Team:EPF-Lausanne/Our Project/Summary
From 2011.igem.org
(→Characterisation) |
(→DIY Microfluidics) |
||
| (11 intermediate revisions not shown) | |||
| Line 1: | Line 1: | ||
{{:Team:EPF-Lausanne/Templates/Header|title=Results Summary}} | {{:Team:EPF-Lausanne/Templates/Header|title=Results Summary}} | ||
| + | |||
| + | This page summarizes our most important results, and is meant only as a reference. For an introduction to and overview of the project, please check out our [[Team:EPF-Lausanne|home page]]. For more information about each part of the project and more results, read the detailed sections for the [[Team:EPF-Lausanne/Our_Project/T7_promoter_variants|selection system]], [[Team:EPF-Lausanne/Our_Project/TetR_mutants|''in vitro'']] & [[Team:EPF-Lausanne/Our_Project/Reporter_Systems|''in vivo'']] characterisation, and [[Team:EPF-Lausanne/Tools/Microfluidics|DIY microfluidics]]. For the list of parts we submitted, please refer to the [[Team:EPF-Lausanne/Our_Project/Data|data page]]. | ||
== Lysis-based transcription factor selection == | == Lysis-based transcription factor selection == | ||
| Line 30: | Line 32: | ||
== Characterisation == | == Characterisation == | ||
| - | We decided to work with TetR, as a proof-of concept trascription factor characterization. We | + | We decided to work with TetR, as a proof-of concept trascription factor characterization. We characterised our mutants both ''in vitro'' and ''in vivo''. We started by testing the wild-type and went on with some of our 12 TetR mutants. The strategy we used consisted of: |
''' 1) producing interesting TetR mutants | ''' 1) producing interesting TetR mutants | ||
| - | The mutants were created by site-specific and PCR-induced mutagenesis | + | The mutants were created by site-specific and PCR-induced mutagenesis. Based on the literature, we introduced point mutations at key amino acids involved in DNA recognition, in an attempt to alter the specificity of the TetR mutants. |
''' 2) determining the binding energy landscape of each mutants | ''' 2) determining the binding energy landscape of each mutants | ||
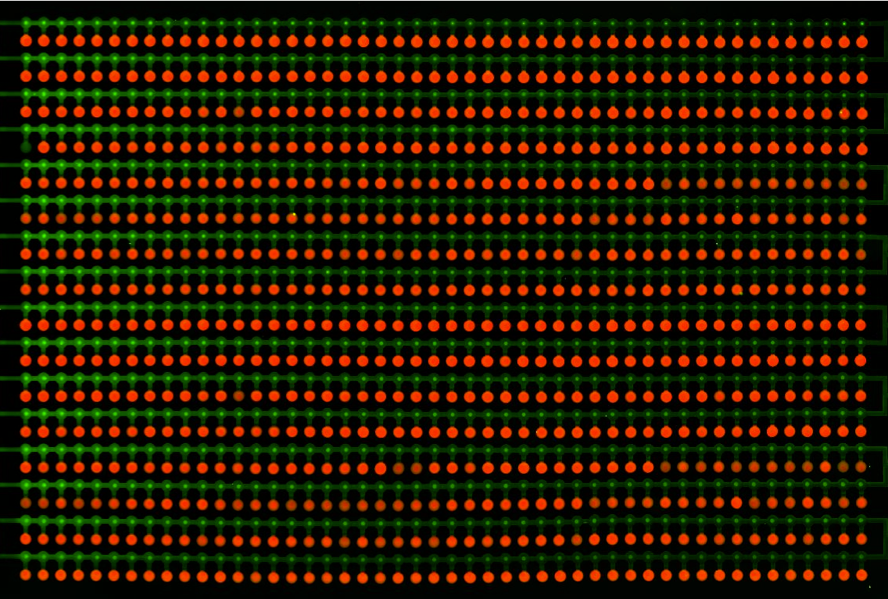
| - | This is the ''in vitro'' characterization, which was performed | + | This is the ''in vitro'' characterization, which was performed thanks to a microfluidic chip. The technique, called MITOMI, allows parallel testing of one mutant with 756 different DNA sequences. The absolute binding energy is then measured, and an enoLOGO is generated. |
