Team:EPF-Lausanne/Our Project/TetR mutants/MITOMI data
From 2011.igem.org
(→Mutant TetRs) |
|||
| (44 intermediate revisions not shown) | |||
| Line 10: | Line 10: | ||
=Wildtype TetR= | =Wildtype TetR= | ||
| - | + | ===In Vitro Characterization=== | |
| + | Using the <html> <a href="https://2011.igem.org/Team:EPF-Lausanne/Tools/MITOMI">MITOMI</a></html> technique we determined the DNA binding landscape of the wild type TetR. To do so, we first designed and generated the library of double-stranded DNA sequences that cover all possible single base substitutions within the TetO operator sequence. Based on that library we measured the dissociation constants of the mutant to all the tetO-like sequences of the library. Then, we determined the specificity of the mutant to the Tet operator sequence, expressed as a position-weight matrix (PWM). | ||
| - | + | The enoLOGO we obtained for the wild-type is: | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
[[File:EPFL2011_WTTetR_WebLogo.png|700px]] | [[File:EPFL2011_WTTetR_WebLogo.png|700px]] | ||
| - | ''' | + | '''enoLOGO reference:'''<p> |
Workman CT, Yin Y, Corcoran DL, Ideker T, Stormo GD, Benos PV. | Workman CT, Yin Y, Corcoran DL, Ideker T, Stormo GD, Benos PV. | ||
enoLOGOS: a versatile web tool for energy normalized sequence logos. | enoLOGOS: a versatile web tool for energy normalized sequence logos. | ||
Nucleic Acids Res. 2005 Jul 1;33:W389-92.</p> | Nucleic Acids Res. 2005 Jul 1;33:W389-92.</p> | ||
| - | |||
| - | |||
===Position Weight Matrix=== | ===Position Weight Matrix=== | ||
| Line 73: | Line 68: | ||
|} | |} | ||
| - | Each row represents the changes in binding energy, ΔΔG (kcal/mol), compared to the reference sequence upon the substitution | + | Each row represents the changes in binding energy, ΔΔG (kcal/mol), compared to the reference sequence upon the substitution of the indicated nucleotide at a certain position within the target DNA element. Values are indicated in kcal/mol. |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
=Mutant TetRs= | =Mutant TetRs= | ||
| Line 84: | Line 75: | ||
<html> <a href=http://partsregistry.org/Part:BBa_K613013>BBa_K613013</a></html> | <html> <a href=http://partsregistry.org/Part:BBa_K613013>BBa_K613013</a></html> | ||
| - | ===In | + | ===In Vitro Characterization=== |
| - | Using the <html> <a href="https://2011.igem.org/Team:EPF-Lausanne/ | + | Using the <html> <a href="https://2011.igem.org/Team:EPF-Lausanne/Tools/MITOMI">MITOMI</a></html> technique we determined the DNA binding landscape of the TetR V36F mutant. To do so, first we designed and generated the library of double stranded DNA sequences that cover all possible single base substitution within the tetO operator sequence. Based on that library we measured the dissociation constants of the mutant to variable tetO-like sequences and determined the specificity of the mutant to the tet operator sequence (expressed as a PWM). |
| - | + | The enoLOGO we obtained for the V36F mutant: | |
[[File:EPFL2011_MITOMI_WebLogo_V36F.png|700px]] | [[File:EPFL2011_MITOMI_WebLogo_V36F.png|700px]] | ||
| - | ''' | + | '''enoLOGO reference:'''<p> |
Workman CT, Yin Y, Corcoran DL, Ideker T, Stormo GD, Benos PV. | Workman CT, Yin Y, Corcoran DL, Ideker T, Stormo GD, Benos PV. | ||
enoLOGOS: a versatile web tool for energy normalized sequence logos. | enoLOGOS: a versatile web tool for energy normalized sequence logos. | ||
Nucleic Acids Res. 2005 Jul 1;33:W389-92.</p> | Nucleic Acids Res. 2005 Jul 1;33:W389-92.</p> | ||
| - | |||
| - | |||
