Team:Washington/Alkanes/Future/DecarbDesign
From 2011.igem.org
(→In Vitro Assays) |
|||
| Line 1: | Line 1: | ||
| - | |||
{{Template:Team:Washington/Templates/Top}} | {{Template:Team:Washington/Templates/Top}} | ||
| + | __NOTOC__ | ||
| + | |||
<big> DECARBONYLASE REDESIGN </big> | <big> DECARBONYLASE REDESIGN </big> | ||
='''Background'''= | ='''Background'''= | ||
Revision as of 21:15, 16 September 2011
DECARBONYLASE REDESIGN
Background
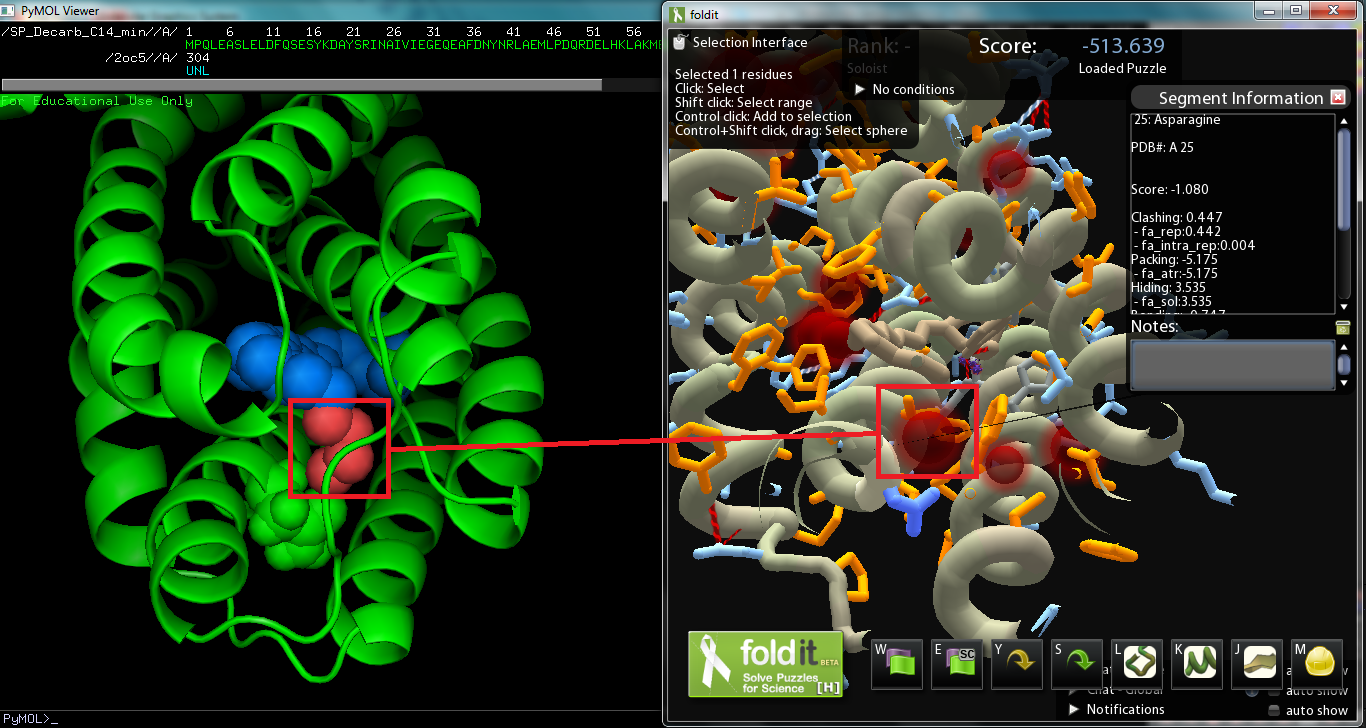
The objective of this sub-project was to modify the ADC protein so that we can synthesize a wider variety of alkanes; in other words, mutate the protein so that it works better on a range of saturated fatty aldehydes. The protein was originally surmised to work best on C18 aldehydes, as the crystal structure derived Pymol model of the protein revealed a C18 carboxylic acid bounded to the metal center (though the vector with AAR and ADC proved to work best on C16 aldehydes while less on C14 and C18 aldehydes). Consequently, we decided to strive to modify ADC to work on shorter chain aldehydes, specifically tetradecanal. The substrate on the original Pymol file was modified to model C14 aldehyde, and then the file was converted to a Fold-it puzzle for human interaction. We decided to avoid changing amino acids near the active site, which binds to the aldehyde group, as we wanted to maintain the basic chemistry, decarbonylation (or perhaps deformylation). There was also the issue of the Fold-it modeling 122-Tyrosine hydrogen bonded to the substrate and another side chain, and according to Fold-it's notifications, serine, threonine, and tyrosine cannot "have more than 1 donor and 2 acceptors." As the amino acid we will change will be around the alkyl chain, there are no hydrogen bonds to be made(hydrogen bonds show up on the interface). If we are to make changes so that the protein binds more favorably to C14 aldehydes, we must improve dispersion intermolecular interactions by increasing the surface interaction between protein and substrate. Comparison with the original Pymol file showed that the removal of the four carbon atoms created a spatial “void area.” Targeting the void area with minimal interference or clashes between atoms, the most promising mutation sites are on adjacent sections of two helices positioned opposite of the substrate’s carbon end. The adjacent sections take up amino acids 21 to 25 and 67 to 71, the former of which, being closer, shows promise as a site to mutate to fill in the void area and avoid interferences.