[[File:EPFL-MITOMI shot.png|400px]] | [[File:EPFL-MITOMI shot.png|400px]] | ||
| Line 48: | Line 50: | ||
[[File:EPFL_Summary_without_LacI.jpg|400px ]] | [[File:EPFL_Summary_without_LacI.jpg|400px ]] | ||
| + | We ran platereader experiments for 6 of our mutants. Increasing concentrations of ATC were added to force expression of RFP, which allowed us to appreciate the repressive effect of TetR on fluorescence expression. The following graph shows wild-type TetR characterization. Increasing ATC concentrations do lead to increased fluorescence, which confirms that tetR was initially repressing RFP: | ||
| - | + | [[File:EPFL_Nadine-exp3-induction.png|600px]] | |
| - | + | ||
| - | [[File:EPFL_Nadine-exp3-induction.png| | + | |
| - | + | ||
''' 4) comparing the results of both characterization | ''' 4) comparing the results of both characterization | ||
| - | Three of our mutants plus the wild-type were tested both ''in vivo'' and ''in vitro''. The results were consistent between the two experiments. The V36F exhibits the same behaviour as the wild-type while the P39K shows no interaction to the DNA at all. The E37A triple mutant, however, has contradictory results: the MITOMI data indicate that this mutant has a change in specificity, with a consensus sequence being different from the wild-type Ptet. However, it still recognizes Ptet well, | + | Three of our mutants plus the wild-type were tested both ''in vivo'' and ''in vitro''. The results were consistent between the two experiments. The V36F exhibits the same behaviour as the wild-type while the P39K shows no interaction to the DNA at all. The E37A triple mutant, however, has contradictory results: the MITOMI data indicate that this mutant has a change in specificity, with a consensus sequence being different from the wild-type Ptet. However, it still recognizes Ptet well, meaning that there is still crosstalk with the endogenous promoter. |
[[File:EPFL_Comparison_char.png|500px]] | [[File:EPFL_Comparison_char.png|500px]] | ||
| + | We were able to change the specificity of some of our mutants; however this change was not big enough to have a fully orthogonal mutant that would not recognize the wild-type consensus sequence. | ||
| + | |||