===Position Weight Matrix=== | ===Position Weight Matrix=== | ||
| Line 146: | Line 135: | ||
Each row represents the changes in binding energy, ΔΔG (kcal/mol), compared to the reference sequence upon the substitution to the indicated nucleotide at certain position within the target DNA element. Values are indicated in kcal/mol. | Each row represents the changes in binding energy, ΔΔG (kcal/mol), compared to the reference sequence upon the substitution to the indicated nucleotide at certain position within the target DNA element. Values are indicated in kcal/mol. | ||
| - | We compared the measured DNA binding affinities of the V36F mutant to the affinities obtained for the wt-TetR and found that V36F mutant interacts with the | + | We compared the measured DNA binding affinities of the V36F mutant to the affinities obtained for the wt-TetR and found that V36F mutant interacts with the TetO as strongly as the wild-type variant. |
[[File:EPFL2011_Scatter_plot_kd_wt_VF36.png|600px]] | [[File:EPFL2011_Scatter_plot_kd_wt_VF36.png|600px]] | ||
| - | |||
| - | |||
| - | |||
| - | |||
==E37A W43S T141A== | ==E37A W43S T141A== | ||
<html> <a href=http://partsregistry.org/Part:BBa_K613015>BBa_K613015</a></html> | <html> <a href=http://partsregistry.org/Part:BBa_K613015>BBa_K613015</a></html> | ||
| - | ===In | + | ===In Vitro Characterization=== |
| - | Using the <html> <a href="https://2011.igem.org/Team:EPF-Lausanne/ | + | Using the <html> <a href="https://2011.igem.org/Team:EPF-Lausanne/Tools/MITOMI">MITOMI</a></html> technique we determined the DNA binding landscape of the TetR E37A W43S T141A mutant. To do so, first we designed and generated the library of double stranded DNA sequences that cover all possible single base substitution within the tetO operator sequence. Based on that library we measured the dissociation constants of the mutant to variable tetO-like sequences and determined the specificity of the mutant to the tet operator sequence (expressed as a PWM). |
| - | + | The measured DNA binding affinities of the E37A W43S T141A mutant show that these mutations alter the sequence recognition, while the symmetry and the spacer affinities are conserved. | |
| + | |||
| + | |||
| + | The enoLOGO we obtained for the E37A W43S T141A mutant: | ||
[[File:EPFL2011_WebLogo_EA37WS43TA41.png|700px]] | [[File:EPFL2011_WebLogo_EA37WS43TA41.png|700px]] | ||
| - | ''' | + | '''enoLOGO reference:'''<p> |
Workman CT, Yin Y, Corcoran DL, Ideker T, Stormo GD, Benos PV. | Workman CT, Yin Y, Corcoran DL, Ideker T, Stormo GD, Benos PV. | ||
enoLOGOS: a versatile web tool for energy normalized sequence logos. | enoLOGOS: a versatile web tool for energy normalized sequence logos. | ||
Nucleic Acids Res. 2005 Jul 1;33:W389-92.</p> | Nucleic Acids Res. 2005 Jul 1;33:W389-92.</p> | ||
| - | |||
===Position Weight Matrix=== | ===Position Weight Matrix=== | ||
| Line 220: | Line 207: | ||
<html> <a href=http://partsregistry.org/Part:BBa_K613016>BBa_K613016</a></html> | <html> <a href=http://partsregistry.org/Part:BBa_K613016>BBa_K613016</a></html> | ||
| - | ===In | + | ===In Vitro Characterization=== |
| - | Using the <html> <a href="https://2011.igem.org/Team:EPF-Lausanne/ | + | Using the <html> <a href="https://2011.igem.org/Team:EPF-Lausanne/Tools/MITOMI">MITOMI</a></html> technique we determined the DNA binding landscape of the TetR P39K mutant. To do so, first we designed and generated the library of double stranded DNA sequences that cover all possible single base substitution within the tetO operator sequence. Based on that library we measured the dissociation constants of the mutant to variable tetO-like sequences and determined the specificity of the mutant to the tet operator sequence (expressed as a PWM). |
