Team:Harvard/Technology/One-Hybrid Selection System
From 2011.igem.org
(→Fine-tuning the one-hybrid selection system) |
|||
| Line 6: | Line 6: | ||
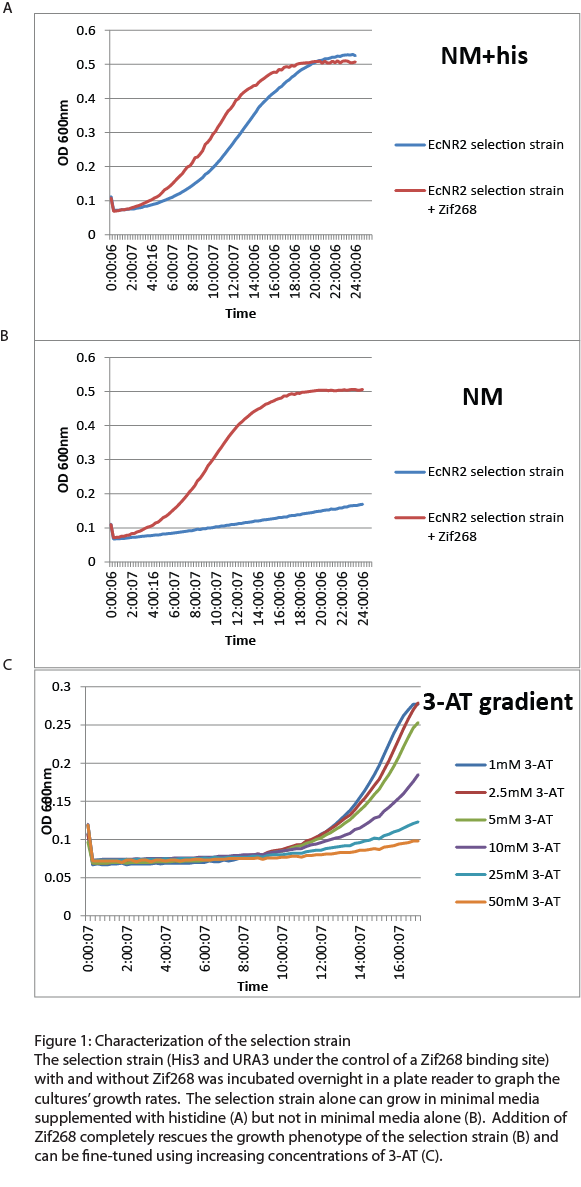
[[File: HARVhybrid1.png|left|300px|Figure 1: Characterization of the selection strain]] | [[File: HARVhybrid1.png|left|300px|Figure 1: Characterization of the selection strain]] | ||
| - | |||
| - | |||
| + | After the one-hybrid His3-URA3 strain was constructed using lambda red and MAGE, we characterized its growth phenotype in preparation for the chip-synthesized zinc fingers. Cultures with the Zif268 binding site either with and without the Zif268 protein were grown overnight in a plate reader to chart their growth under various conditions by measuring the absorbance levels at 600nm. As expected, the strain without Zif268 grew normally in complete media but failed to grow in NM media without histidine (Figure 1A). Zif268 successfully resuced the growth phenotype in NM (Figure 1B) and reached similar saturation levels to cultures grown in NM+histidine. Addition of 5-FOA killed the Zif268 cultures because of their expression of URA3 but did not have a great effect on the selection strain alone, showing that the zinc finger binding site promoter is not inherently leaky. 3-AT, the competitive inhibitor of His3, also fine-tuned selection: Zif268 cultures grew less as 3-AT concentration increased. Overall, the strain showed the proper phenotypes and successfully illustrated selection for zinc finger binding. | ||
| + | [[File: HARVhybrid2.png|right|300px|Figure 2: Simulation of chip conditions]] | ||
For the zinc fingers synthesized from the chip, the selection strain would need to be able to recognize low numbers of hits among a high level of background. Approximately 9000 different zinc fingers were designed for each binding site, and all 9000 are unlikely to successfully bind to the target, so we tested the sensitivity of the selection strain by diluting Zif268 into zinc finger plasmids that would not bind to the Zif268 site (Figure 2). Overnight growth in a plate reader showed that our selection strain was sensitive enough to detect Zif268 when diluted as low as one to one million in as high a concentration of 3-AT as 10mM. This is more than sufficient to pick out one hit among the 9000 chip-synthesized zinc fingers. | For the zinc fingers synthesized from the chip, the selection strain would need to be able to recognize low numbers of hits among a high level of background. Approximately 9000 different zinc fingers were designed for each binding site, and all 9000 are unlikely to successfully bind to the target, so we tested the sensitivity of the selection strain by diluting Zif268 into zinc finger plasmids that would not bind to the Zif268 site (Figure 2). Overnight growth in a plate reader showed that our selection strain was sensitive enough to detect Zif268 when diluted as low as one to one million in as high a concentration of 3-AT as 10mM. This is more than sufficient to pick out one hit among the 9000 chip-synthesized zinc fingers. | ||
Revision as of 19:47, 21 September 2011
Fine-tuning the one-hybrid selection system
After the one-hybrid His3-URA3 strain was constructed using lambda red and MAGE, we characterized its growth phenotype in preparation for the chip-synthesized zinc fingers. Cultures with the Zif268 binding site either with and without the Zif268 protein were grown overnight in a plate reader to chart their growth under various conditions by measuring the absorbance levels at 600nm. As expected, the strain without Zif268 grew normally in complete media but failed to grow in NM media without histidine (Figure 1A). Zif268 successfully resuced the growth phenotype in NM (Figure 1B) and reached similar saturation levels to cultures grown in NM+histidine. Addition of 5-FOA killed the Zif268 cultures because of their expression of URA3 but did not have a great effect on the selection strain alone, showing that the zinc finger binding site promoter is not inherently leaky. 3-AT, the competitive inhibitor of His3, also fine-tuned selection: Zif268 cultures grew less as 3-AT concentration increased. Overall, the strain showed the proper phenotypes and successfully illustrated selection for zinc finger binding.
For the zinc fingers synthesized from the chip, the selection strain would need to be able to recognize low numbers of hits among a high level of background. Approximately 9000 different zinc fingers were designed for each binding site, and all 9000 are unlikely to successfully bind to the target, so we tested the sensitivity of the selection strain by diluting Zif268 into zinc finger plasmids that would not bind to the Zif268 site (Figure 2). Overnight growth in a plate reader showed that our selection strain was sensitive enough to detect Zif268 when diluted as low as one to one million in as high a concentration of 3-AT as 10mM. This is more than sufficient to pick out one hit among the 9000 chip-synthesized zinc fingers.
We designed an additional one-hybrid selection system utilizing TolC, an SDS pump, under the control of a zinc finger binding site. The strain was successfully made and showed the proper phenotypic rescue when Zif268 was present, but SDS concentrations were not as titratable as 3-AT, and it was not able to recognize hits among high background levels as well as the His3-URA3 system. Due to TolC’s inferior sensitivity, we decided to transform the chip-synthesized zinc fingers only into the His3-URA3 selection strain.
 "
"