Team:Groningen/modeling results
From 2011.igem.org
| Line 53: | Line 53: | ||
==Visulisation of out model== | ==Visulisation of out model== | ||
| - | Because we did not manage to finish our | + | Because we did not manage to finish our complete device we decided to try and model our device in the Cumulus system without any measurement data. One of the main problems in assembling our device in the wetlab seemed to be the defective hybB promoter. For this reason in this model we replaced it with a LacI promoter. In order to operate our system we also assume that we can remove both arabinose and IPTG from the medium in an effcient way. As you can see with these small modifications we get a perfectly working system. |
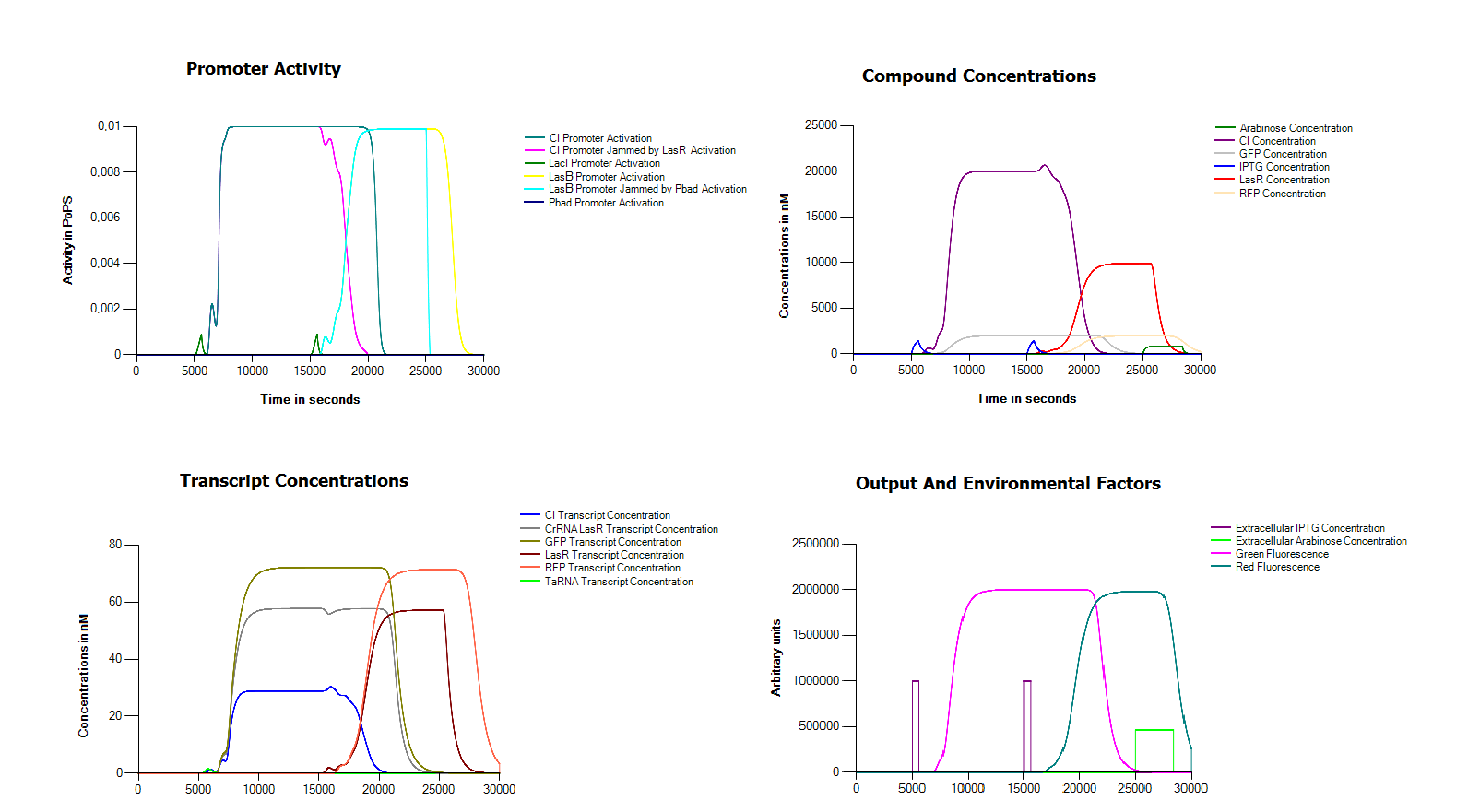
[[File:Groningen2011FullSimulation.png|thumb|centre|600px|This figure shows a simulation of our full model. As you can see in the bottomleft corner we induce twice with IPTG for ten minutes, once after 5000 seconds and a second time after 15000 seconds, finally reset the system with a long exposure to arabinose in the medium after 25000]] | [[File:Groningen2011FullSimulation.png|thumb|centre|600px|This figure shows a simulation of our full model. As you can see in the bottomleft corner we induce twice with IPTG for ten minutes, once after 5000 seconds and a second time after 15000 seconds, finally reset the system with a long exposure to arabinose in the medium after 25000]] | ||
Revision as of 13:08, 21 September 2011
Results
A simple characterization example
In its most simple use the Cumulus system can be used to characterize promoters. The example below shows how this can be done for the Pbad Promoter (BBa_I0500) with flowcytometry data. The measurements points in this case are generated by comparing as measurement of an unknown part against two measurements from a well characterized one.
