Team:EPF-Lausanne/Our Project/T7 promoter variants/selection
From 2011.igem.org
Lysis Selection System
Lysis selection system Main | Lysis Characterization | DNA Recovery | DNA Selection | T7 Promoter VariantsContents |
DNA Selection
Introduction
To round out the experiments for DNA recovery, it is essential that we show that the recovered DNA is in fact plasmid DNA from the relevant lysed cells. This is important because in our selection scheme we are only interested in the plasmids from cells expressing the lysis cassettes, and not the DNA from their neighbors (probably the majority of cells in a real selection).
Experimental Setup
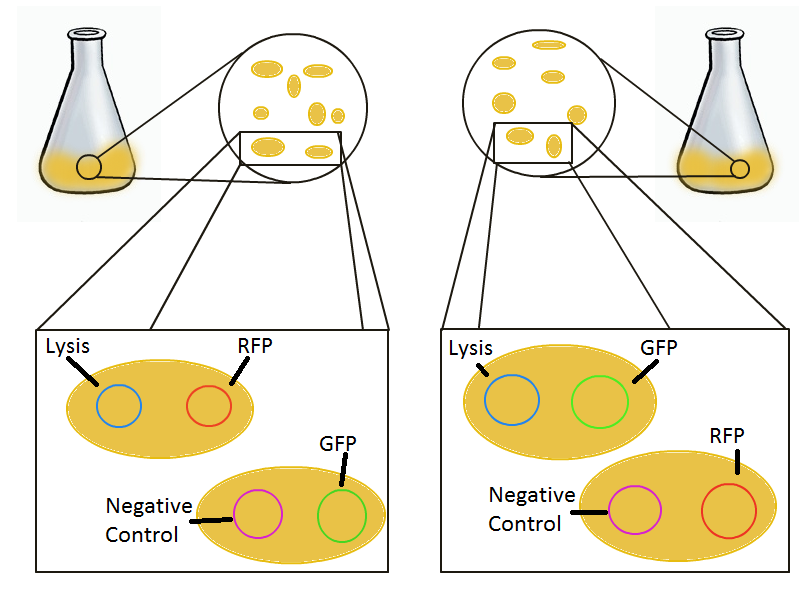
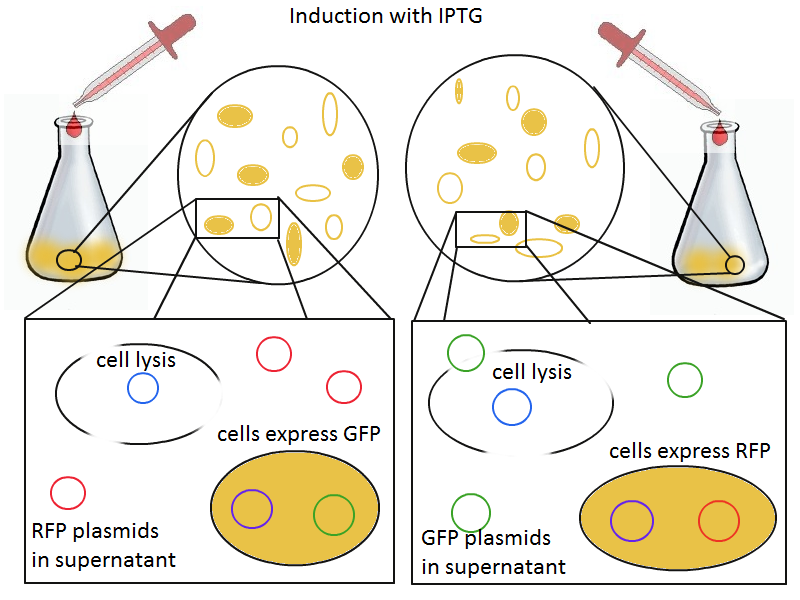
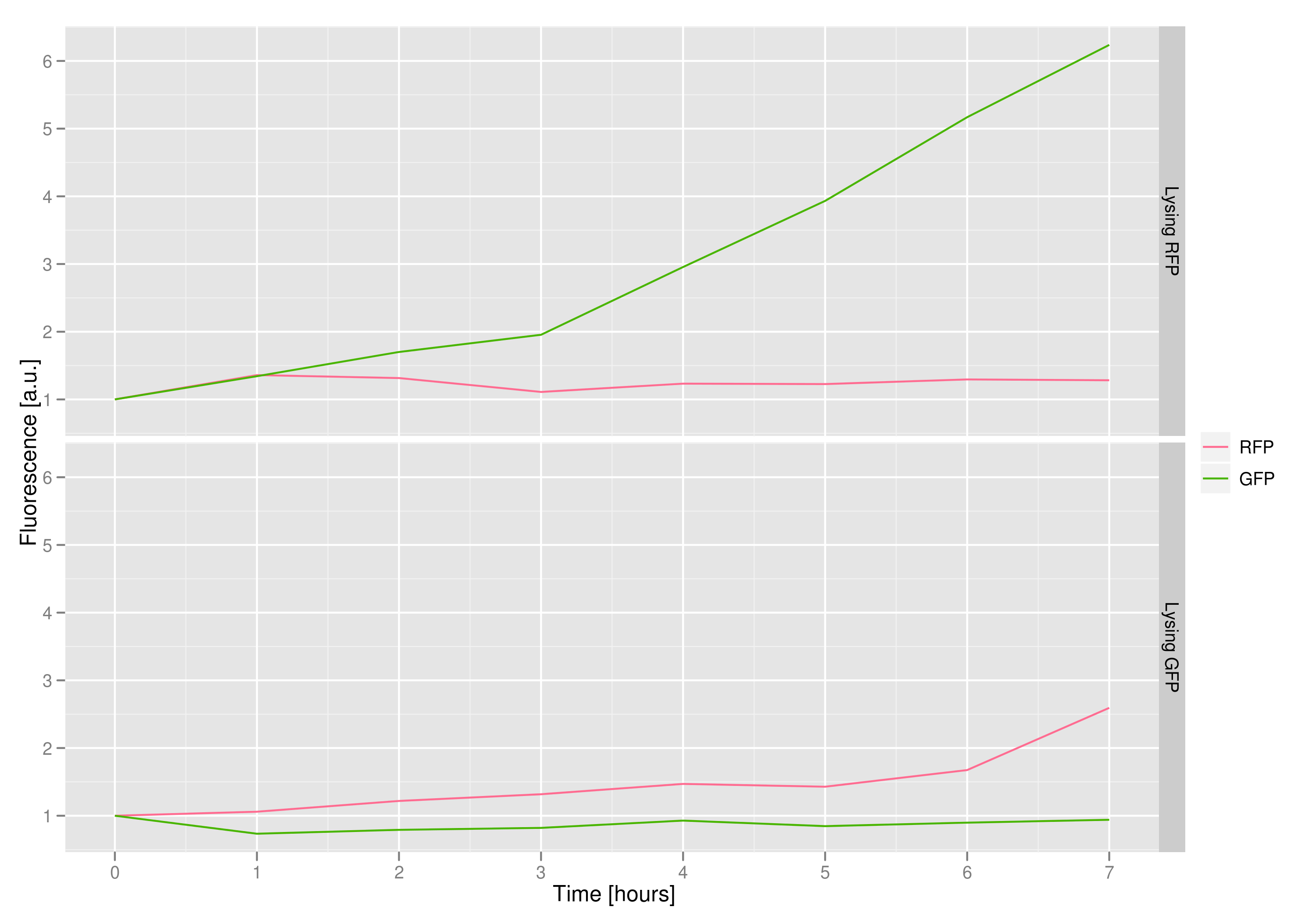
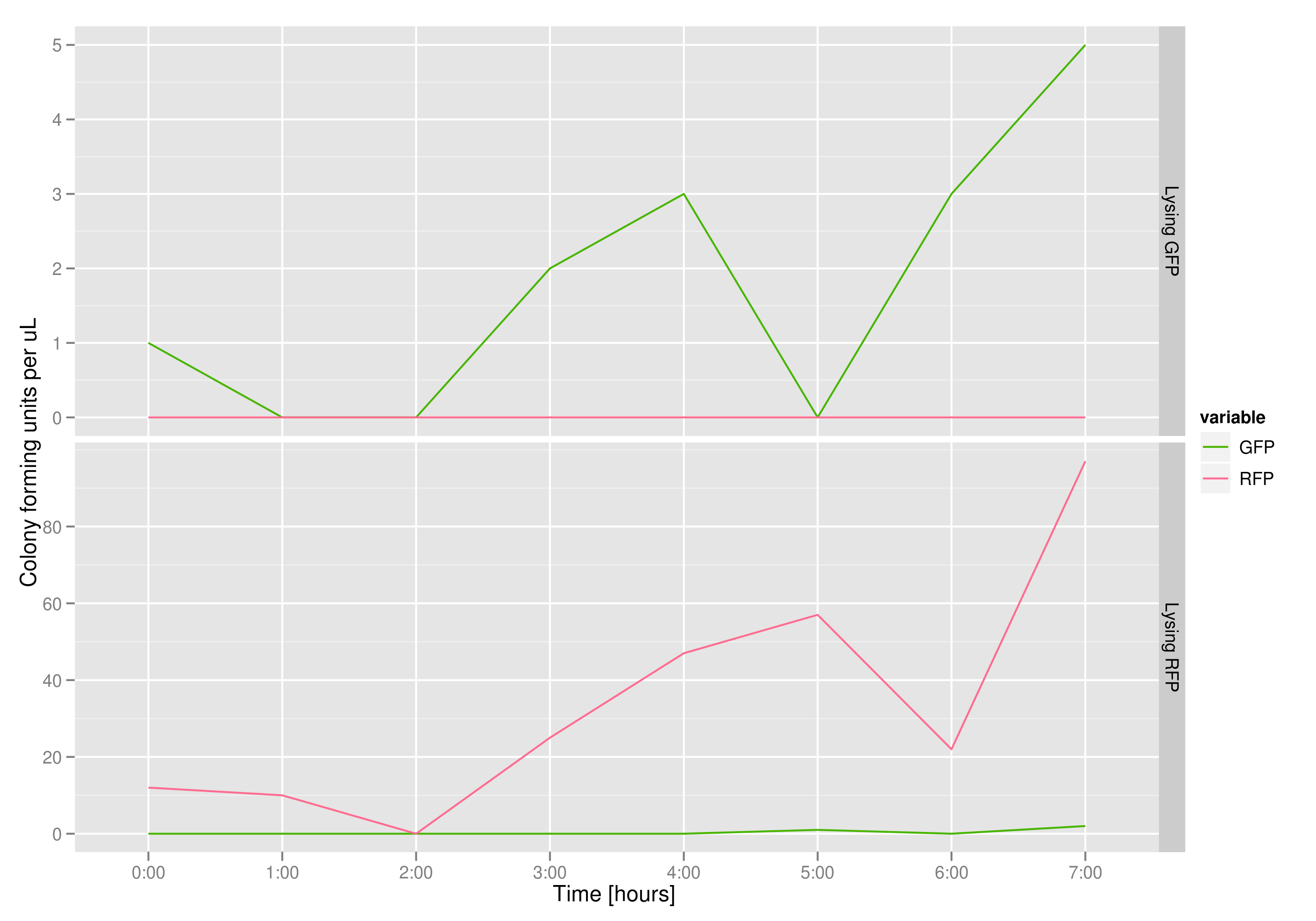
To that end, we again set up an experiment involving two flasks but this time each flask contains two types of cells. For one flask, one culture is a co-transformation of a lysis plasmid (C2) and a RFP-containing plasmid and the other is a co-transformation of a negative control plasmid (C11) with a GFP-containing plasmid. In the other flask, the reverse is true: lysis (C2) is with GFP and negative control (C11) is with RFP. This co-culture experiment was prepared by mixing equal amounts of cells from an overnight culture in one big flask. After a 1h culture, we induced lysis with 500 µM IPTG.
