Team:Paris Liliane Bettencourt/Notebook/2011/09/15/
From 2011.igem.org

Contents |
Baptiste, Hovannes & Ouriel
Microscopy experiments of today.
As we begin to master totally the Nikon microscope we chose to run three settings for the 3610 gfp-/3610 gfp+ experiments. The aim is to see which way is the best to plate them on our slides and more important at which growth state. We start for the same overnight culture and then create three microscopic slides from our two 3610 strains:
- One directly from the overnight culture in stationnary phase.
- One diluting until the OD reached 0.1. We then wait for the OD to grow to 0.8 (roughly 2 hours) and we concentrate it as usual (4000 rpm during 10 min and then put 200 microliters of LB on the pellet).
- One by diluting 1/500 the overnight culture. We then wait for the OD to grow to 0.8 (roughly 4 hours) and we concentrate it as usual (4000 rpm during 10 min and then put 200 microliters of LB on the pellet).
First conclusions on our methods
Despite what we thought, the autofocus programme on the Nikon is not perfect. We need to adjust it a little bit to our needs. It gave us slightly blurry image for the second serie of slide (dilution to an OD of 0.1) and catastrophic results for the stationnary phase slide. Fortunately, the stationnary phase means that bacteria do not grow and we were able to interpret the results even missing over an hour of images. Our slide for the 1/500 dilution was very faulty. We could not see anything clearly and we will have to retry it tomorrow. Letting the drop dry or not is the primary issue here. We will try both ways tomorrow to see which seems better.
Stationnary phase
