Team:EPF-Lausanne/Tools/MITOMI
From 2011.igem.org
| Line 9: | Line 9: | ||
[[File:Mitomi.PNG|800x348px]] | [[File:Mitomi.PNG|800x348px]] | ||
| + | |||
| + | The control inputs control the valves that close the | ||
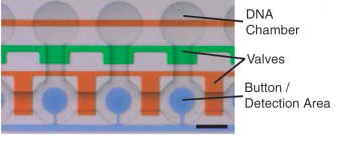
| + | Detailed view of the chambers and of the movable membrane used for mechanical trapping, referred to as the 'Button' . | ||
| + | |||
[[File:Mitomi chamber.PNG]] | [[File:Mitomi chamber.PNG]] | ||
| - | |||
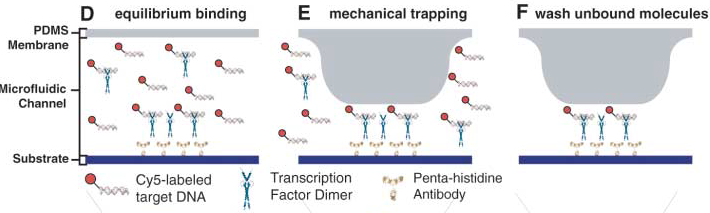
'''Advantages:''' The movable memrane shown in the scheme below enables measurements of transient interactions, exhibiting sub micromolar affinities which are generally problematic. | '''Advantages:''' The movable memrane shown in the scheme below enables measurements of transient interactions, exhibiting sub micromolar affinities which are generally problematic. | ||
Revision as of 16:01, 8 August 2011
MITOMI
Mitomi is a high troughput microfluidic system designed to mesure low affinity molecular interactions by mechanically trapping them.
Purpose: We applied this technology during our project for characterizing transcription factor binding energy landscapes in absolute terms by determining dissociation constants (Kd) for a comprehensive set of target DNA sequences.
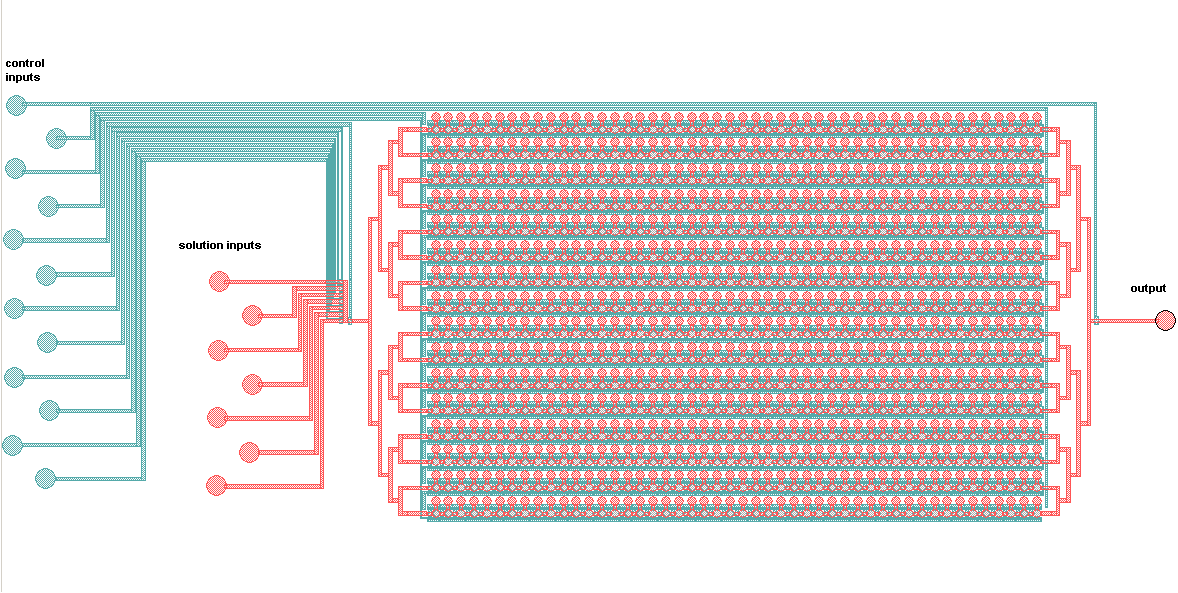
Schematic of the Mitomi platform:
The control inputs control the valves that close the Detailed view of the chambers and of the movable membrane used for mechanical trapping, referred to as the 'Button' .
Advantages: The movable memrane shown in the scheme below enables measurements of transient interactions, exhibiting sub micromolar affinities which are generally problematic.
 "
"