| + | == DIY Microfluidics == | ||
| + | |||
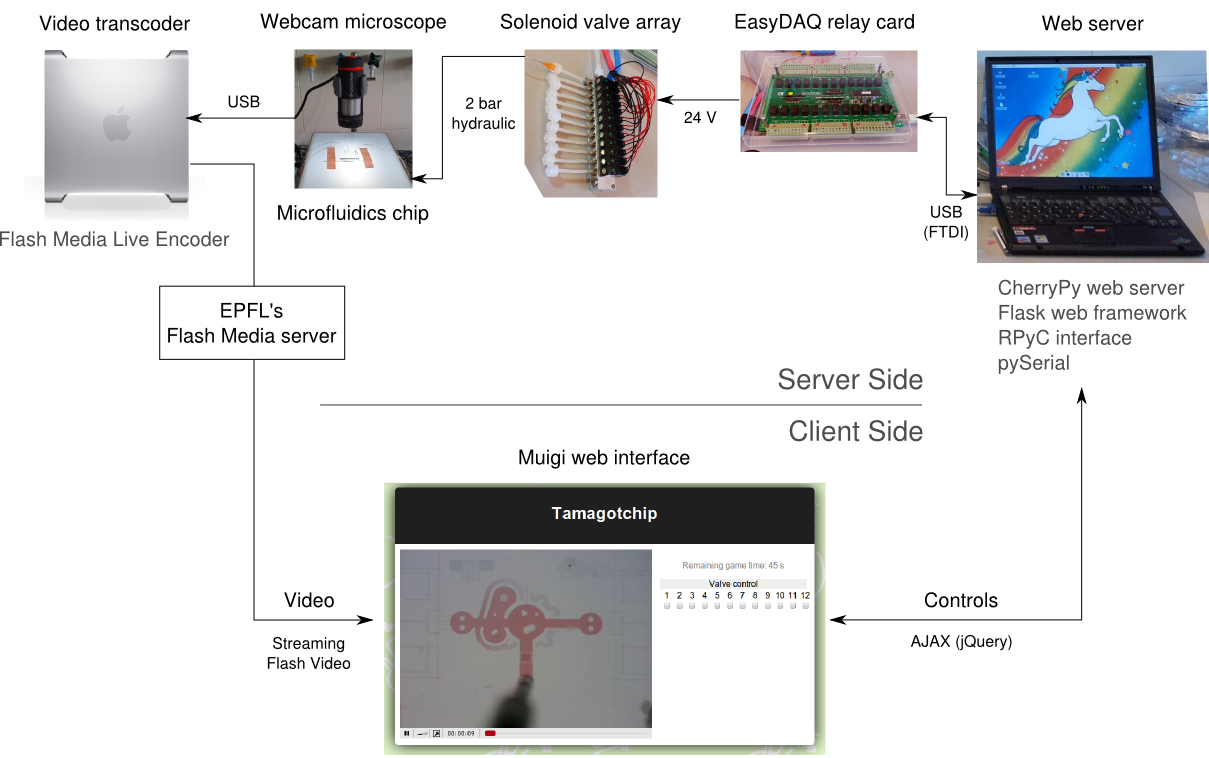
| + | [[File:EPFL-Muigi-schematic.png|thumb|300px|For less than €800, we built a microfluidics setup for 12 valves that anybody can control from their web browser.]] | ||
| + | |||
| + | In parallel to the work on the transcription factor pipeline, we built a computer-controlled microfluidic injection setup from off-the shelf components, to demonstrate that microfluidics can be used for CHF 1000 (about €800). To help future teams from other universities get started with microfluidics, we wrote on our wiki instructions to replicate the setup. | ||
| + | |||
| + | In addition to this, we programmed a web interface for the machine: from any web browser, users can view a live video feed from the microscope that images the chip, and control the valves by clicking buttons on the web page. Concurrent use is managed by a queue system. The web site was mostly used at the Jamboree in Amsterdam to illustrate the principle of microfluidics to other iGEMers, then it stayed live for most of October, allowing anybody to try the setup. The demonstration chip had channels in the shape of the iGEM logo, and was filled with a red and a blue dye to highlight flow and mixing. | ||
{{:Team:EPF-Lausanne/Templates/Footer}} | {{:Team:EPF-Lausanne/Templates/Footer}} | ||
Latest revision as of 22:59, 28 October 2011
Results Summary
This page summarizes our most important results, and is meant only as a reference. For an introduction to and overview of the project, please check out our home page. For more information about each part of the project and more results, read the detailed sections for the selection system, in vitro & in vivo characterisation, and DIY microfluidics. For the list of parts we submitted, please refer to the data page.
Lysis-based transcription factor selection
We implemented a prototype of our lysis selection system by experimentally validating four essential aspects of the strategy:
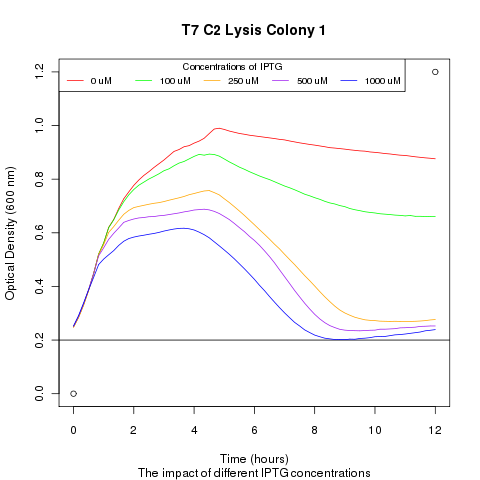
1) demonstrating that we could effectively lyse cells:
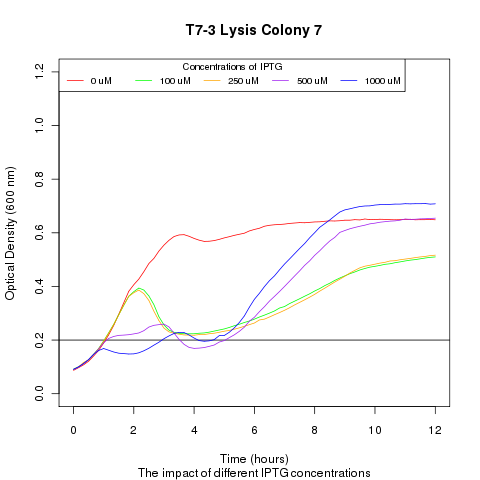
We ran platereader experiments involving growing cells to stationary growth phase and then induced with IPTG. The resulting optical density indicates that lysis occurs, with increased lysing efficiency as a function of increased IPTG concentration.
