Team:EPF-Lausanne/Notebook/July2011
From 2011.igem.org
| Line 61: | Line 61: | ||
Also, we have some light smaller bands in colonies 1 and 4, indicating that the miniprep is perhaps not so pure. | Also, we have some light smaller bands in colonies 1 and 4, indicating that the miniprep is perhaps not so pure. | ||
| - | |||
== Friday, 8th of July 2011 == | == Friday, 8th of July 2011 == | ||
| Line 70: | Line 69: | ||
The fluorescence is insufficiently clear for accurate conclusions, therefore the PCR and gel will be repeated with different concentrations on Monday. | The fluorescence is insufficiently clear for accurate conclusions, therefore the PCR and gel will be repeated with different concentrations on Monday. | ||
| + | |||
| + | |||
| + | {{:Team:EPF-Lausanne/Templates/Footer}} | ||
Revision as of 16:50, 8 July 2011
Notebook: July 2011
Contents |
Friday, 1st of July 2011
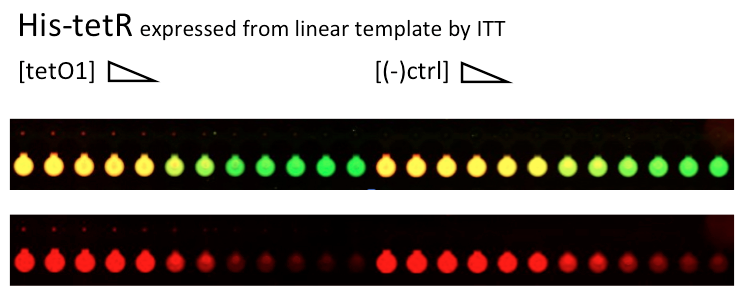
Alina and Lilia did MITOMI on His-tetR expressed from linear template, it was loaded on chip in ITT expression mix. DNA was spotted on June 29 in different concentrations for both: consensus (tetO1) sequence and a random sequence ((-)control).
Images: Green fluorescence comes from green-lysine and red fluorescence comes from DNA (cy-5 labeled).
Although we don't observe much protein bound to the anti His-tag antibody, we can still see that it does bound TetO1 sequence and did not bind any (-)control sequence. Some possible reasons for low protein fluorescence under the buttons: the expression yield from the linear template with T7 promoter might be too low, the His-tag to antibody binding is not strong enough or the concentration of antibody is too low.
Friday, 1st of July 2011
Alina and Lilia amplified the His-tetR linear template. After PCR sample was loaded on 1% agarose gel with Rad Safe, “1Kb Plus DNA Ladder” was used.
We also prepared about thirty LB-agar plates with ampicilin, they are in the BM fridge.
Tween experiment
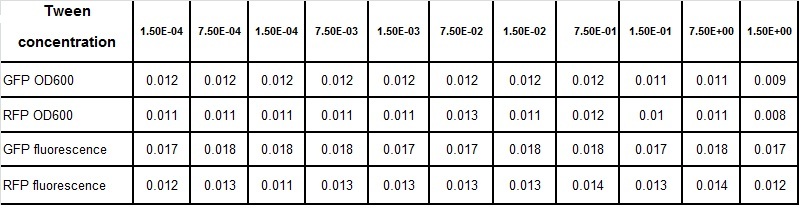
Clara and Henrike tested the right dose of Tween 20 that can be used for the "chemostat" experiment without affecting the cell growth. Tween is a detergent that would prevent the cells from sticking and thus clogging the microfluidic channels. We tested 12 different dilutions of Tween 12 both for GFP and RFP e-coli which can be potentially used for the "chemostat" experiment. We read the OD(absorption @600nm) and fluorescence every 10 minutes, for 18 hours at 37°C.