| - | + | ||
| + | The enoLOGO we obtained for the P39K mutant: | ||
[[File:EPFL2011_P39K_WebLogo.png|700px]] | [[File:EPFL2011_P39K_WebLogo.png|700px]] | ||
| + | |||
| + | The strong difference of the binding affinities between the P39K mutant and the wtTetR, might be due to the altered recognition of the P39K. This mutant was shown to have a new recognition specificity for the tetO-4C operator in [http://www.sciencedirect.com/science/article/pii/S0022283697915400 Helbl et al, 1998]. | ||
| Line 233: | Line 223: | ||
enoLOGOS: a versatile web tool for energy normalized sequence logos. | enoLOGOS: a versatile web tool for energy normalized sequence logos. | ||
Nucleic Acids Res. 2005 Jul 1;33:W389-92.</p> | Nucleic Acids Res. 2005 Jul 1;33:W389-92.</p> | ||
| + | Helbl, V., and Hillen, W. (1998). Stepwise selection of TetR variants recognizing tet operator 4C with high affinity and specificity. J Mol Biol 276, 313-318. | ||
| + | |||
| + | |||
===Position Weight Matrix=== | ===Position Weight Matrix=== | ||
| Line 284: | Line 277: | ||
<html> <a href=http://partsregistry.org/Part:BBa_K613017>BBa_K613017</a></html> | <html> <a href=http://partsregistry.org/Part:BBa_K613017>BBa_K613017</a></html> | ||
| - | ===In | + | ===In Vitro Characterization=== |
| - | Using the <html> <a href="https://2011.igem.org/Team:EPF-Lausanne/ | + | Using the <html> <a href="https://2011.igem.org/Team:EPF-Lausanne/Tools/MITOMI">MITOMI</a></html> technique we determined the DNA binding landscape of the TetR Y42F mutant. To do so, first we designed and generated the library of double stranded DNA sequences that cover all possible single base substitution within the tetO operator sequence. Based on that library we measured the dissociation constants of the mutant to variable tetO-like sequences and determined the specificity of the mutant to the tet operator sequence (expressed as a PWM). For the Y42F mutant we observed the decrease of the specificity compared to the wild-type tetR sequence. |
| - | + | The enoLOGO we obtained for the Y42F mutant: | |
[[File:EPFL2011_WebLogo_Y42F.png|700px]] | [[File:EPFL2011_WebLogo_Y42F.png|700px]] | ||
| - | ''' | + | The measured binding affinities show no symmetry conservation and the contribution of the outer bases equals that of the inner. These experiment will have to be repeated in order to get a clearer idea of the binding affinities. |
| + | |||
| + | |||
| + | '''enoLOGO reference:'''<p> | ||
Workman CT, Yin Y, Corcoran DL, Ideker T, Stormo GD, Benos PV. | Workman CT, Yin Y, Corcoran DL, Ideker T, Stormo GD, Benos PV. | ||
enoLOGOS: a versatile web tool for energy normalized sequence logos. | enoLOGOS: a versatile web tool for energy normalized sequence logos. | ||
Nucleic Acids Res. 2005 Jul 1;33:W389-92.</p> | Nucleic Acids Res. 2005 Jul 1;33:W389-92.</p> | ||
| - | + | ===Position Weight Matrix=== | |
| - | + | ||
{| {{table}} | {| {{table}} | ||
| align="center" style="background:#f0f0f0;"|'''PO''' | | align="center" style="background:#f0f0f0;"|'''PO''' | ||
| Line 343: | Line 338: | ||
|} | |} | ||
| - | Each row represents the changes in binding energy, ΔΔG, compared to the reference sequence upon the substitution | + | Each row represents the changes in binding energy, ΔΔG, compared to the reference sequence upon the substitution at the indicated nucleotide at a certain position within the target DNA element. Values are indicated in kcal/mol. |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