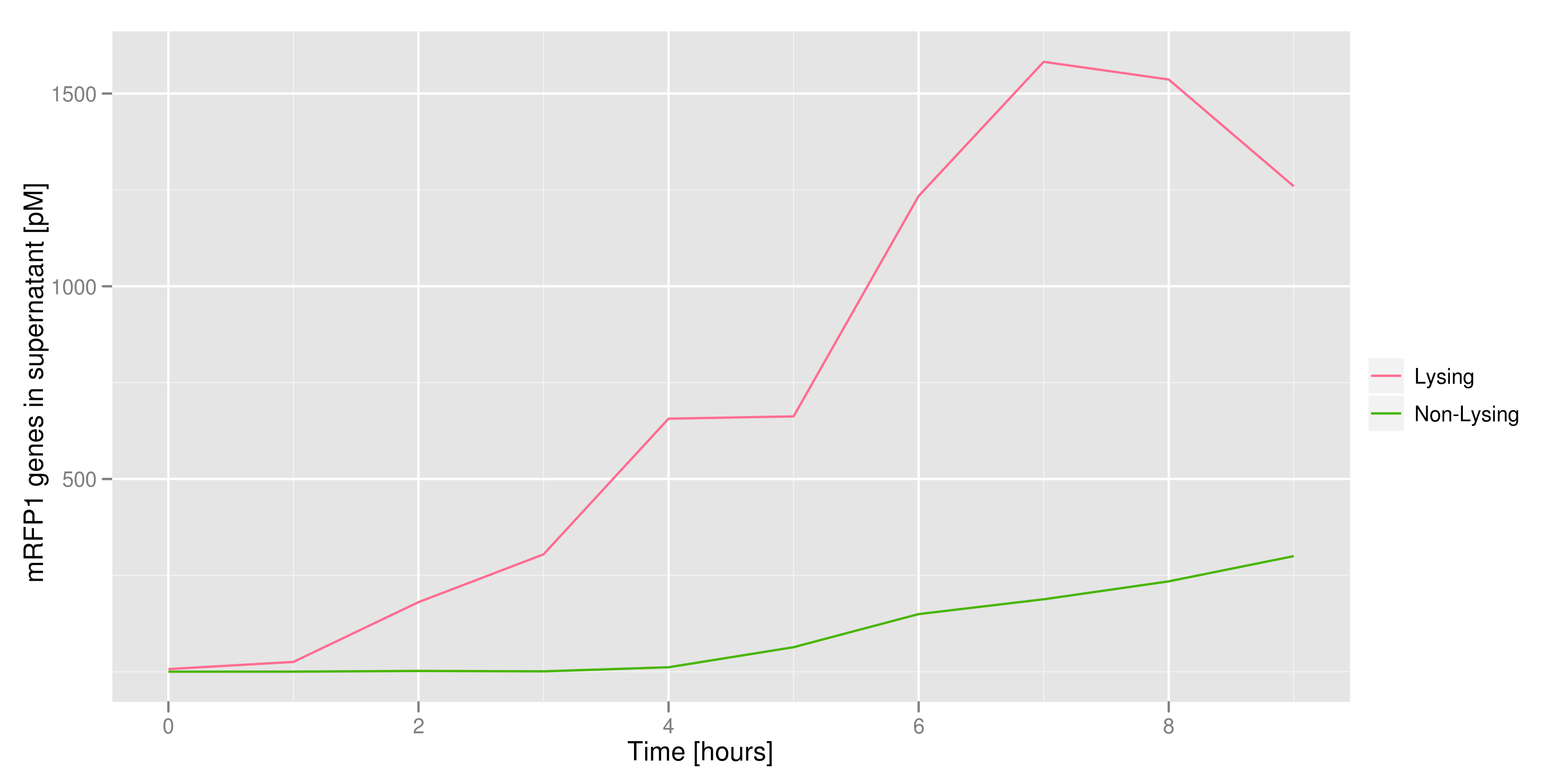
2) demonstrating that DNA could be recovered from the supernatant
We made a large culture of cells containing the lysis cassette and an RFP-containing plasmid and induced with IPTG. We collected samples of the supernatant every hour and quantified the amount of DNA in those samples by transforming the DNA and counting CFUs and by qPCR. Both quantifications revealed the same result: RFP plasmids were recovered in increasing amounts as time went on.
