Team:Washington/Magnetosomes/Magnet Results
From 2011.igem.org
(→A table of cloning efficiency) |
|||
| Line 102: | Line 102: | ||
Please see our <i>mamAB</i> genes description [https://2011.igem.org/Team:Washington/Magnetosomes/mamDescriptions page]. | Please see our <i>mamAB</i> genes description [https://2011.igem.org/Team:Washington/Magnetosomes/mamDescriptions page]. | ||
| - | ==''' | + | =='''Table of computationally predicted cloning efficiencies ''' == |
| - | + | The following results and analyses are preliminary. | |
| - | The following is a list of computationally predicted cloning efficiencies for each gene we cloned, from the RBS tool | + | The following is a list of computationally predicted cloning efficiencies for each gene we cloned, from the publicly available RBS tool. |
{| class="wikitable" | {| class="wikitable" | ||
Revision as of 22:12, 21 October 2011
What’s in the Magnetosome Toolkit?
- A set of 10 gene clusters from the essential mamAB operon of strain AMB-1
- Our favorite genes as translational fusions with superfolder gfp in pGA vectors
- A table compiling individual gene functions from our literature search
- A table of cloning efficiency
Superfolder GFP-magnetosome gene protein fusions
The two genes we characterized as fusions with superfolder GFP are mamK and mamI. They each perform core functions of magnetosome formation. MamK is a bacterial actin-like cytoskeleton protein required for proper alignment of the magnetosomes in a chain. MamI is a membrane-localized protein required for magnetosome vesicle formation that has also been shown to localize on the MamK filament. For more information, see the mamAB description page. Using our two genes of interest, we created C-terminal sfGFP fusions so we could track the localization of each gene separately within E.coli.
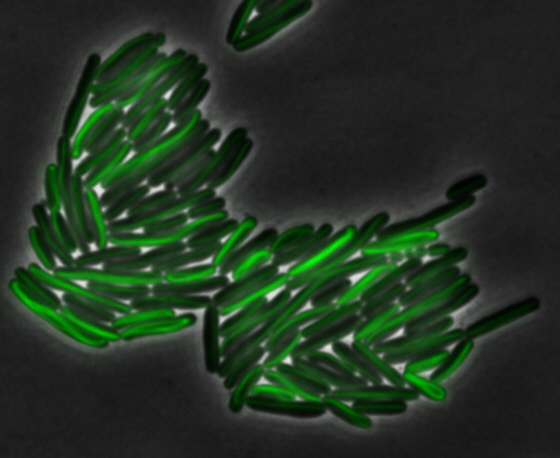
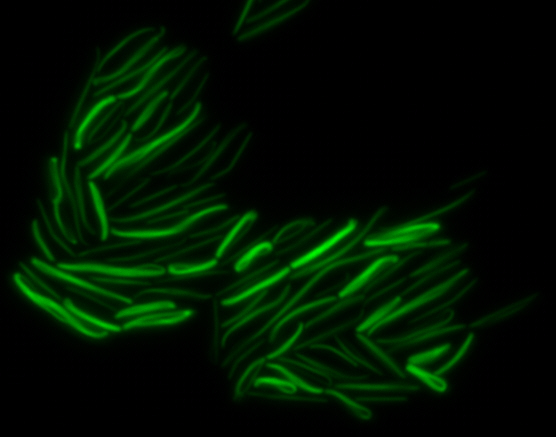
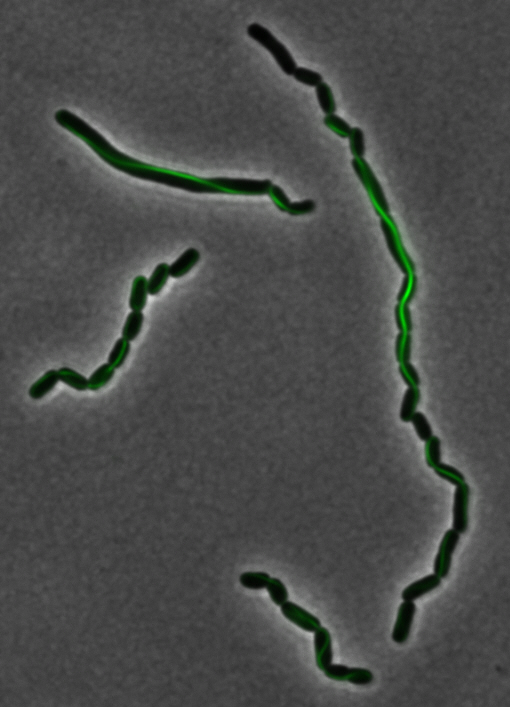
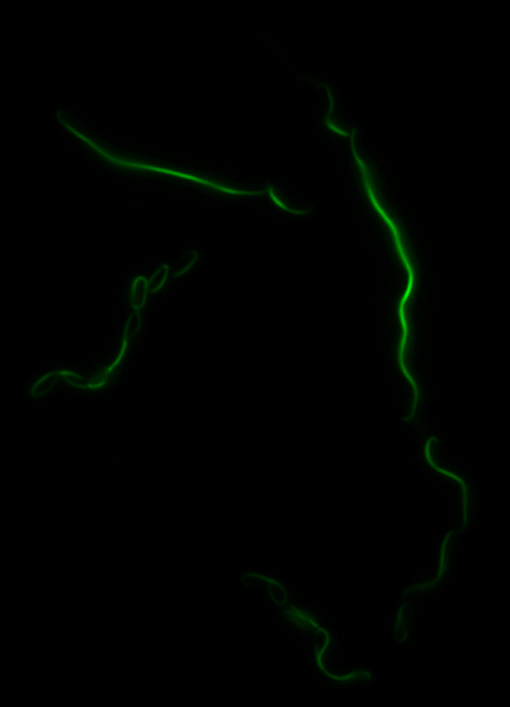
sfGFP-MamK: Scaffold formation
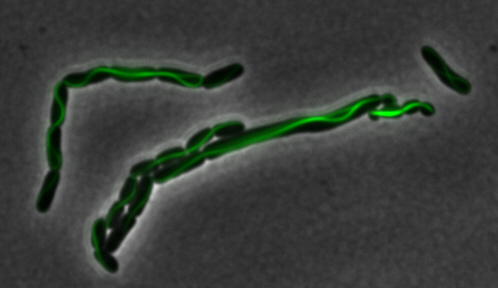
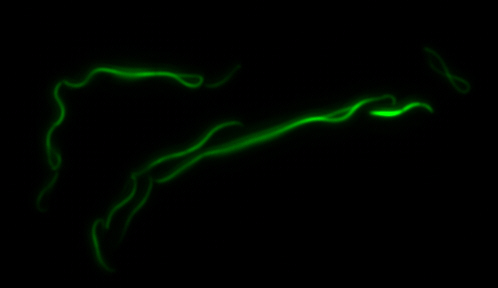
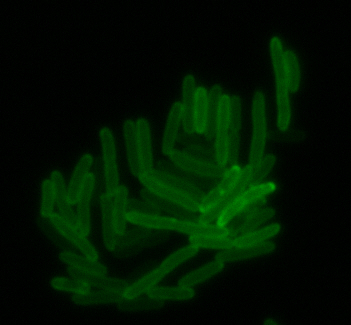
The results we obtained with our sfGFP fusions inside E. coli were comparable to those done through other studies in the host organism Magnetospirillum magneticum. Within AMB-1, MamK is a filament which runs along the long axis of the bacteria. In the our images of sfGFP-MamK, scaffold-like structures can be clearly seen running through the length of most of the cells, in some cases looping back within a single cell for "figure-8" shaped filaments. In other cases, the filaments seem to prevent the cells from dividing properly, resulting in long chains of E. coli. In our experimental result, there was an over- expression of mamK which connected the E.coli cells together.