==P39Q Y42M== | ==P39Q Y42M== | ||
<html> <a href=http://partsregistry.org/Part:BBa_K613019>BBa_K613019</a></html> | <html> <a href=http://partsregistry.org/Part:BBa_K613019>BBa_K613019</a></html> | ||
| - | ===In | + | ===In Vitro Characterization=== |
| - | Using the <html> <a href="https://2011.igem.org/Team:EPF-Lausanne/ | + | Using the <html> <a href="https://2011.igem.org/Team:EPF-Lausanne/Tools/MITOMI">MITOMI</a></html> technique we determined the DNA binding landscape of the TetR P39Q Y42M mutant. To do so, first we designed and generated the library of double stranded DNA sequences that cover all possible single base substitution within the tetO operator sequence. Based on that library we measured the dissociation constants of the mutant to variable tetO-like sequences and determined the specificity of the mutant to the tet operator sequence (expressed as a PWM). For the P39Q Y42M mutant we observed the decrease of the specificity compared to the wild-type tetR sequence. |
| - | + | The enoLOGO we obtained for the P39Q Y42M mutant: | |
[[File:EPFL2011_P39QY42M_WebLogo.png|700px]] | [[File:EPFL2011_P39QY42M_WebLogo.png|700px]] | ||
| + | '''enoLOGO reference:'''<p> | ||
| + | Workman CT, Yin Y, Corcoran DL, Ideker T, Stormo GD, Benos PV. | ||
| + | enoLOGOS: a versatile web tool for energy normalized sequence logos. | ||
| + | Nucleic Acids Res. 2005 Jul 1;33:W389-92.</p> | ||
| - | + | ===Position Weight Matrix=== | |
{| {{table}} | {| {{table}} | ||
| Line 405: | Line 399: | ||
| 17||0.0634094||0||0.455977||0.0582705 | | 17||0.0634094||0||0.455977||0.0582705 | ||
|} | |} | ||
| + | |||
| + | |||
Each row represents the changes in binding energy, ΔΔG, compared to the reference sequence upon the substitution to the indicated nucleotide at certain position within the target DNA element. Values are indicated in kcal/mol. | Each row represents the changes in binding energy, ΔΔG, compared to the reference sequence upon the substitution to the indicated nucleotide at certain position within the target DNA element. Values are indicated in kcal/mol. | ||
| - | =Uncharacterized | + | =Uncharacterized Mutant TetRs= |
| + | |||
| + | |||
| + | Unfortunately we did not have time to finish the characterization of the following parts: | ||
| + | |||
| + | ==Y42F K108E== | ||
| + | |||
| + | <html> <a href=http://partsregistry.org/Part:BBa_K613018>BBa_K613018</a></html> | ||
| + | |||
| + | ==V36F W43S== | ||
| + | <html> <a href=http://partsregistry.org/Part:BBa_K613014>BBa_K613014</a></html> | ||
==P39Q Y42M L197S== | ==P39Q Y42M L197S== | ||
Latest revision as of 21:49, 26 October 2011
MITOMI Data
In vitro Main | Why TetR? | Mutant TetRs | MITOMI Data | In-vivo & In-vitro outline
To find out more about the MITOMI method of characterization, please click here.
The raw data from the successful experiments can be found here.
Contents |
Wildtype TetR
In Vitro Characterization
Using the MITOMI technique we determined the DNA binding landscape of the wild type TetR. To do so, we first designed and generated the library of double-stranded DNA sequences that cover all possible single base substitutions within the TetO operator sequence. Based on that library we measured the dissociation constants of the mutant to all the tetO-like sequences of the library. Then, we determined the specificity of the mutant to the Tet operator sequence, expressed as a position-weight matrix (PWM).