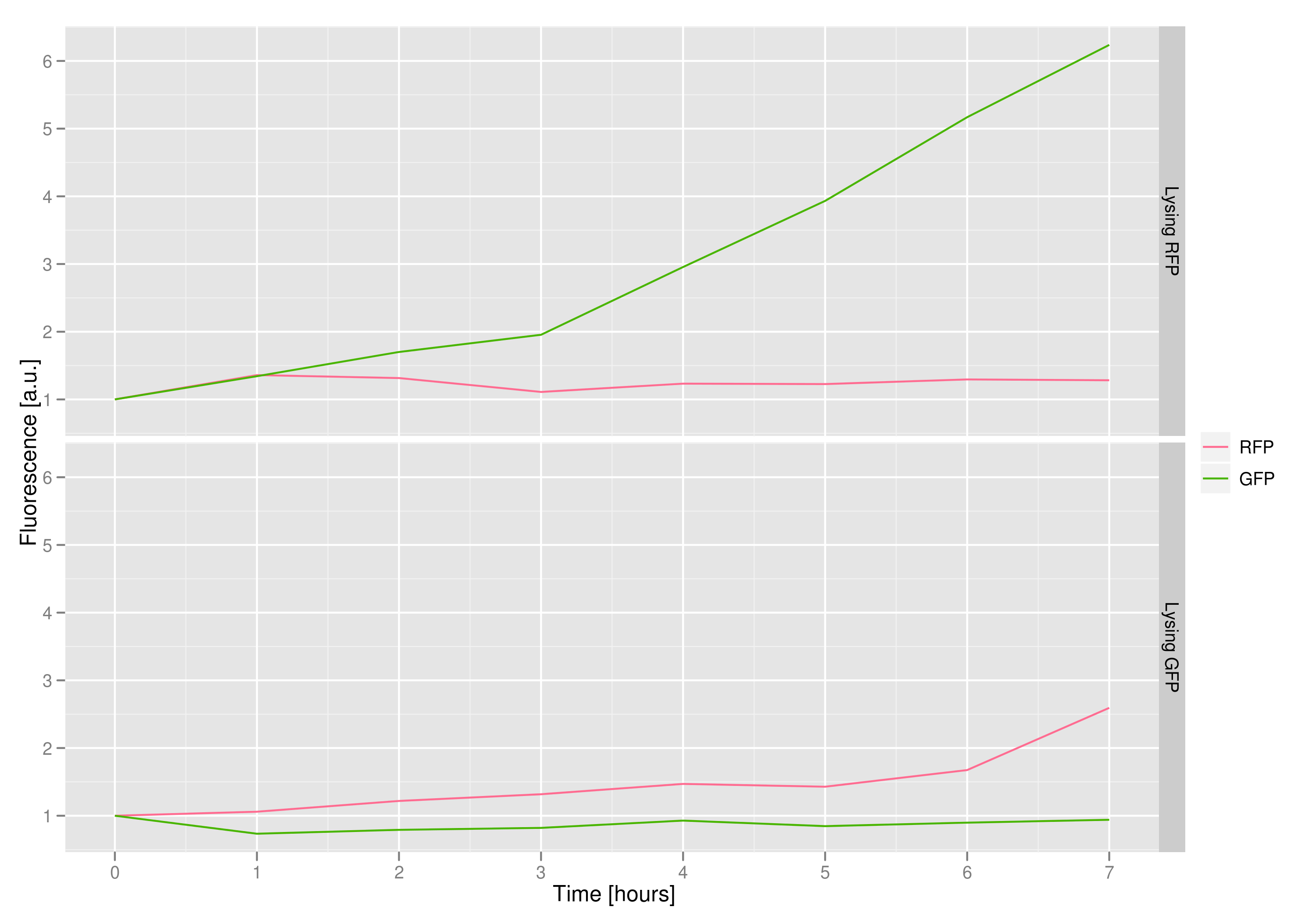
3) demonstrating that the appropriate cells (those with the desired trait or mutant DNA) would lyse, leaving the other cells intact
We made a co-culture of cells. One culture had the lysis cassette and an RFP plasmid, while the other had a GFP plasmid and no lysis cassette. Upon induction with IPTG, the qPCR and transformation methods of the previous experiment showed that RFP was recovered in large quantities and only trace quantities of GFP are detected.
