Team:Paris Bettencourt/Modeling/tRNA diffusion
From 2011.igem.org
| Line 210: | Line 210: | ||
<h4>Limits</h4> | <h4>Limits</h4> | ||
| - | + | </html> | |
[[File:TRNA_scheme.png|thumb|center|upright=3.0|tRNA Amber genetic design]] | [[File:TRNA_scheme.png|thumb|center|upright=3.0|tRNA Amber genetic design]] | ||
Revision as of 08:52, 8 September 2011

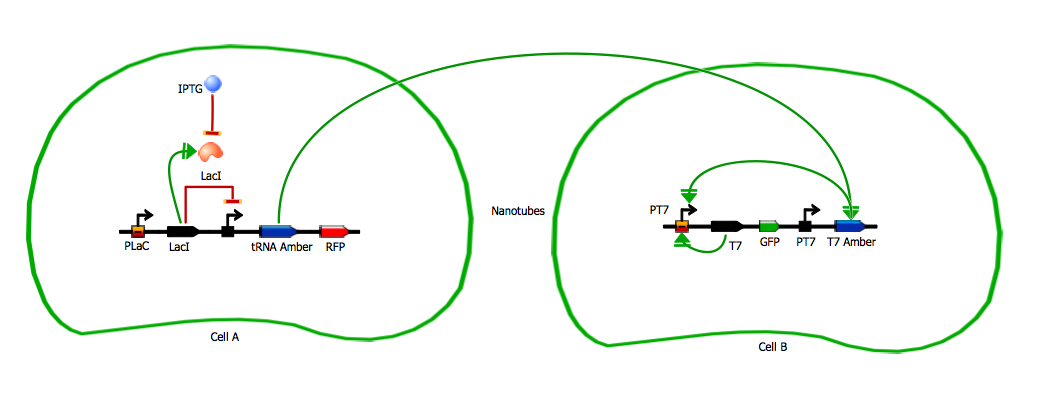
Model for tRNA amber diffusion system
Summary
Design
Model
LacI
We use LacI as a repressor for the emitter gene construct. LacI repression can be cancelled by IPTG. This way we can induce production of RFP and tRNA amber by adding IPTG on the cells.


Inactivated LacI can not repress the pLAC promoter anymore. Note that we consider that the reaction between IPTG and LacI fires without any delay. This assumption is justified by the fact that this reaction is much faster than any other in our gene network.
Emitter gene construct - comS
The emitter gene construct is modeled by the following equations:

The reporter for the emitter gene construct (RFP) is modeled by the following equations:

Receiver gene construct and amplification loop - T7 amber and T7 amplification loop
The receiver and amplification gene construct is modeled by the following equations:

The reporter for the receiver and amplification gene construct (GFP) is modeled by the following equations:

Parameters
This design relies on tRNA amber as the signaling molecule going through the nanotubes.
The parameters used in this model are:
| Parameter | Description | Value | Unit | Reference |
|---|---|---|---|---|
 |
Active LacI concentration (LacI which is not inactivated by IPTG) | NA | molecules per cell |
Notation convention |
 |
IPTG concentration | NA | molecules per cell |
Notation convention |
 |
Inactived LacI concentration | NA | molecules per cell |
Notation convention |
 |
Total LacI concentration | TBD | molecules per cell |
Steady state for equation |
 |
T7 RNA polymerase (emitter, T7') concentration | NA | molecules per cell |
Notation convention |
 |
mRNA associated with T7' concentration | NA | molecules per cell |
Notation convention |
 |
T7 RNA polymerase (auto-amplification, T7'') concentration | NA | molecules per cell |
Notation convention |
 |
mRNA associated with T7'' concentration | NA | molecules per cell |
Notation convention |
 |
Maximal production rate of pVeg promoter (constitutive) | ??? | molecules.s-1 or pops |
Estimated |
 |
Maximal production rate of pLac promoter | 0.02 | molecules.s-1 or pops |
Estimated |
 |
Maximal production rate of pT7 promoter | 0.02 | molecules.s-1 or pops |
Estimated |
 |
Dissociation constant for IPTG to LacI | 1200 | molecules per cell |
Aberdeen 2009 wiki |
 |
Dissociation constant for LacI to LacO (pLac) | 700 | molecules per cell |
Aberdeen 2009 wiki |
 |
Dissociation constant for T7 RNA polymerase to pT7 | 3 | molecules per cell |
Estimated ADD EXPLANATION |
 |
Translation rate of proteins | 1 | s-1 | Estimated ADD EXPLANATION |
 |
Dilution rate in exponential phase | 2.88x10-4 | s-1 | Calculated with a 40 min generation time. See explanation |
 |
Degradation rate of mRNA | 2.88x10-3 | s-1 | Uri Alon (To Be Confirmed) |
 |
Delay due tT7 RNA polymerase production and maturation | 300 | s | http://mol-biol4masters.masters.grkraj.org/html/Prokaryotic_DNA_Replication13-T7_Phage_DNA_Replication.htm |
 |
Delay due to mRNA production | 30 | s | http://bionumbers.hms.harvard.edu/bionumber.aspx?s=y&id=104902&ver=5&hlid=58815 2kb/(50b/s) --> approximation: all our contructs are around 2kb |
Results & discussions
Limits
The amber suppressor tRNA diffusion. The idea of the system is to pass tRNA amber molecules through the nanotubes. At every moment of time in the receiver cell there is a certain amount of transcribed mRNA-T7 among the others mRNA. The behavior of tRNA amber that arrived in a receiver cell is random, so in order to describe its interaction with mRNA-T7 and its further translation we can reason in terms of probability.
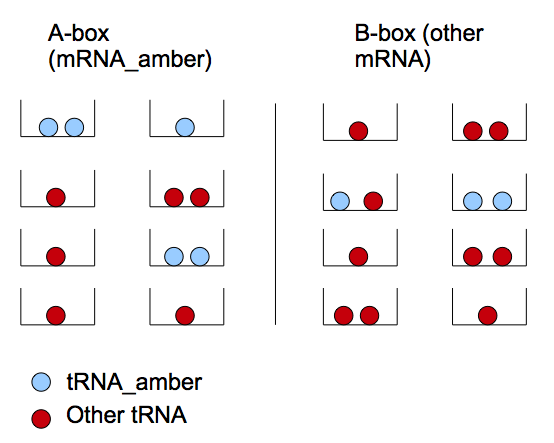
We can reason in two steps : first a tRNA amber molecule gets close to a mRNA molecule. Then, it binds it's anti-codon with a codon of the mRNA. This reasoning is similar to the problem of boxes and balls. There are two types of boxes: 'a' of the first type and 'b' of the second (which corresponds to the set of mRNA-T7 and mRNA-non-T7), and there are 't' balls(tRNA amber). All the balls are randomly distributed in the boxes. If there are two or more balls in some box of the first type (two or more tRNA amber per mRNA-T7) then a T7 molecule will be produced with a chance P_0.
We have defined two models for this system which both rely on the following assumptions :
- Each mRNA is defined as a 'box'
- All the tRNA molecules are uniformly distributed in the boxes.
- The number of tRNA_amber diffused through the nanotubes is much more smaller than the one of the mRNA. Thus the chance that three or more tRNA amber will "find" one mRNA-T7 is negligible comparing to the one of two tRNA amber (finding a mRNA-T7). In our model we will consider that at one moment of time each mRNA interacts with 0, 1 or 2 tRNA ambers.
- The tRNA_amber placed in a correct box are always used
We have defined two models for this system which both rely on the following assumptions :
- Each mRNA is defined as a 'box'
- All the tRNA molecules are uniformly distributed in the boxes.
- The number of tRNA_amber diffused through the nanotubes is much more smaller than the one of the mRNA. Thus the chance that three or more tRNA amber will "find" one mRNA-T7 is negligible comparing to the one of two tRNA amber (finding a mRNA-T7). In our model we will consider that at one moment of time each mRNA interacts with 0, 1 or 2 tRNA ambers.
- The tRNA_amber placed in a correct box are always used
Let's define two types of boxes: mRNA_amber ('A') and other type of mRNA ('B'). We note the mRNA_amber producing T7 as mRNA*_amber. The latter appears if we have two tRNA_amber in one box. This model treats the repartition of tRNA_amber in the different boxes.
We note P(x) the probability of having x A-boxes containing two tRNA_amber. Thus P(x=1) corresponds to the probability of finding a couple of tRNA_amber in an A-box, thus to produce x T7 molecules. 't' is the number of tRNA_amber in the cell.
We note:
- P(x=1)= (probability that 2 balls choose A-boxes) * (probability that these 2 balls choose the same A-box) + (probability that 3 balls choose A boxes) * (probability that 2 balls out of 3 choose the same A-box and the third doesn't) + ... + (probability that t balls choose an A-box) * (probability that 2 of these t balls choose the same A-box and the rest don't).
- P(x=2)= (probability that 4 balls choose A-boxes) * (probability that these 4 balls choose 2 A-boxes, one A-box per pair of balls) + (probability that 5 balls choose A-boxes) * (probability that 4 balls out of 5 choose 2 A-boxes, one A-box per pair of balls and the fifth doesn't) + ... + (probability that t balls choose A-boxes) * (probability that 4 out of these t balls choose [t/2] A-boxes, one A-box per pair of balls and the rest don't).
- ...
- P(x=i)= (probability that 2i balls choose A-boxes) * (probability that these 2i balls choose i A-boxes, one A-box per pair of balls) + (probability that (2i + 1) balls choose A-boxes) * (probability that 2i balls out of (2i + 1) choose i A-boxes, one A-box per pair of balls and the rest don't) + ... + (probability that t balls choose A-boxes) * (probability that 2i out of these t balls choose [t/2] A-boxes, one A-box per pair of balls and the rest don't).
Hence:
 "
"