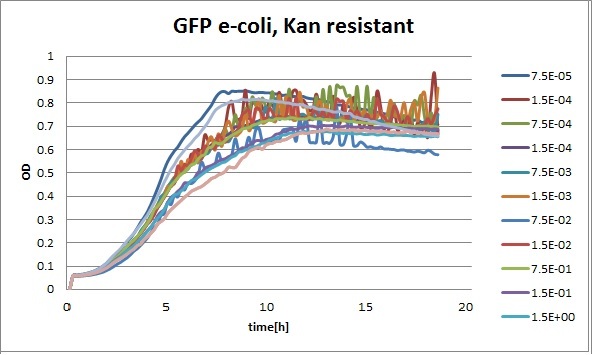
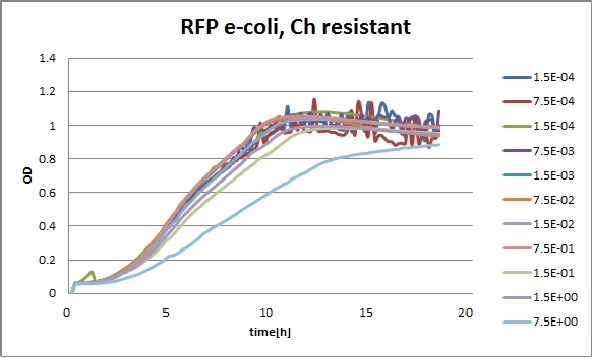
Growth curves for all the different concentrations vs time
The legend shows the percentage of tween 20 that was added to the medium.
- We can observe that the growth factor is smaller for Chloramphenicol resistant e-coli due to the fact that Chloramphenicol is stronger than Kanamycin.
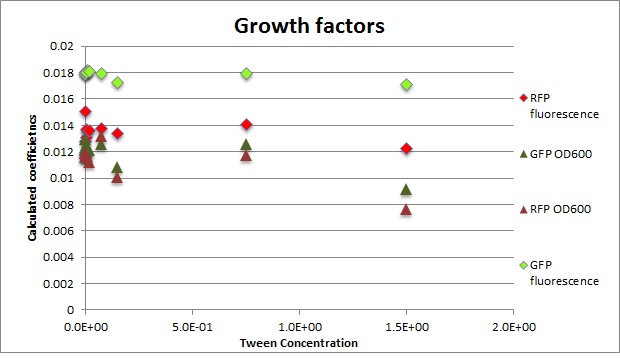
Calculated growth factor for the previous curves(Lilia)
Growth factor
We can observe that tween did not have a major effect on the cells' growth.
I suggest we use 0.075% tween in media for the experiments
Thursday, 7th of July 2011
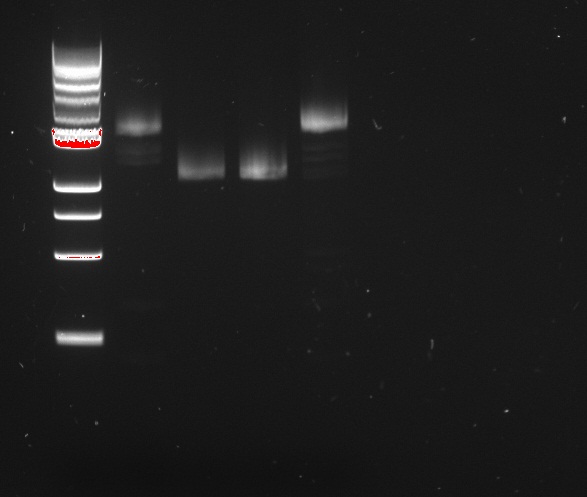
Vincent and Nadine made a digestion on the minipreps from the 4 colonies that were transformed with the Gibson-extended J61002 plasmid. If Gibson was successful, only Pst1 will cut and we should have one band of 3094 bp. If Gibson didn't work, then Pst1 and Spe1 will cut and we should have 2 bands of 890 and 2060 bp. The digestion was made according to last year's protocol, with 1.5 hour incubation. The ladder on the gel is 1kb.
Colonies 1 and 4 have the expected band over 3kb, which is coherent with the observation made that they did express RFP. But colonies 2 and 3 have only one band... It is likely that in these cells the plasmid recombined in a shorter way, without RFP but with the resistance; otherwise the plasmid sequence is not what we think. We'll check tomorrow on the J61002 plasmid if we do get 2 bands on the starting plasmid. Also, we have some light smaller bands in colonies 1 and 4, indicating that the miniprep is perhaps not so pure.
Friday, 8th of July 2011
We ran the PCR on the tetR linear template, using Clara's primers for site-specific mutagenesis.
Six PCRs were run. The first amplifies the common sequence of the mutants: everything up to the mutated sites. The six other reactions amplified the second half of the gene, inducing specific mutations in tetR.
The fluorescence is insufficiently clear for accurate conclusions, therefore the PCR and gel will be repeated with different concentrations on Monday.
 "
"