Methods
Several aldehyde decarbonylase (ADC) mutants were designed using Fold-It, a computational design program. These ADC mutants were made using the Kunkel Mutagenesis method. Also, the sequences of these mutants were later verified to carry the correct mutations. After creating several ADC mutants, three different types of assays were developed and the mutants were later tested and analyzed using Gas chromatography–mass spectrometry (GC-MS) to see if our mutants can produce high abundance of C13 alkanes. The three different assays that were set up are: cell lysis assay, protein purification assay, and in vivo assay.
In Vitro Assays
- Cell Lysis Assay
Cell lysis assay is the first assay we developed. One aldehyde decarbonylase (ADC) colony was picked and grown in the 37°C shaker until OD600 reached the value of 0.8. When it reached the value, culture was induced with IPTG and cell pellet was collected by spinning down after three more hours of shaking. This cell pellet was resuspended in a solution that contains sodium phosphate buffer, protease inhibitor, DNAses and lysozyme. This resuspended cell was lysed using 10% triton and then it was centrifuged to separate the cell membrane to the bottom and the cell’s inner machinery to the top. This top layer was transferred to a glass vial and used to set up in vitro reactions. In vitro reactions were set up by adding certain amount of cells, feredoxin, feredoxin reductase, NADPH and aldehyde. However, this assay had a one big flaw. It didn't yield consistent alkane production every time even though we kept variables very minimal. So our team decided to move on and develop a different assay, protein purification assay.
- Protein Purification Assay
This time, we decided to increase the concentration of ADC and decrease the number of contaminants in the assay. We grow up cells to express 10 times more ADC proteins and then we purify the proteins by lysing the cells and binding the target ADC proteins to the BioRad columns and then eluting them out, resulting in pure ADC proteins. Then, we setted up in vitro reactions by adding purified ADC mutant protein, standard buffer(Hepes at pH8, NaCl, glycerol, TCEP and water) and feredoxin, feredoxin reductase, NADPH, and aldehyde into an eppendorf. After certain time period has passed, it was quenched in ethyl acetate and the top organic layer that results from quencing was extracted to an insert in a GC-MS vial. This was later analyzed in the GC-MS. With only a week or two (at most) left of experiment time, we basically have to give up on a second round of mutations. Also, the GCMS results with the purified proteins showed no alkane (product of reaction) peaks at all. This could be explained by the fact that the freezer in which we stored glycerol stocks of our purified protein was actually -25C instead of the displayed -50C. However, since we did not know when the freezer began malfunctioning, so it may be just as likely that aldehyde decarbonylase cannot be stored and reused. So we moved onto working with in vivo assay which we started to develop at the same time we started to work on protein purification assay.
In Vivo Assay
We took the base construct RED-PSB1C3 (Acyl-ACP Reductase in PSB1C3 high constitutive vector) and made the 1C3-const-RED-rbs-mutant decarb construct (1C3 construct with RED, ribosomal binding site, and the mutant decarbonylase genes made with Kunkel (site directed) mutagenesis. Then we transformed the competent cells (BL-21) to take up construct plasmids with our mutated ADC genes. We then grow the transformed competent cells in TB with chloramphenicol and then plate it on TB-chloramphenicol agar plates. Individual colonies were taken to inoculate water, and the colony waters were taken to run colony PCRs to confirm the uptake of the correct plasmid constructs that are with mutated ADC genes. Once verified, the corresponding colony waters of the successful colonies were grown in TB-chloramphenicol media overnight. The media are then miniprepped to obtain DNA of the original plasmid construct taken up by BL-21. The miniprepped plasmid construct is then transformed into another form of the competent cells (MG1655) and then plated on TB-chloramphenicol agar to have samples of different genetic variety of samples. Six isolated colonies are picked on each plate (to average out genetic differences) to be grown in special alkane production media M9 for 48 hours. After that, alkane is extracted with ethyl acetate and the samples are then analyzed with GCMS machine.
In vivo (in the cell, start from glucose) : cells’ membrane will keep proteins in smaller volume, but the main point is that we can compare with native ADC cell results from other groups (the cells probably will make more alcohol anyway).
Current Status/Results
- Discuss mutations made and construct submitted
- Show protein gel of expression
- Discuss in vitro assay and how it hasn't worked yet (even for WT)
- Discuss plan for future testing
Parts Submitted
- Plasmids with mutations
- ADC_AB4
- 1C2
- 7A5
- 8BE
- 9A1
- 10B5
 "
"