Here is a video showing the filaments connecting the growing E coli:
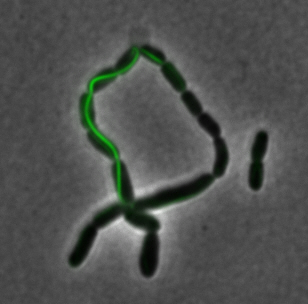
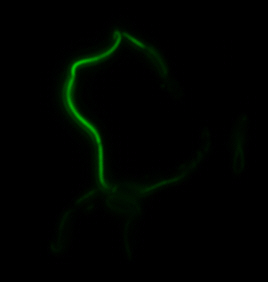
sfGFP-MamI: Membrane localization
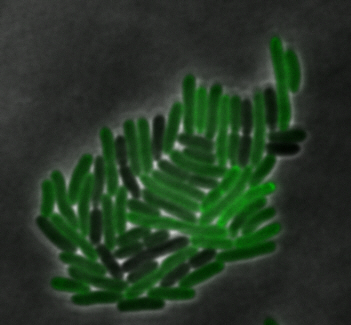
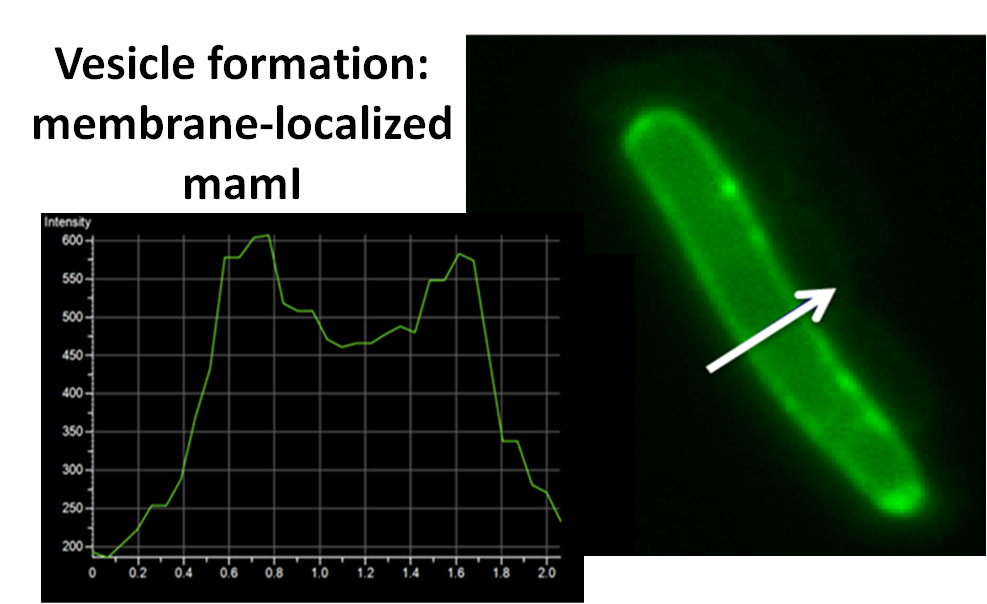
For mamI, the gene product localizes to the cell membrane, consistent with its known role in inner membrane vesicle invagination. The membrane localization is easily seen by the fluorescence profile analysis seen on the panel on the right. The graph shows that the fluorescence levels peak near the cell membrane and decrease to a minimum in the middle of the cytoplasm.
