifically related to magnetosome scaffolding/alignment since they are the essential foundation for magnetosome development. In addition, the creation of a scaffold to which other genes localize is highly applicable to systems in synthetic biology. (for more information, please see our Future Directions page)
The genes we focused on are mamK and mamI since they have known functions related to localization of the magnetosome. Specifically, MamK is a bacterial actin-like cytoskeleton protein required for proper alignment of the magnetosomes in a chain. MamK is also shown to localize the MamI, which when lost inhibits vesicle formation.
(for other gene functions, please see the iGEM Toolkits parts submitted page)
Toolkit Contruction
Please see our Result Summary page for to see how far we were able to get this summer!
References:
- Matsunaga, T., Okamura, Y., Fukuda, Y., Wahyudi, A.T., Murase, Y., Takeyama, H. (2005). Complete genome sequence of the facultative anaerobic Magnetotactic bacterium Magnetospirillum sp. strain AMB-1. DNA research; 12: 157-166. Doi:10.1093/dnares/dsi002.
- Murat, D., Quinlan, A., Vali, H., Komeili, A. (2010). Comprehensive genetic dissection of the magnetosome gene island reveals the step-wise assembly of a prokaryotic organelle. PNAS; 107 (12): 5593-5598. Doi:10.1073/pnas.0914439107.
- Murat, D., Quinlan, A., Vali, H., Komeili, A. (2010). Supporting Information. PNAS; 107 (12): 5593-5598. Doi:10.1073/pnas.0914439107.
- Quinlan, A., Murat, D., Vali, H., Komeili, A. (2011).The HtrA/DegP family protease MamE is a bifunctional protein with roles in magnetosome protein localization and magnetite biomineralization. Molecular Microbiology; 80 (4): 855-1131. Doi:10.1111/j.1365-2958.2011.07631.x.
- Richter, M., Kube, M., Bazylinski, D.A., Lombardot, T.,Glockner, F.O., Reinhardt, R., Shuler, D. (2007). Comparative genome analysis of four Magnetotactic bacteria reveals a complex set of group-specific genes implicated in magnetosome biomineralization and function. Journal of Bacteriology; 189(13): 4899-4910. Doi:10.1128/JB.00119-07.
- Rioux, J.B., Philippe, N., Pereia, S., Pignol, D., Wu, L.F., Ginet, N. (2010). A second actin-like mamK protein in Magnetospirillum magneticum AMB-1 encoded outside the genomic magnetosome island. PLoS ONE; 5(2): e9151. Doi:10.1371/journal.pone.0009151.
 "
"