Team:Paris Bettencourt/Experiments/T7 diffusion
From 2011.igem.org

T7 diffusion experiments
Cloning
The cloning for this system, that seems simple at the beginning took us a very long time, because of many issues we encountered along the cloning.
The first row of problems come from wrong parts sent by the registry. We lost almost one month and an half in building the system. (see the corresponding page). We synthetized ourself the pHyperSpank promoter (K143055), but we didn't have the time to incorporate it into our biggest construction. Instead, we cloned a pVeg SpoVG promoter (K143053) that have a constitutive expression.
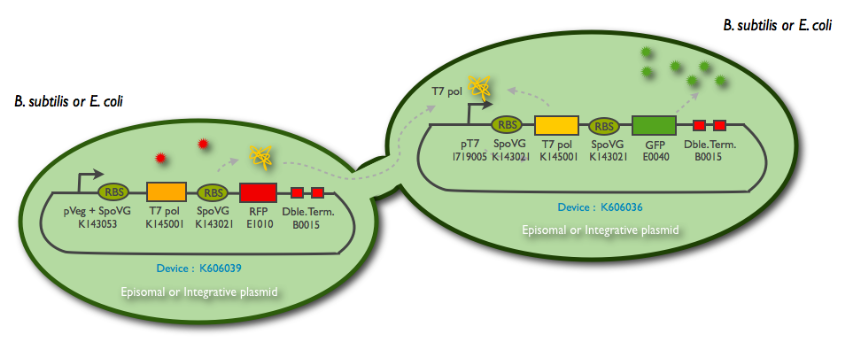
The following schematic represents what we have indeed managed to clone:
This summary shows the parts we have characterized, and the ones we have sent to the registry. But we have successfully managed to clone them into an integration (K090403) and a replicative (pHM3) vector for subtilis, as well as the combination of the emittor and the receiver inside the same plasmid.
Diffusion experiments
We are around the corner of bringing this experiment under the microscope. We already have built the microfluidic device, and it is just waiting for our cells.
At the moment, we have some difficulties with the transformation of our B. subtilis strain but we hope we could show you great results and images for the Amsterdam iGEM Jamboree!!
We have characterized this part transforming it into a BL21 strain. This strain carries insite the chromosome a T7 polymerase under the control of an IPTG inducible promoter. This strain is mostly used for protein expression for protein purification application.
Characterization of the T7 promoter
Growth
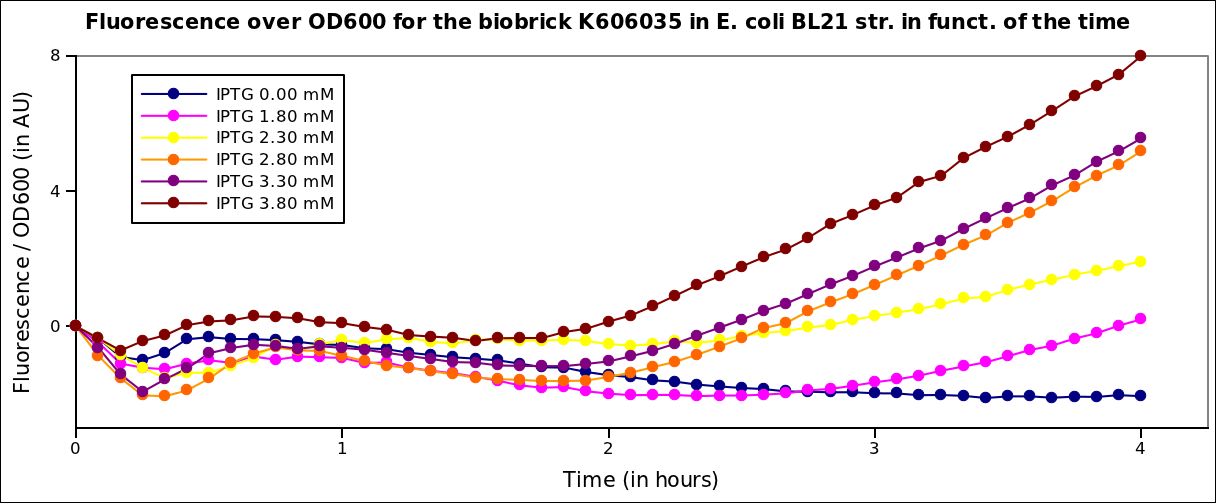
The measurements had been carried on in a TECAN i-control machine, at 37°C under transcient shaking, for 4h, for several colonies and several range of IPTG concentration. The OD 600nm and the fluorescence of the GFP (exc: 470nm / meas:515 nm) was measured every 5 min, and the ratio of the two was calculated.