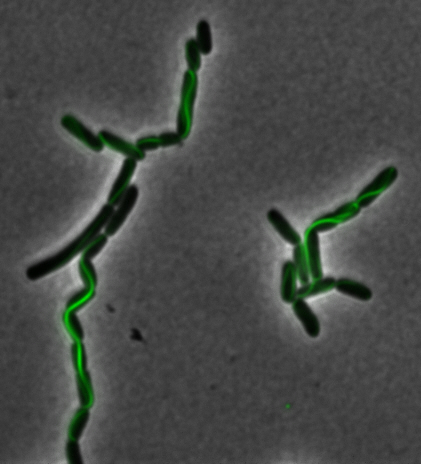
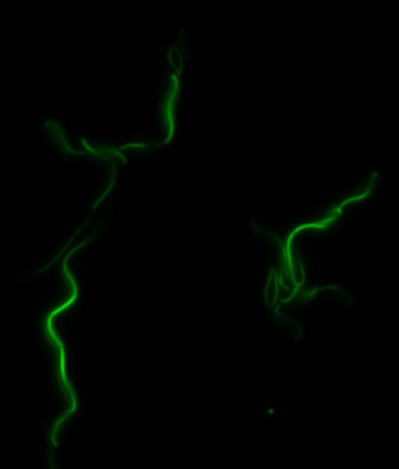
Construction of the R5 region of the Magnetosome Island in E.coli
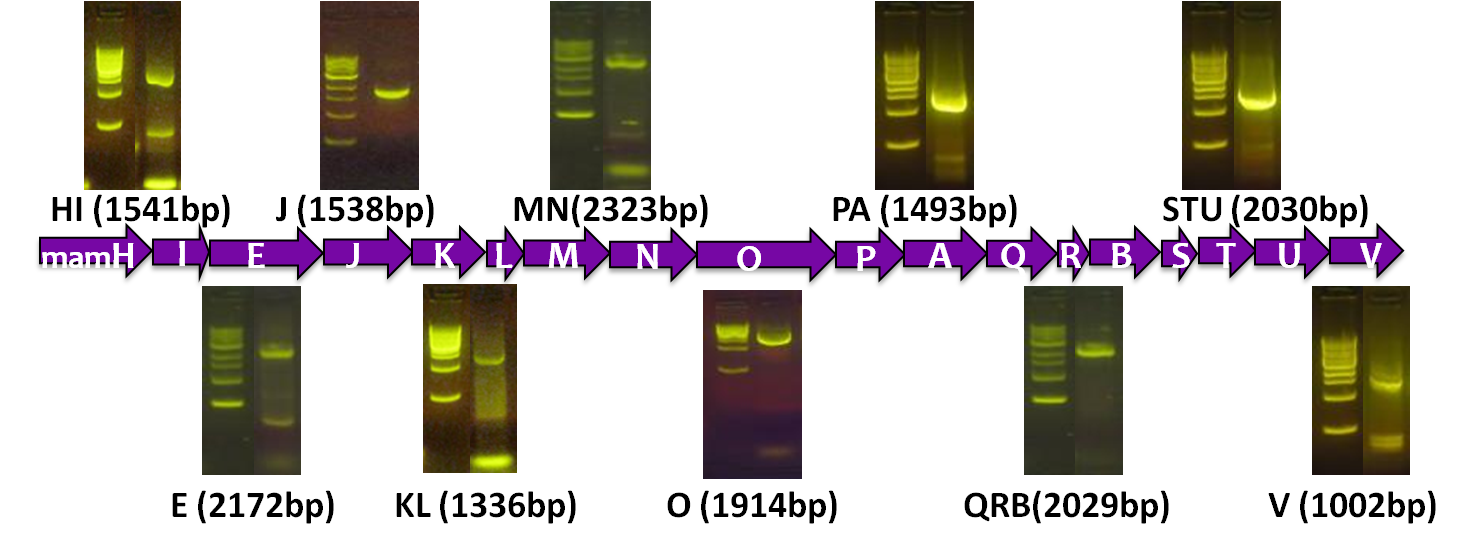
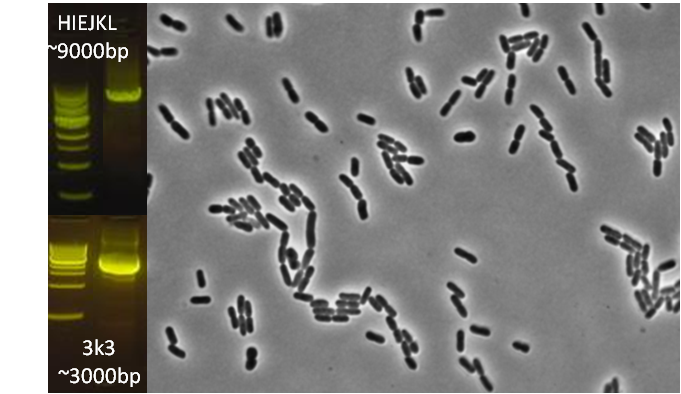
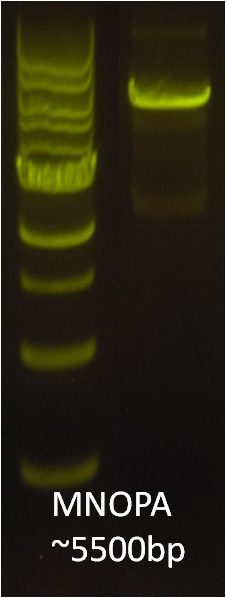
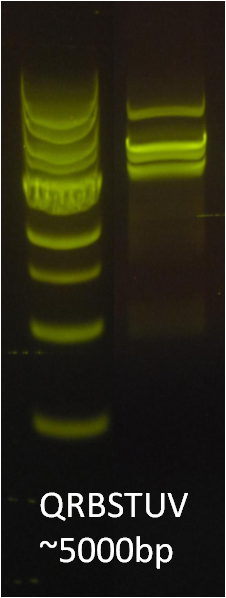
After verifying that the construction of the sfGFP-MamK scaffold worked as expected, we proceeded to create a full assembly of the mamAB operon by building three super-assemblies: mamHIEJKL, mamMNOPA, and mamQRBSTUV. The PCR products of these intermediate assemblies are shown below. The mamHIEJKL and mamQRBSTUV have been partially sequence-confirmed, and we are currently working on designing primers to fill in the gap sequences. Despite these gaps, when cells with the mamHIEJKL construct were imaged, they appeared to be forming chains.
 :
:
A set of the 18 genes from the mamAB operon essential for magnetosome formation
Before piecing together the 16 kb genome of the mamAB gene cluster within the magnetosome island (MAI), we extracted out the genes in the following groups:
| Gene groups | Length (bp) |
|---|---|
| mamHI | 1541 |
| mamE | 2172 |
| mamJ | 1538 |
| mamKL | 1336 |
| mamMN | 2323 |
| mamO | 1914 |
| mamPA | 1493 |
| mamQRB | 2029 |
| mamSTU | 2030 |
| mamV | 1002 |
A table of individual gene functions
Please see our mamAB genes description page.
Table of computationally predicted cloning efficiencies
The following results and analyses are preliminary.
The following is a list of computationally predicted cloning efficiencies for each gene we cloned, from the publicly available RBS tool.
| mam gene | AMB-1 | E.Coli |
|---|---|---|
| mamH | 10821.63 | 10821.63 |
| mamI | 88.93 | 88.93 |
| mamE | 1.59 | 1.62 |
| mamJ | 105.9 | 105.9 |
| mamK | 1056.92 | 1056.92 |
| mamL | 584.21 | 584.21 |
| mamM | 930.1 | 930.1 |
| mamN | 413.73 | 230.48 |
| mamO | 138.73 | 138.73 |
| mamP | 198.85 | 198.85 |
| mamA | 14.29 | 14.29 |
| mamQ | 22.41 | 12.49 |
| mamR | XXX | XXX |
| mamB | 16.56 | 16.56 |
| mamS | 32.86 | 25.09 |
| mamT | 81.16 | 81.16 |
| mamU | 33.58 | 33.58 |
| mamV | 1.61 | 1.61 |
 "
"