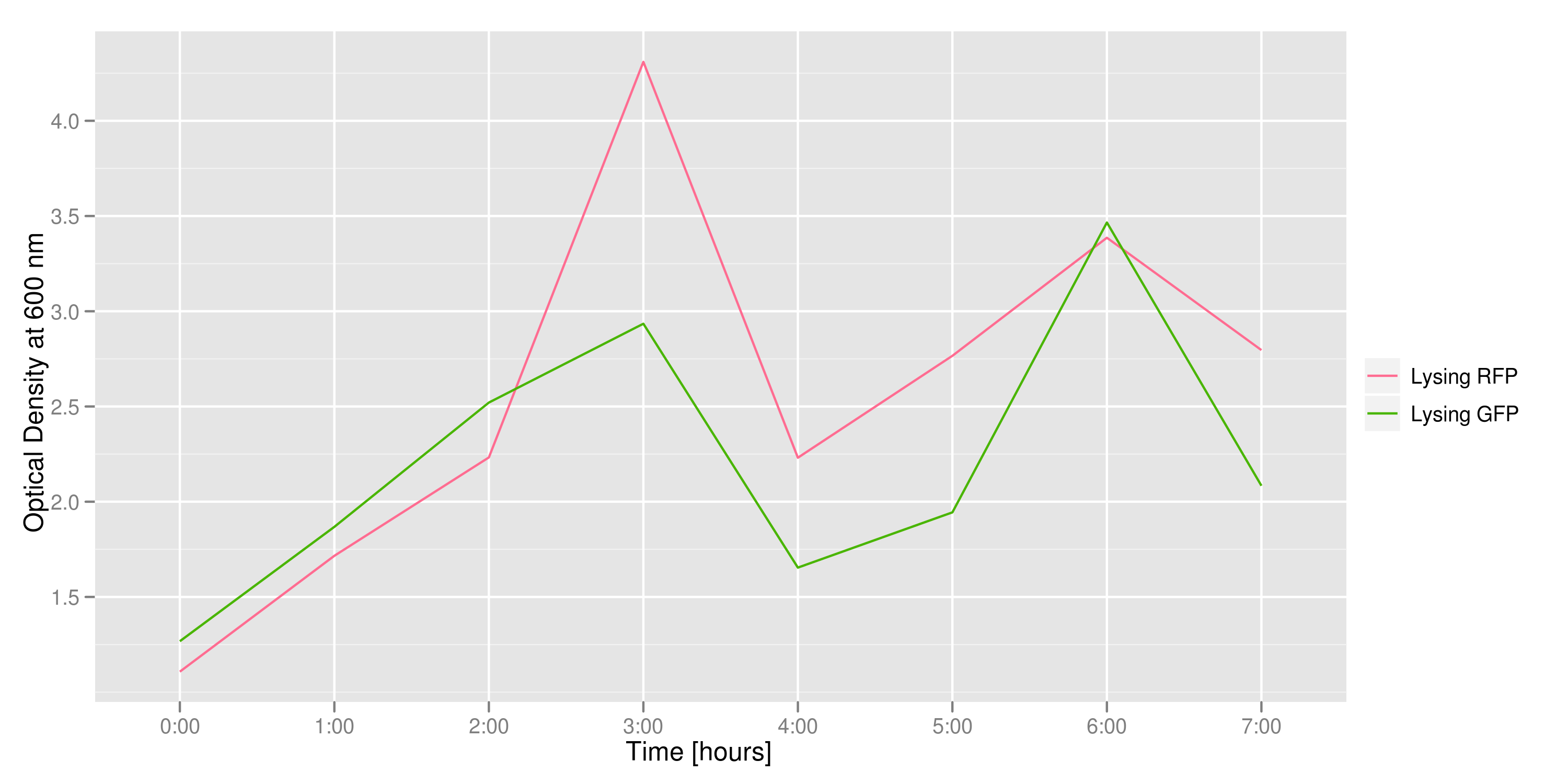
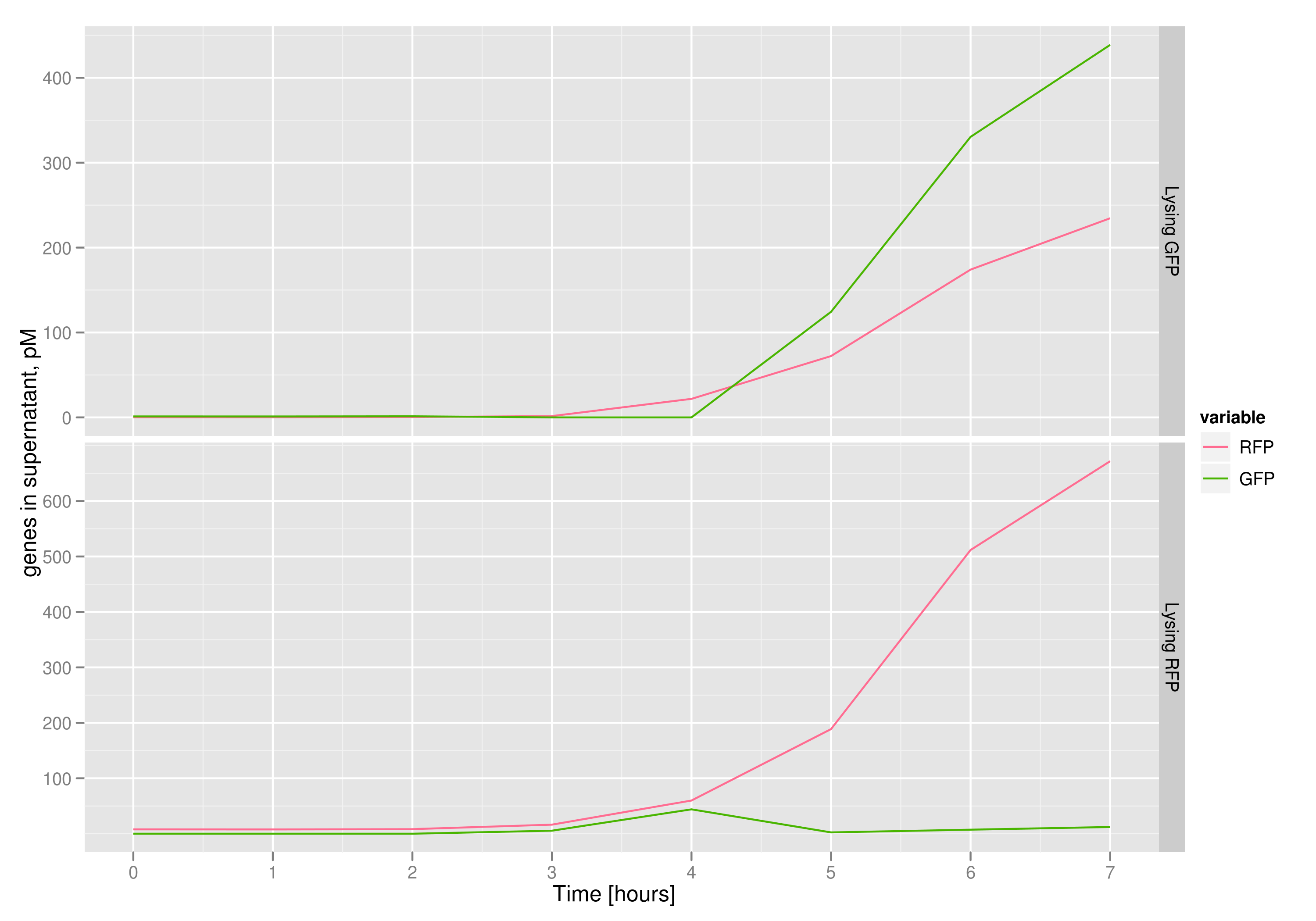
We measured optical density and took two supernatant samples every hour. One supernatant sample was centrifuged and sterile filtered to remove cell debris. It would be used for qPCR and transformation -- our two methods for quantifying the amount of recovered plasmid DNA. The other sample was used to keep track of fluorescence in the cells over the course of the experiment: the non-lysed cells were expected to fluoresce according to the appropriate gene (RFP or GFP, depending on the culture).
Results
 "
"