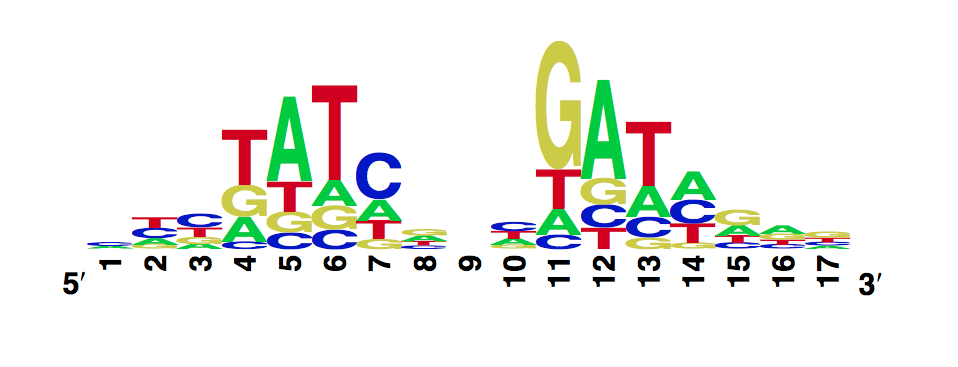
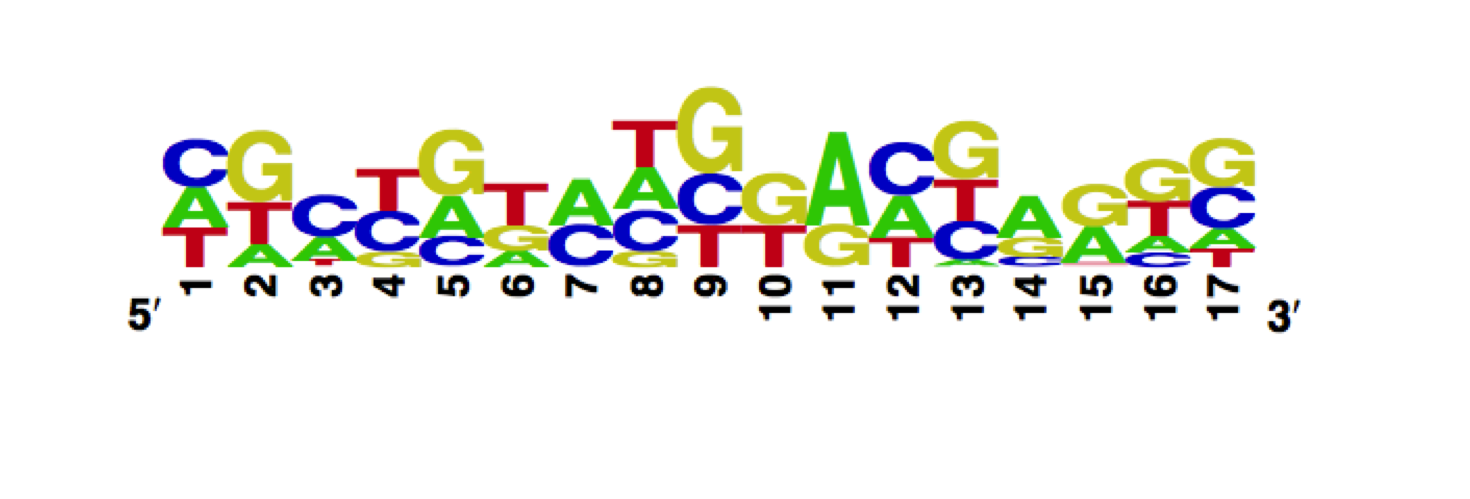
The enoLOGO we obtained for the wild-type is:
Workman CT, Yin Y, Corcoran DL, Ideker T, Stormo GD, Benos PV. enoLOGOS: a versatile web tool for energy normalized sequence logos. Nucleic Acids Res. 2005 Jul 1;33:W389-92.
Position Weight Matrix
| PO | A | T | C | G |
| 1 | 0.164072 | 0.519334 | -0.0468541 | 0.22966 |
| 2 | 0.642569 | 0.164072 | 0.188965 | 0.943314 |
| 3 | 1.04481 | 0.558693 | 0.164072 | 0.848264 |
| 4 | 0.959646 | 0.164072 | 2.17536 | 0.718959 |
| 5 | 0.164072 | 1.19894 | 1.79469 | 1.56685 |
| 6 | 1.46463 | 0.164072 | 1.74384 | 1.54845 |
| 7 | 0.966397 | 1.07557 | 0.164072 | 1.72883 |
| 8 | 0.164072 | 0.504312 | 0.755195 | 0.148902 |
| 9 | 0.0876645 | 0.164072 | 0 | 0.0156484 |
| 10 | 0.289722 | 0.164072 | 0.00707563 | 0.843474 |
| 11 | 1.77568 | 1.3447 | 2.34804 | 0.164072 |
| 12 | 0.164072 | 1.72354 | 1.62782 | 1.49176 |
| 13 | 0.877518 | 0.164072 | 1.30879 | 1.9052 |
| 14 | 0.164072 | 0.544642 | 0.3387 | 1.7537 |
| 15 | 0.540091 | 0.8821 | 1.0861 | 0.164072 |
| 16 | -0.224147 | 0.280358 | 0.442769 | 0.164072 |
| 17 | 0.31238 | -0.0651059 | 0.164072 | -0.303844 |
Each row represents the changes in binding energy, ΔΔG (kcal/mol), compared to the reference sequence upon the substitution of the indicated nucleotide at a certain position within the target DNA element. Values are indicated in kcal/mol.
