Team:DTU-Denmark/Project testing sRNA
From 2011.igem.org
Project
Contents |
Introduction
The project is inspired by three components that regulate chitobiose transport in E. coli: (1) chitoporin chiP (alias ybfM) that facilitates uptake of chitosugars from the medium into the cell; (2) sRNA chiX (alias sroB, micM) which regulates chiP post-transcriptionally; and (3) chiXR sRNA which is transcribed from intergenic region in chbBCARG operon and regulates chiX sRNA. ChiX sRNA regulates chiP expression through binding to the Shine-Dalgarno sequence on the chiP mRNA and thus inhibits recruitment of 30S ribosome subunit and translation cannot occur. Only when chitobiose is present in the medium can the chiXR be transcribed and subsequently its transcript can bind to chiX sRNA, releasing chiP repression.
Motivation
The biological experiments of the project is a proof of concept, showing that the E. coli sRNA sroB and the intergenic sRNA (trap-RNA) in the chbBCARG gene can be used to control gene expression by targeting the ybfM Shine-Dalgarno, and that these sRNAs can be rationally designed to target other Shine-Dalgarnos.
The experimental part consists of 3 distinct areas:
- Construction of plasmids
- Strain construction
- Improving the araBAD promoter
Construction of plasmids
In order to quantitatively describe the natural trap-RNA system the following three plasmids were constructed:
- PchiP-lacZ plasmid
- To mimic and test repression of the chiP gene its upstream region of 599 bp was fused to reporter genes lacZ and GFP and inserted on the pSLD39 plasmid. This constructs were controlled by constitutive promoter. LacZ codes for $\beta$-galactosidase and its activity was tested using $\beta$-Gal assay while GFP activity was tested using Biolektor.
- Ptet-chiX plasmid
- ChiX, which is 84 bp long, was inserted into pSB4K plasmid together with regulatory element from tet operon. Expression from Ptet promoter was induced using anhydrotetracycline.
- PBAD-chiXR plasmid
- Introgenic region from chbBCARG operon 284 bp long was inserted into pBAD18 plasmid. Expression from that plasmid was controlled by regulatory element from ara operon with arabinose as inducer.
Strain construction
The strain used in testing our system is based on the E. coli strain W3110, a wild-type strain of Escherichia coli K-12[3] with the lacZYA, chbBCARG operons and chiP (alias ybfM), chiX (alias sroB, micM) genes deleted from the chromosome. The three latter code for utilization of chitobiose (a sugar), chitobiose permease and the regulation of the chitobiose permease respectively.
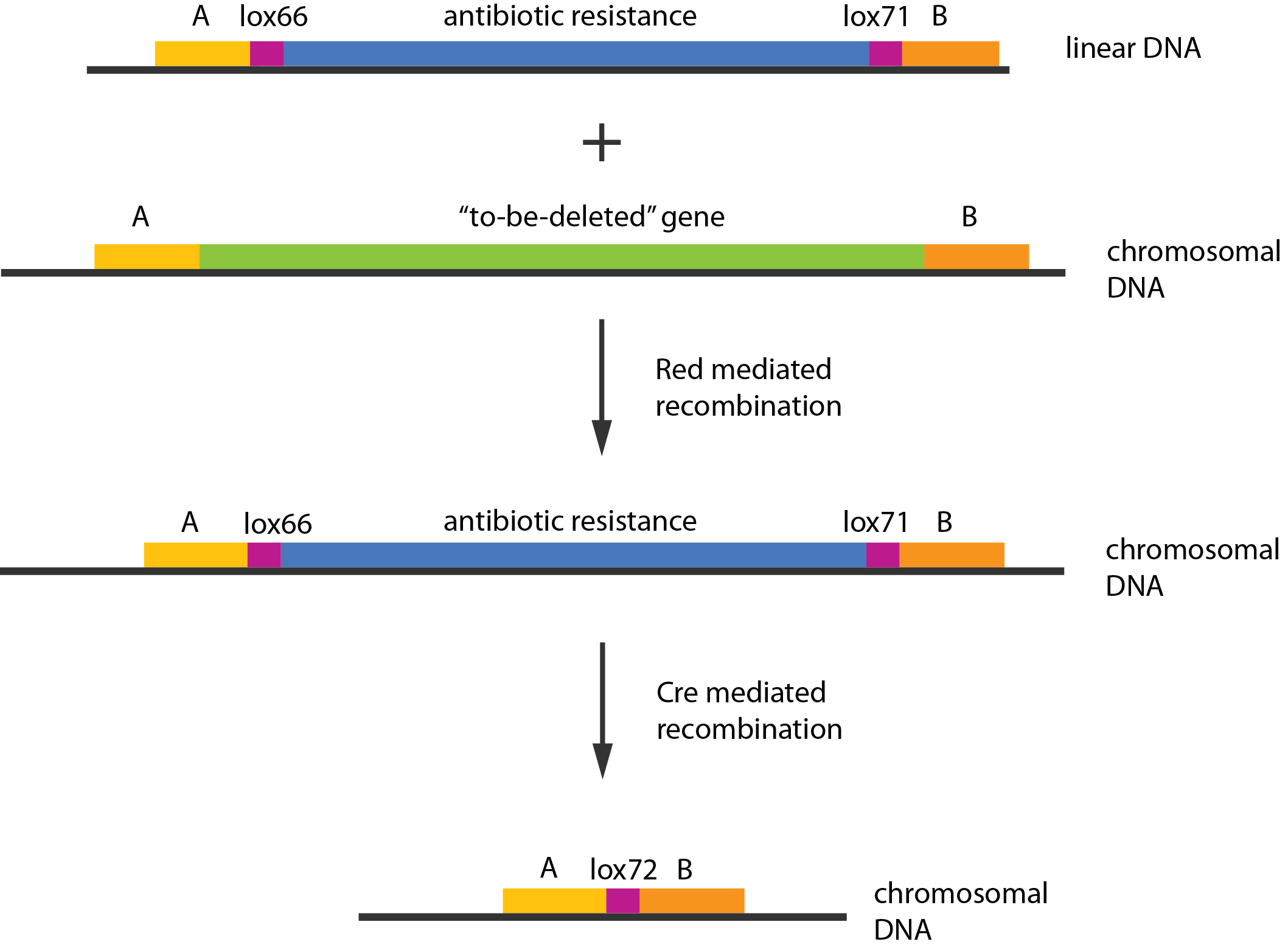
A procedure known as redswap[1] was used to delete these genes. Schematically the procedure works in the following manner:
- Transform competent cells with Red and cre helper plasmids.