4) demonstrating that promoter strength (i.e. transcription factor- DNA binding) has a direct impact on the amount and speed of lysing
We ran a platereader experiment with multiple cultures, each culture containing a plasmid with a different T7 promoter mutant driving the lysis cassette. Induction with IPTG shows that having different promoters results in varying lysis strengths and speeds. This approach with promoter mutants supplies an alternate way of showing that favorable transcription-factor mutants will lyse cells more rapidly and efficiently then their peers. The resulting supernatant will therefore contain a proportionally high number of the better TF mutant DNA, which can be recovered and sequenced.
Characterisation
We decided to work with TetR, as a proof-of concept trascription factor characterization. We characterised our mutants both in vitro and in vivo. We started by testing the wild-type and went on with some of our 12 TetR mutants. The strategy we used consisted of:
1) producing interesting TetR mutants
The mutants were created by site-specific and PCR-induced mutagenesis. Based on the literature, we introduced point mutations at key amino acids involved in DNA recognition, in an attempt to alter the specificity of the TetR mutants.
2) determining the binding energy landscape of each mutants
This is the in vitro characterization, which was performed thanks to a microfluidic chip. The technique, called MITOMI, allows parallel testing of one mutant with 756 different DNA sequences. The absolute binding energy is then measured, and an enoLOGO is generated.
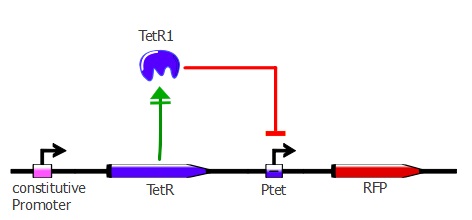
3) determining the binding of each mutant to the wild-type TetO sequence in vivo
For this characterization step, we set up a reporter construct: TetR constitutively expressed and RFP with a wild-type Ptet promoter, as you can see on the illustration below. The more RFP is expressed in the cells, the less the TetR mutant recognizes Ptet.