The offset values for these curves was adjusted for better visualisation. The values given are in arbitrary units.
First we see an inflexion in the curve, that is due to the stong influence of the IPTG on the metabolism of the cells. Then, the bacteria start growing again. We see a clear increase of the fluorescence with the IPTG concentration, that is to say with the quantity of T7 polymerase in the cell.
Comparison of the growth with the traduction saturated cells
As a positive control we have saturated a strain with a lot of IPTG. After 1h50 of growth, we compare the fluorescence of the gradient of IPTG with the saturated cells. We see a clear increase of the fluorescence whereas the saturated strains are quite stable.
The offset of the curves is here renormalize at the inflexion point of the growth, when the cells start have passed the IPTG stress and are growing again.
Characterization of the T7 signal amplification leakage in E. coli
We characterized T7 autoloop in E.coli, when hosted in the plasmid pSB1C3 (K606036). The idea was to see if the system was leaky inside a synthetic biology plasmid, that is to say, a plasmid holding 4 terminators before any construct.
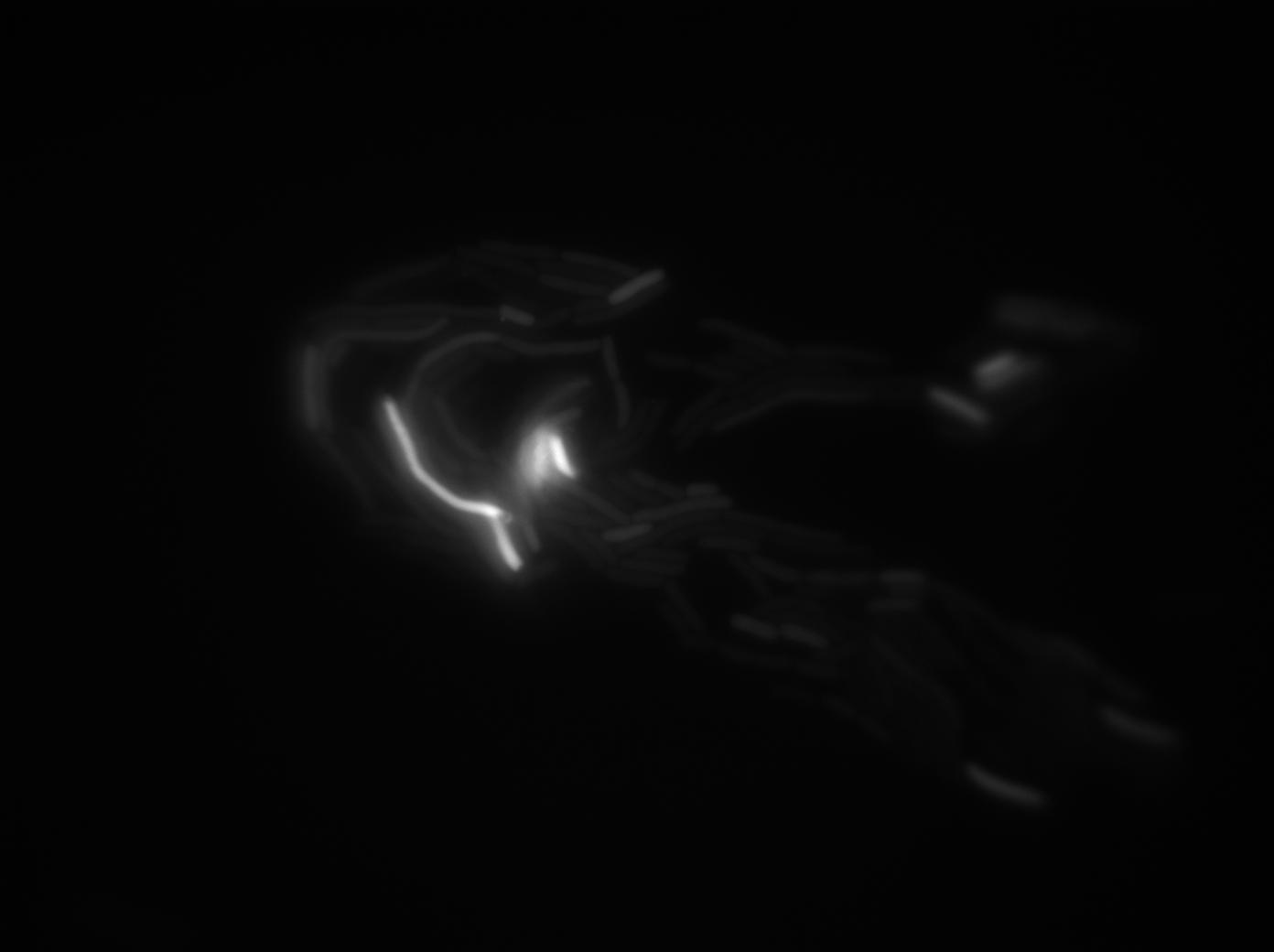
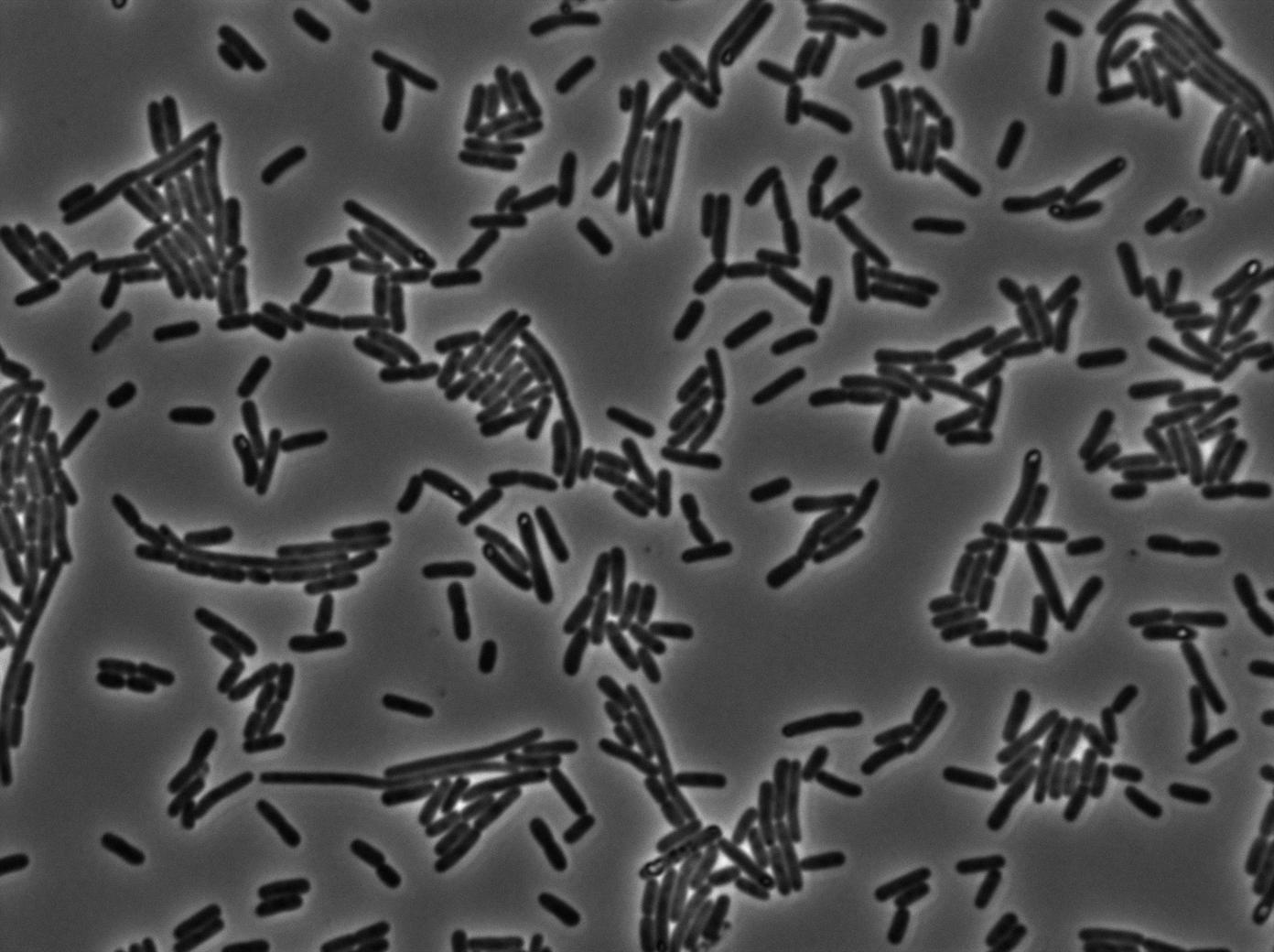
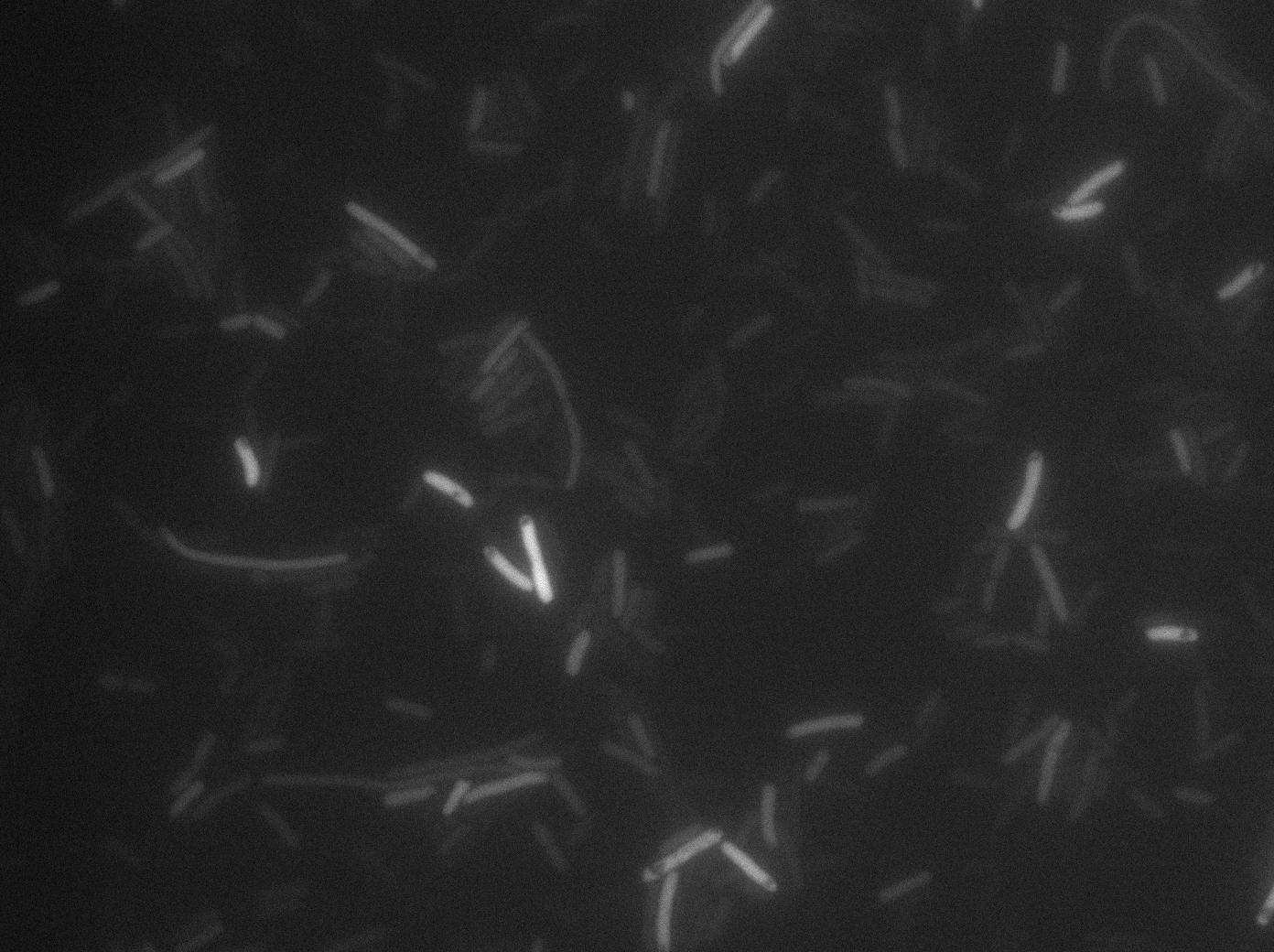
We found out that there is a leakage, but it is small. The cells in which the positive feedback loop is activated stop dividing and glow with a very strong signal. Here are the pictures commented:
The first pictures show that RFP construct has been well done. Indeed, we can see that some cells are glowing with RFP fluorescence. This also shows that the system is not as leaky as we expected for. In deed, the promoter regulating RFP expression is 'Pveg' which is a constitutive promoter.
Finallyr, the autoloop system is working very well because when leak occurs, cells glow highly.
This pictures show that GFP system gives more disparity than the RFP construct. This is probably due to the LB that already glows in green. It is also noticeable that B.subtilis is naturally shining green. Moreover, the absence of double terminator could explain the leakiness of our system. However, we expected this cells to glow in RFP but they did not. This shows that the full construct has failed.
The last pictures show less disparity that the previous one. This shows that double terminator actually proceeds on the phenotype. We also can notice that the cells glow in red a bit. However we can not say if it is an artifact or if the system is actually well constructed with the terminator and RFP construct.
 "
"