Setting up the paramter space
The Code blow shows how we set the initial parameters for the Pbad Promoter. In it you see parameters gettign added to a parameter set. The arguments of the operation are the part name, followed by a the parameter name and then finally a series of three numbers, the first being the estimated parameter value, the minimal value we think it can take and the maximal value we think it can take. Defining these are the responsibility of the user.
Dim ParameterSetting As ParameterSetting = New ParameterSetting()
ParameterSetting.Add("Pbad Promoter", "LeakageActivation", 0.0001, 0, 0.01)
ParameterSetting.Add("Pbad Promoter", "MaximumActivation", 0.02, 0.01, 0.02)
ParameterSetting.Add("Pbad Promoter", "KMid", 20000, 10000, 40000)
ParameterSetting.Add("Pbad Promoter", "Cooperativety", 2, 1, 3)
ParameterSetting.Add("Arabinose", "Halflife", 2520, 600, 3600)
ParameterSetting.Add("GFP", "Halflife", 2520, 600, 3600)
ParameterSetting.Add("GFP Transcript", "Halflife", 240, 180, 300)
ParameterSetting.Add("GFP Transcript", "RIBS", 0.02, 0, 0.1)
ParameterSetting.Add("GFP Transcript", "Delay", 300, 120, 600)
ParameterSetting.Add("Pbad Sequence", "CopyNumber", 10, 9, 11)
ParameterSetting.Add("Pbad Sequence", "TranscriptionTime", 300, 120, 600)
ParameterSetting.Add("Extracellular Arabinose", "TransportRate", 0.00001, 0, 1)
Setting up the model
The second code segment shows how the actual model is constructed. As you can see it is build from smaller modeling components as described here. The first argument a part receives is always the ParameterSetting that was constructed above, from this it the component will draw the parameters that govern its behavior.
Dim Arabinose As Compound = New Protein(ParameterSetting, "Arabinose") Dim GFP As Compound = New Protein(ParameterSetting, "GFP") Dim GFPGene As Gene = New Gene(ParameterSetting, "GFPGene", GFP) Dim PbadPromoter As AdditiveHillPromoter = New AdditiveHillPromoter(ParameterSetting, "Pbad Promoter", "Arabinose") Dim GFPTranscript As Transcript = New RegularTranscript(ParameterSetting, "GFP Transcript", GFPGene) Dim PbadSequence As Sequence = New Sequence(ParameterSetting, "Pbad Sequence", PbadPromoter, GFPTranscript) Dim Sequences As List(Of Sequence) = New List(Of Sequence) Sequences.Add(PbadSequence) Dim Compounds As List(Of Compound) = New List(Of Compound) Compounds.Add(Arabinose) Compounds.Add(GFP) Dim Environmentals As List(Of Environmental) = New List(Of Environmental) Environmentals.Add(New ExtracellularConcentration(ParameterSetting, "Arabinose")) Dim Cell As Cell = New Cell(Sequences, Compounds, Environmentals) Return New Model(Cell)
Running the model
The movie blow shows how the algoritm searches movie
Visulisation of out model
Because we did not manage to finish our complete device we decided to try and model our device in the Cumulus system without any measurement data. One of the main problems in assembling our device in the wetlab seemed to be the defective hybB promoter. For this reason in this model we replaced it with a LacI promoter. In order to operate our system we also assume that we can remove both arabinose and IPTG from the medium in an effcient way. As you can see with these small modifications we get a perfectly working system.
Speedup
The Cumulus system allows for large speedups by parallelization through both the cloud and added machines. We tried this with some typical cloud intances and some local machines in the lab, the results are listed below. The benchmarking was done the the Pbad characterization task described above.
 "
"