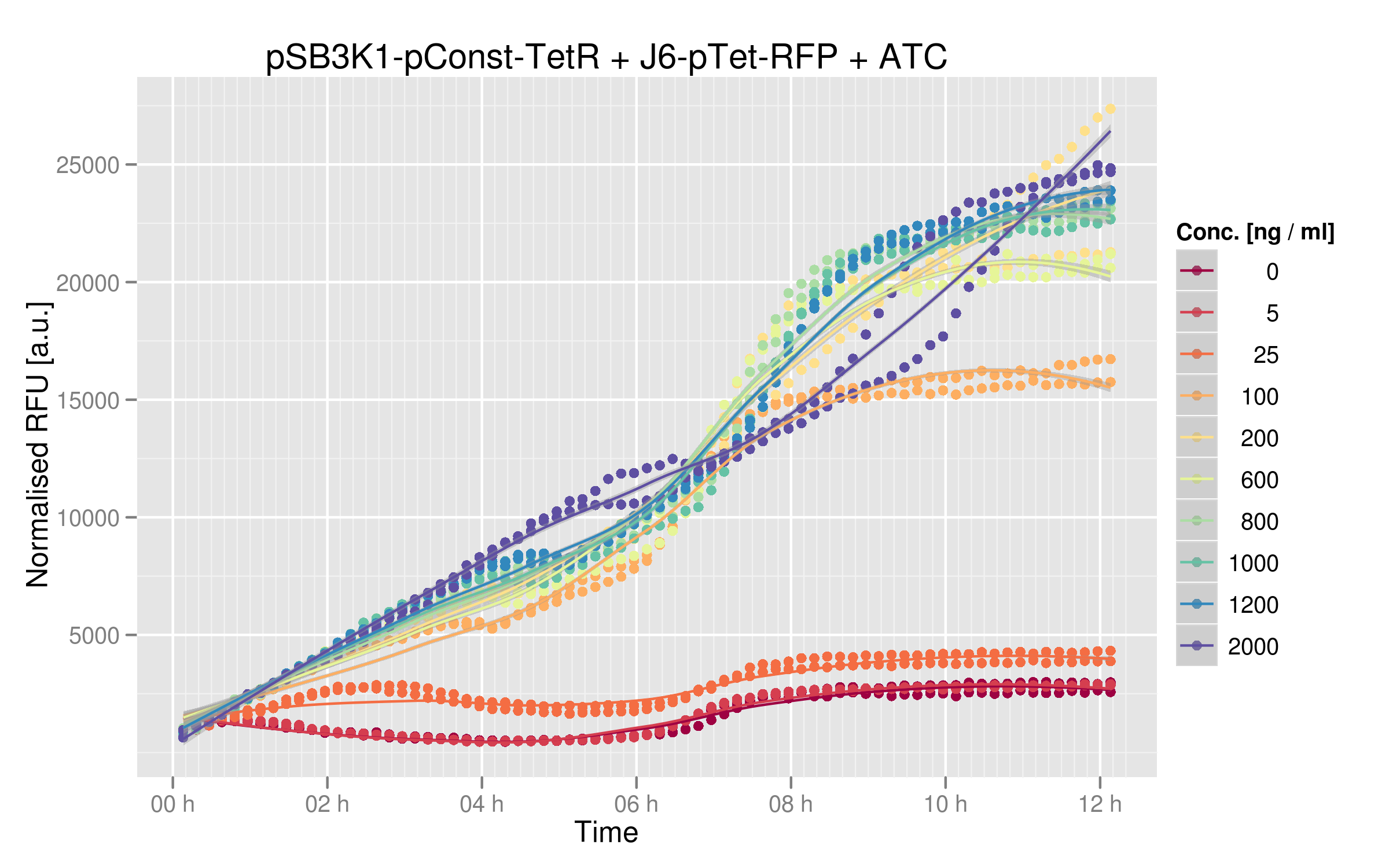
We ran platereader experiments for 6 of our mutants. Increasing concentrations of ATC were added to force expression of RFP, which allowed us to appreciate the repressive effect of TetR on fluorescence expression. The following graph shows wild-type TetR characterization. Increasing ATC concentrations do lead to increased fluorescence, which confirms that tetR was initially repressing RFP:
4) comparing the results of both characterization
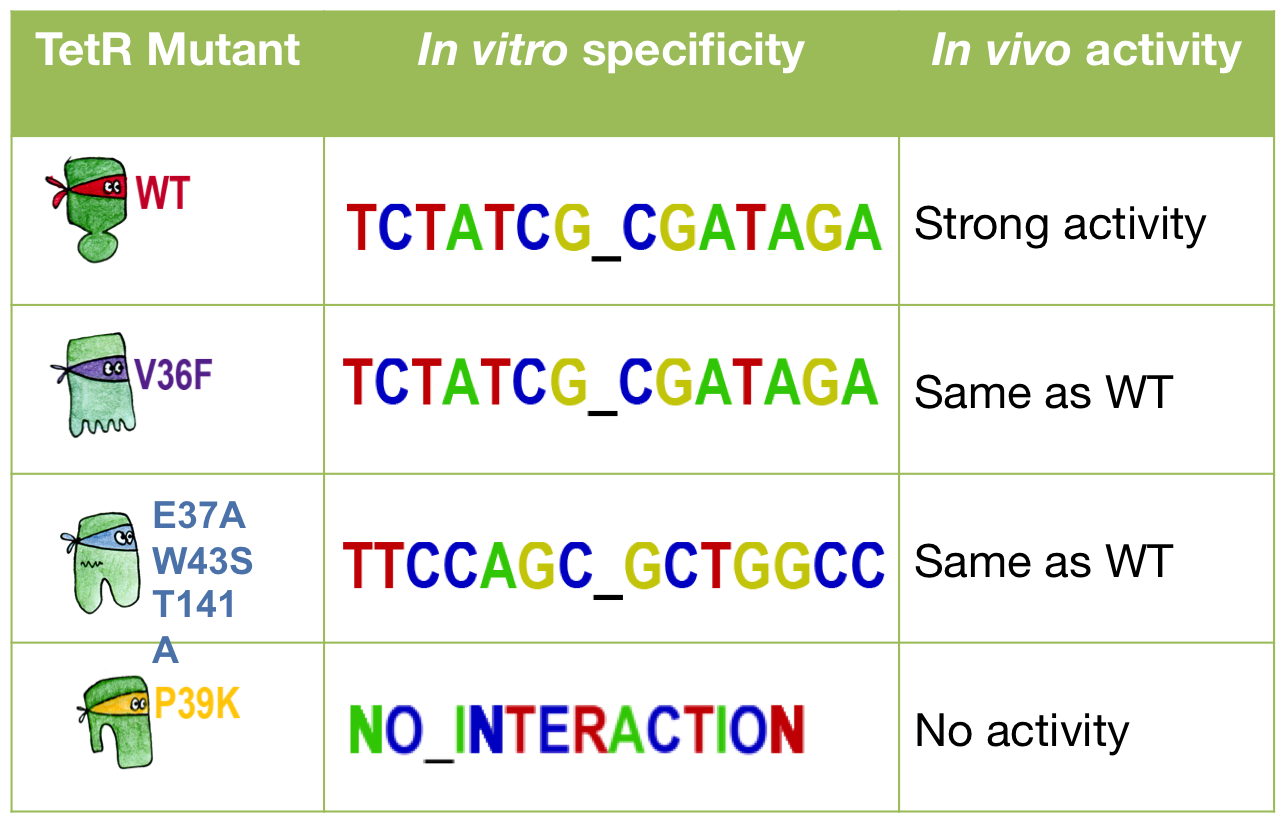
Three of our mutants plus the wild-type were tested both in vivo and in vitro. The results were consistent between the two experiments. The V36F exhibits the same behaviour as the wild-type while the P39K shows no interaction to the DNA at all. The E37A triple mutant, however, has contradictory results: the MITOMI data indicate that this mutant has a change in specificity, with a consensus sequence being different from the wild-type Ptet. However, it still recognizes Ptet well, meaning that there is still crosstalk with the endogenous promoter.
We were able to change the specificity of some of our mutants; however this change was not big enough to have a fully orthogonal mutant that would not recognize the wild-type consensus sequence.
DIY Microfluidics
In parallel to the work on the transcription factor pipeline, we built a computer-controlled microfluidic injection setup from off-the shelf components, to demonstrate that microfluidics can be used for CHF 1000 (about €800). To help future teams from other universities get started with microfluidics, we wrote on our wiki instructions to replicate the setup.
In addition to this, we programmed a web interface for the machine: from any web browser, users can view a live video feed from the microscope that images the chip, and control the valves by clicking buttons on the web page. Concurrent use is managed by a queue system. The web site was mostly used at the Jamboree in Amsterdam to illustrate the principle of microfluidics to other iGEMers, then it stayed live for most of October, allowing anybody to try the setup. The demonstration chip had channels in the shape of the iGEM logo, and was filled with a red and a blue dye to highlight flow and mixing.
 "
"