- Make the cells competent and induce the Red recombinase.
- Transform with linear piece of DNA.
- Select for gain of resistance.
- Move to medium inducing the Cre recombinase.
- Screen for loss of resistance.
- Verify the construct.
- Repeat step 2-7 as needed.
- Cure the helper plasmids.
The Red helper plasmids encodes the $\lambda{}$ phage Red recombination system. This system allows recombination events between sequences with around 40 nucleotides of homology. The cre helper plasmid encodes the Cre recombinase. This recombinase recognises 34bp loxP sequences and promotes recombination between them.
An example of a linear piece of DNA that could be used in redswap, as well as the simplified representation of the procedure, can be seen in figure \ref{linDNA}.
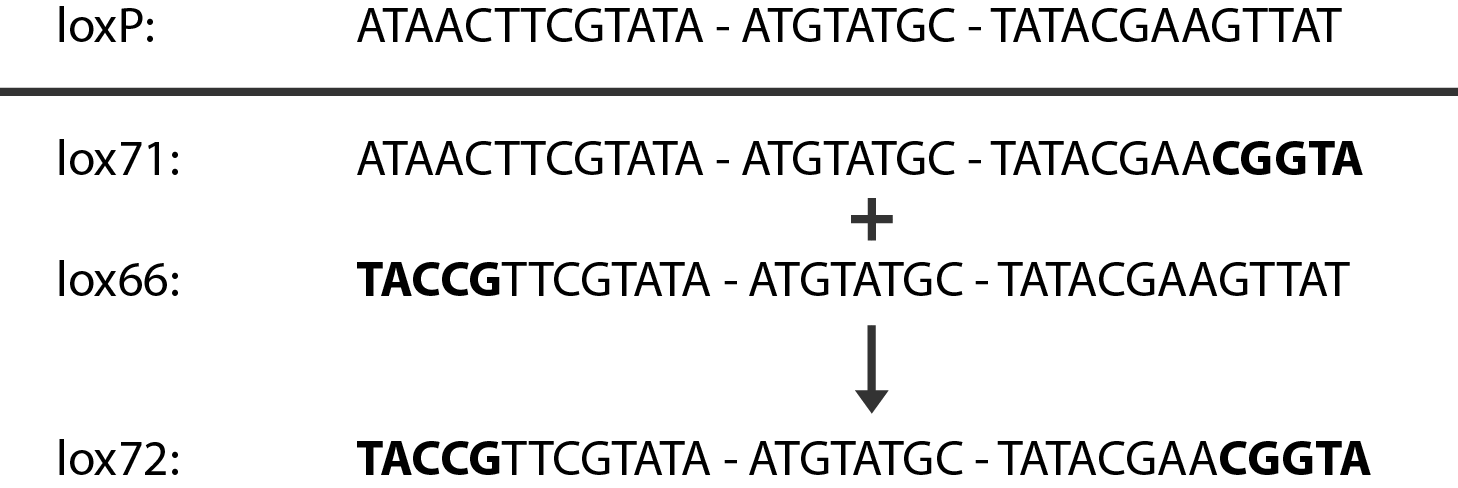
In case of two or more subsequent gene deletions using redswap a problem of recombination between an old loxP site and newly introduced ones arises. Remember, that after each round of redswap one loxP site is left in the genome. As a result recombination using Cre recombinase in second or next redswap might not occur between loxP sites flanking a resistance gene. To circumvent this problem two mutated loxP sequences are used - lox66 and lox71[4]. In spite of the mutations these lox sites are recognized by Cre which can remove the intervening region. Finally, from lox66 and lox71 a new site is formed, lox72, which has a reduced affinity to Cre recombinase. Thus, more than one gene deletions can be subsequently conducted.
References
[1] Datsenko, K.A. & Wanner, B.L. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proceedings of the National Academy of Sciences 97, 6640-6645(2000).
[2] Figueroa-Bossi, Nara, Martina Valentini, Laurette Malleret, and Lionello Bossi. “Caught at its own game: regulatory small RNA inactivated by an inducible transcript mimicking its target.” Genes & Development 23, no. 17 (2009): 2004 -2015.
[3] Hayashi, K. et al. Highly accurate genome sequences of Escherichia coli K-12 strains MG1655 and W3110. Molecular Systems Biology 2, 2006.0007(2006).
[4] Lambert, J.M., Bongers, R.S. & Kleerebezem, M. Cre-lox-based system for multiple gene deletions and selectable-marker removal in Lactobacillus plantarum. Applied and environmental microbiology 73, 1126-35(2007).
[5] Overgaard, Martin, Jesper Johansen, Jakob Møller‐Jensen, and Poul Valentin‐Hansen. “Switching off small RNA regulation with trap‐mRNA.” Molecular Microbiology 73, no. 5 (September 2009): 790-800.
 "
"