As expected, no growth and no diffusion was observed.
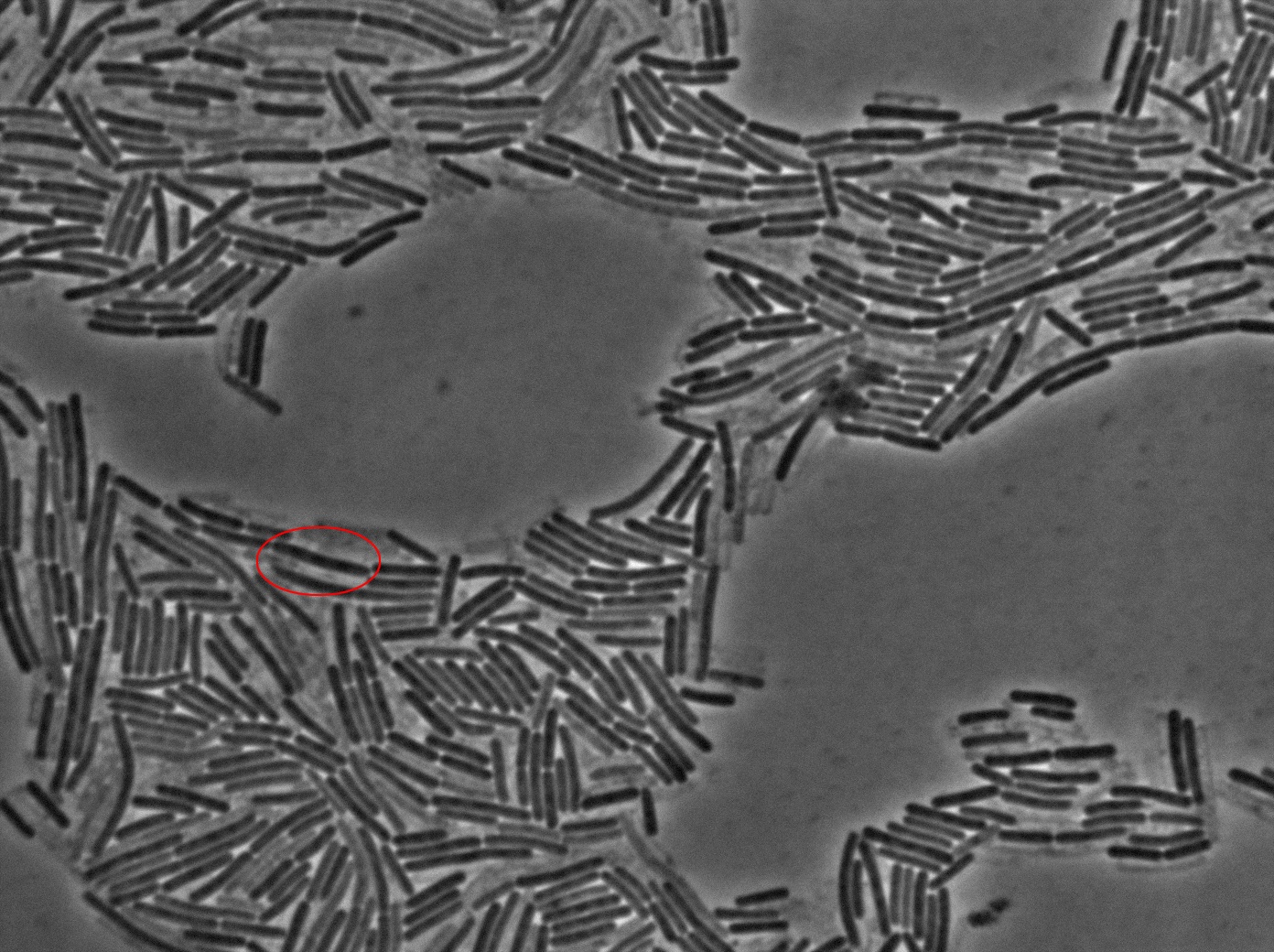
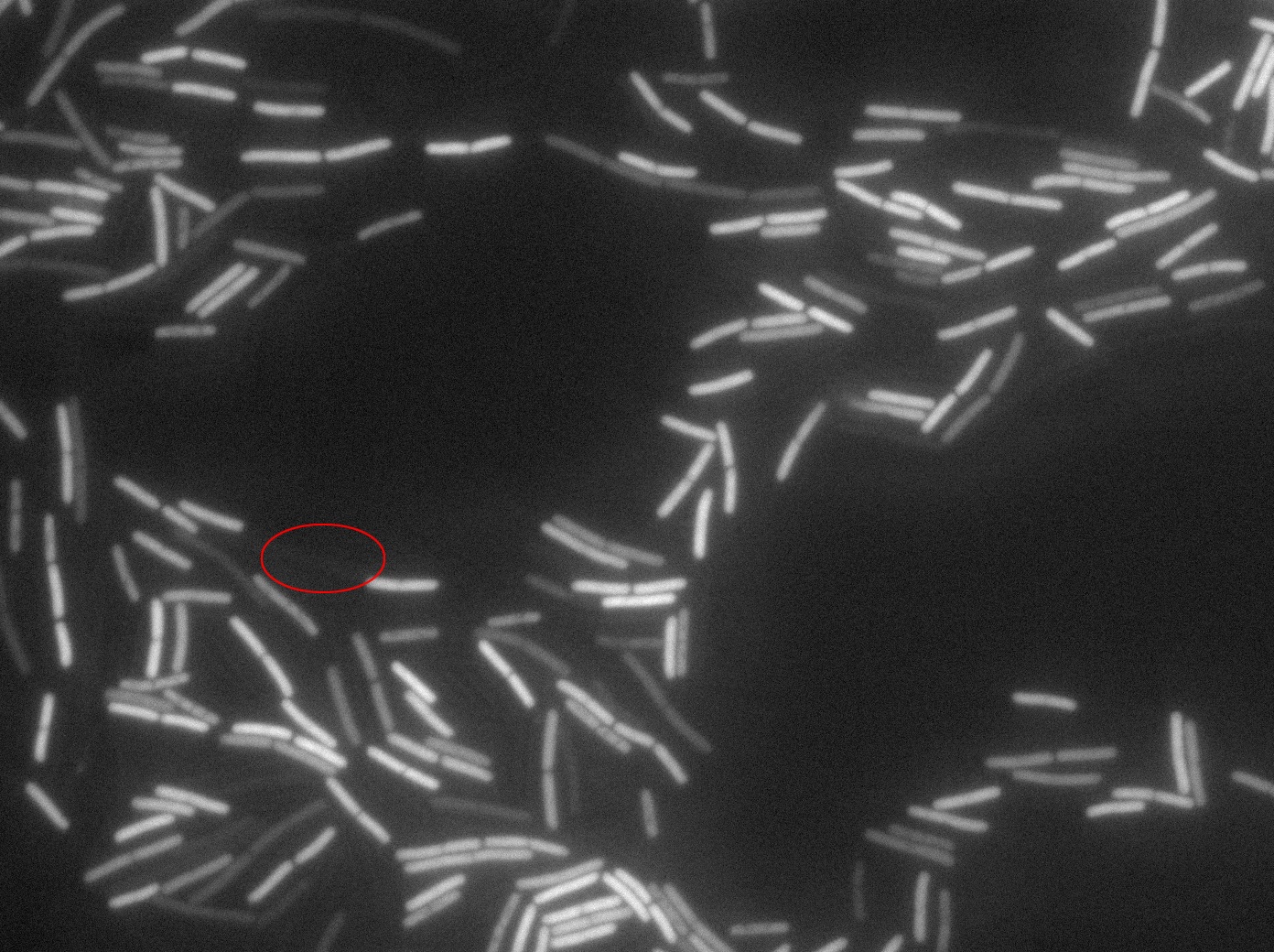
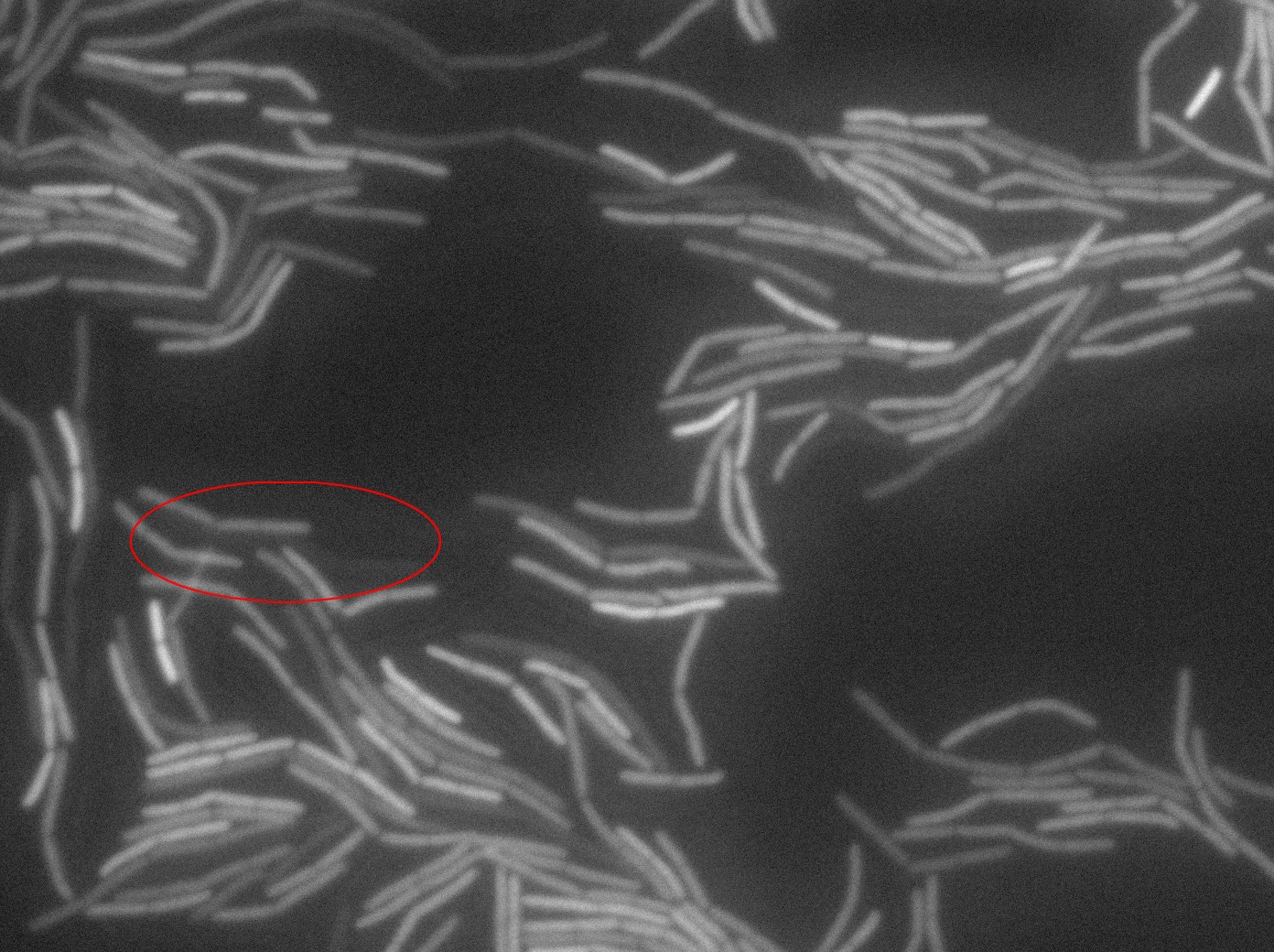
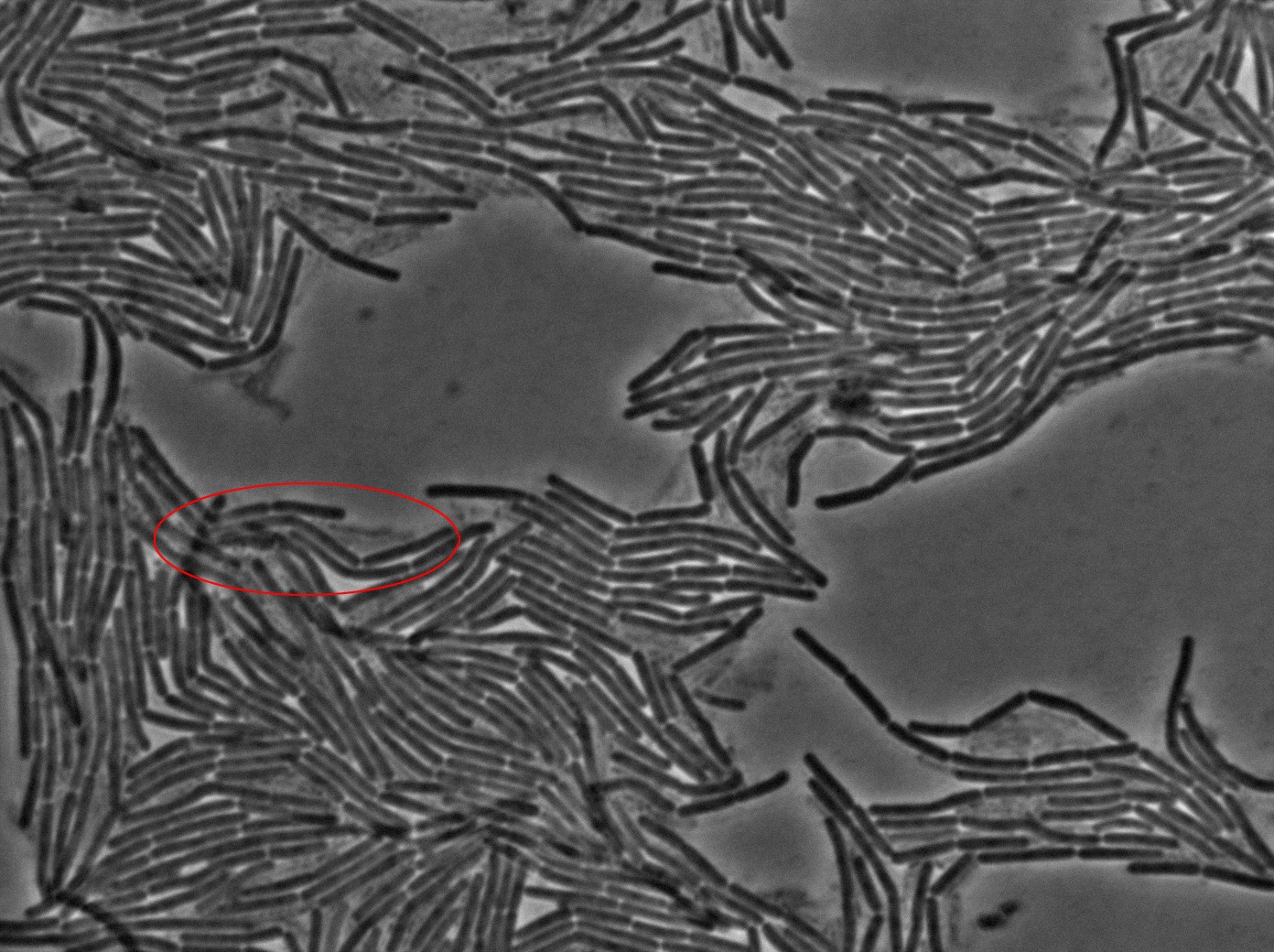
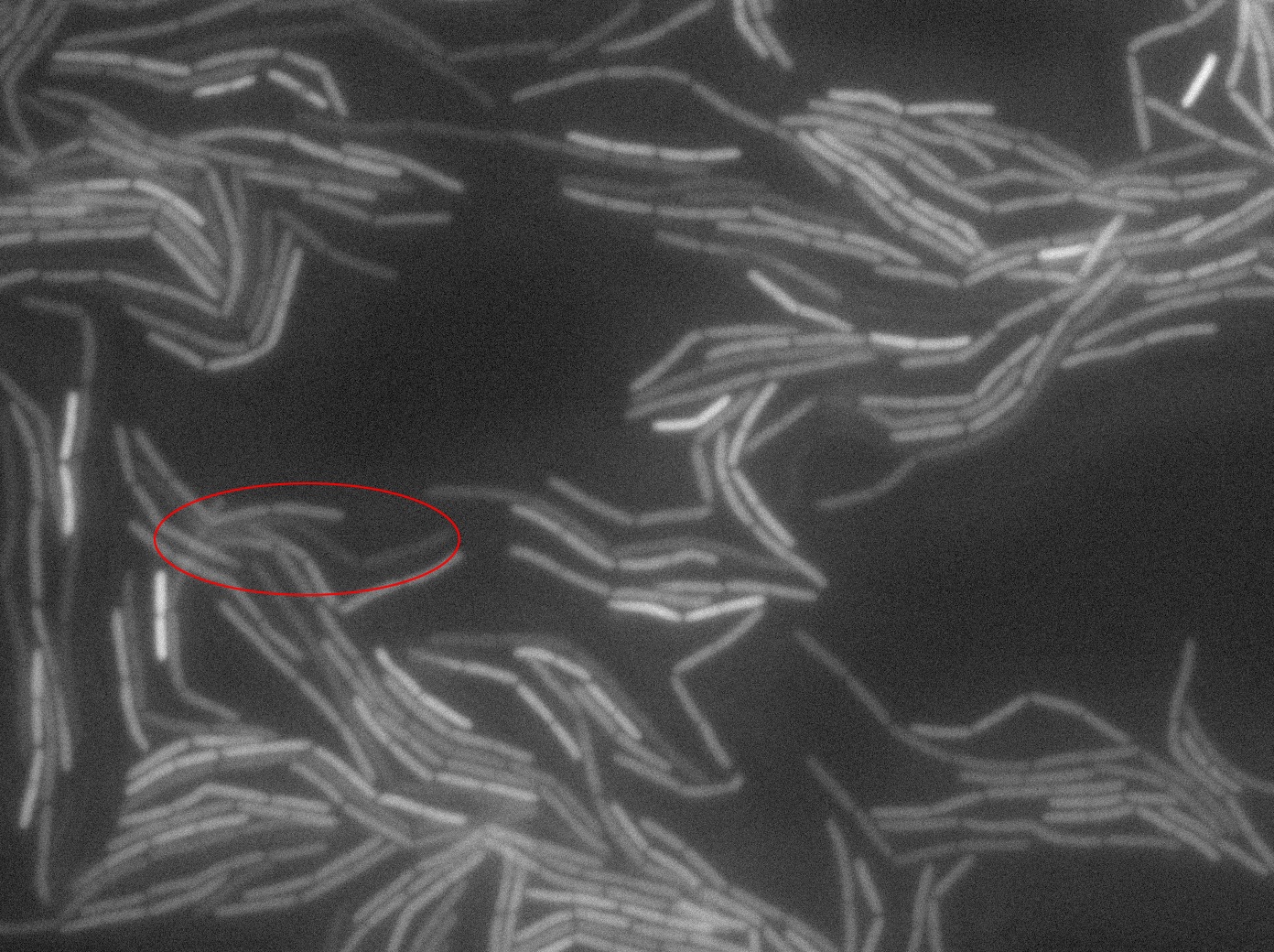
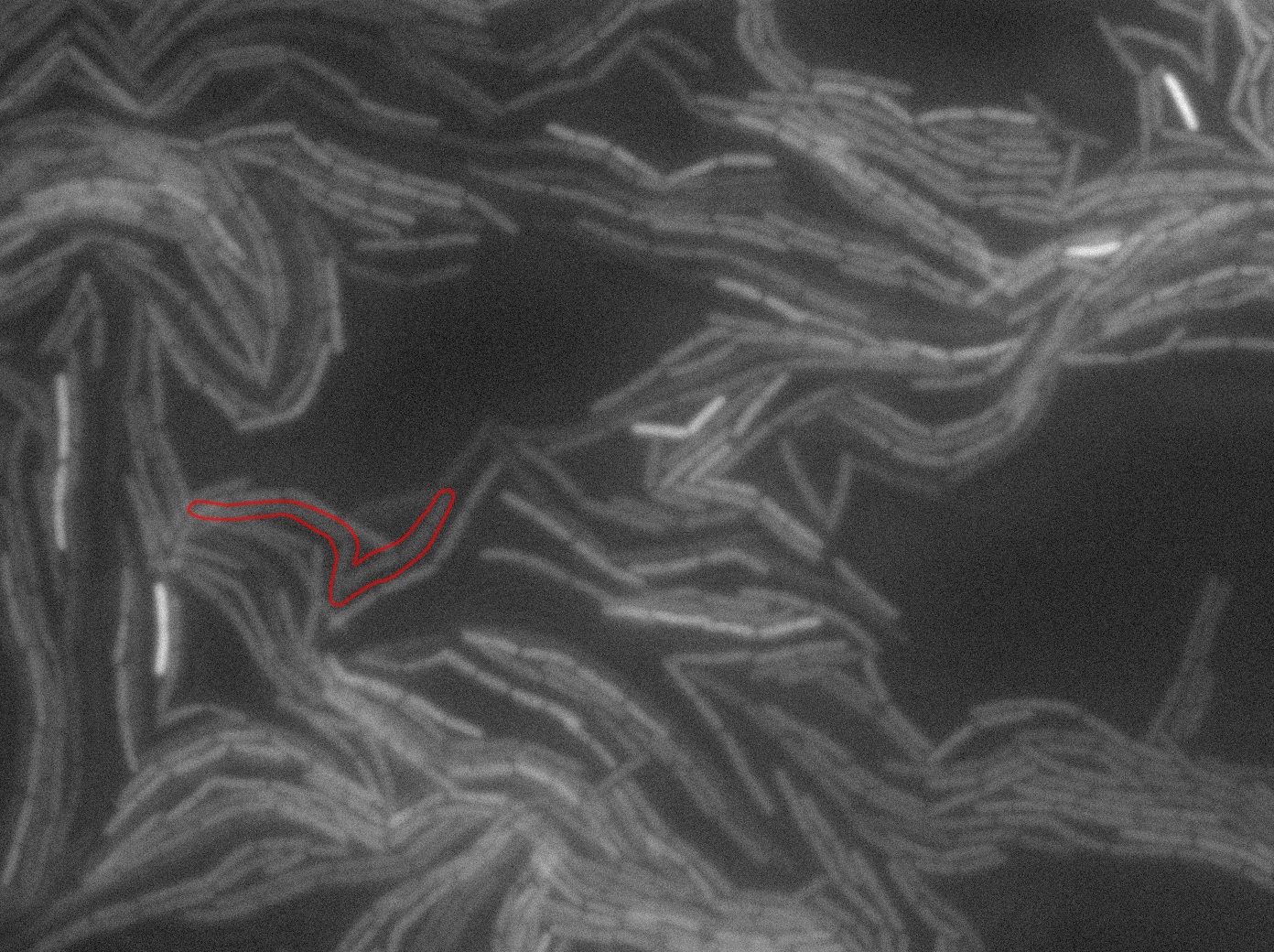
Dilution to an OD of 0.1
The slide was one of the best we've done so far. We were able to see clearly zones with a monolayer of roughly 50/50 GFP and non-GFP cells. And we finally found some evidence of deiffusion in one of the followed sites! The cells gaining fluorescence are circled in red. Note that all the other fluorescent cells have lost fluorescence during the experiment. the ones circled are gaining fluorescence. We also ran positive controls but took pictures only at the end of the experiment.
Kevin
Following step for characterisation of tetO array
Double transformation works ! Tomorrow microscopy.
cyrille
Treatment of the ligations done yesterday. PVeg T7a TT + pT7 gfp TT worked. 16 col putted in culture.
For the other ligations, the results of the positive control indicates that the transformation failed. 1 clone was found for p/eg tRNA rfp, and 3 pVeg tRNA tt rfp they were insulated. PCR colonie failed. The cultures will be minkprepped tomorow.
The ligation that are supposed to have failed were done again. As we were short on pHM3, the S99 was not done again.
Miniprep of pHM3 overday.
Suspicion on pHM3 on the lost of replication origin for subtilis due to the analysis of the sequencing. 7 miniprep of the original pHM3 were launched. Double digestion in HincIII and PstI was help to see if the region was indeed lost. HindIII cut several times in the replication origin region.
 "
"