Mutant TetRs
V36F
In Vitro Characterization
Using the MITOMI technique we determined the DNA binding landscape of the TetR V36F mutant. To do so, first we designed and generated the library of double stranded DNA sequences that cover all possible single base substitution within the tetO operator sequence. Based on that library we measured the dissociation constants of the mutant to variable tetO-like sequences and determined the specificity of the mutant to the tet operator sequence (expressed as a PWM).
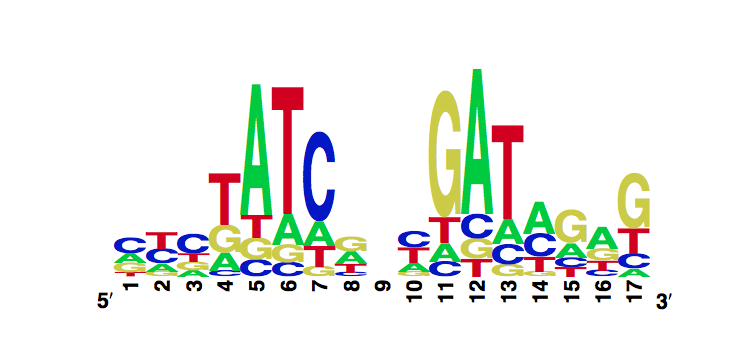
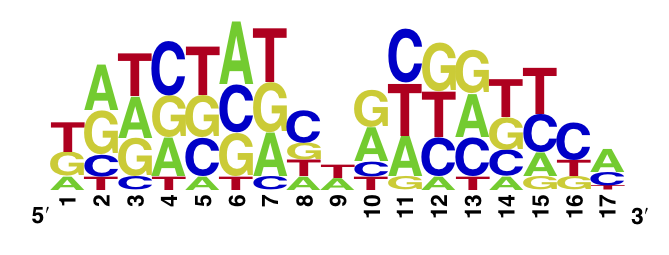
The enoLOGO we obtained for the V36F mutant:
Workman CT, Yin Y, Corcoran DL, Ideker T, Stormo GD, Benos PV. enoLOGOS: a versatile web tool for energy normalized sequence logos. Nucleic Acids Res. 2005 Jul 1;33:W389-92.
Position Weight Matrix
| PO | A | T | C | G |
| 1 | 0.164072 | 0.519334 | 0 | 0.22966 |
| 2 | 0.642569 | 0.164072 | 0.188965 | 0.943314 |
| 3 | 1.04481 | 0.558693 | 0.164072 | 0.848264 |
| 4 | 0.959646 | 0.164072 | 2.17536 | 0.718959 |
| 5 | 0.164072 | 1.19894 | 1.79469 | 1.56685 |
| 6 | 1.46463 | 0.164072 | 1.74384 | 1.54845 |
| 7 | 0.966397 | 1.07557 | 0.164072 | 1.72883 |
| 8 | 0.164072 | 0.504312 | 0.755195 | 0.148902 |
| 9 | 0.0876645 | 0.164072 | 0 | 0.0156484 |
| 10 | 0.289722 | 0.164072 | 0.00707563 | 0.843474 |
| 11 | 1.77568 | 1.3447 | 2.34804 | 0.164072 |
| 12 | 0.164072 | 1.72354 | 1.62782 | 1.49176 |
| 13 | 0.877518 | 0.164072 | 1.30879 | 1.9052 |
| 14 | 0.164072 | 0.544642 | 0.3387 | 1.7537 |
| 15 | 0.540091 | 0.8821 | 1.0861 | 0.164072 |
| 16 | 0 | 0.280358 | 0.442769 | 0.164072 |
| 17 | 0.31238 | 0 | 0.164072 | 0 |
Each row represents the changes in binding energy, ΔΔG (kcal/mol), compared to the reference sequence upon the substitution to the indicated nucleotide at certain position within the target DNA element. Values are indicated in kcal/mol.
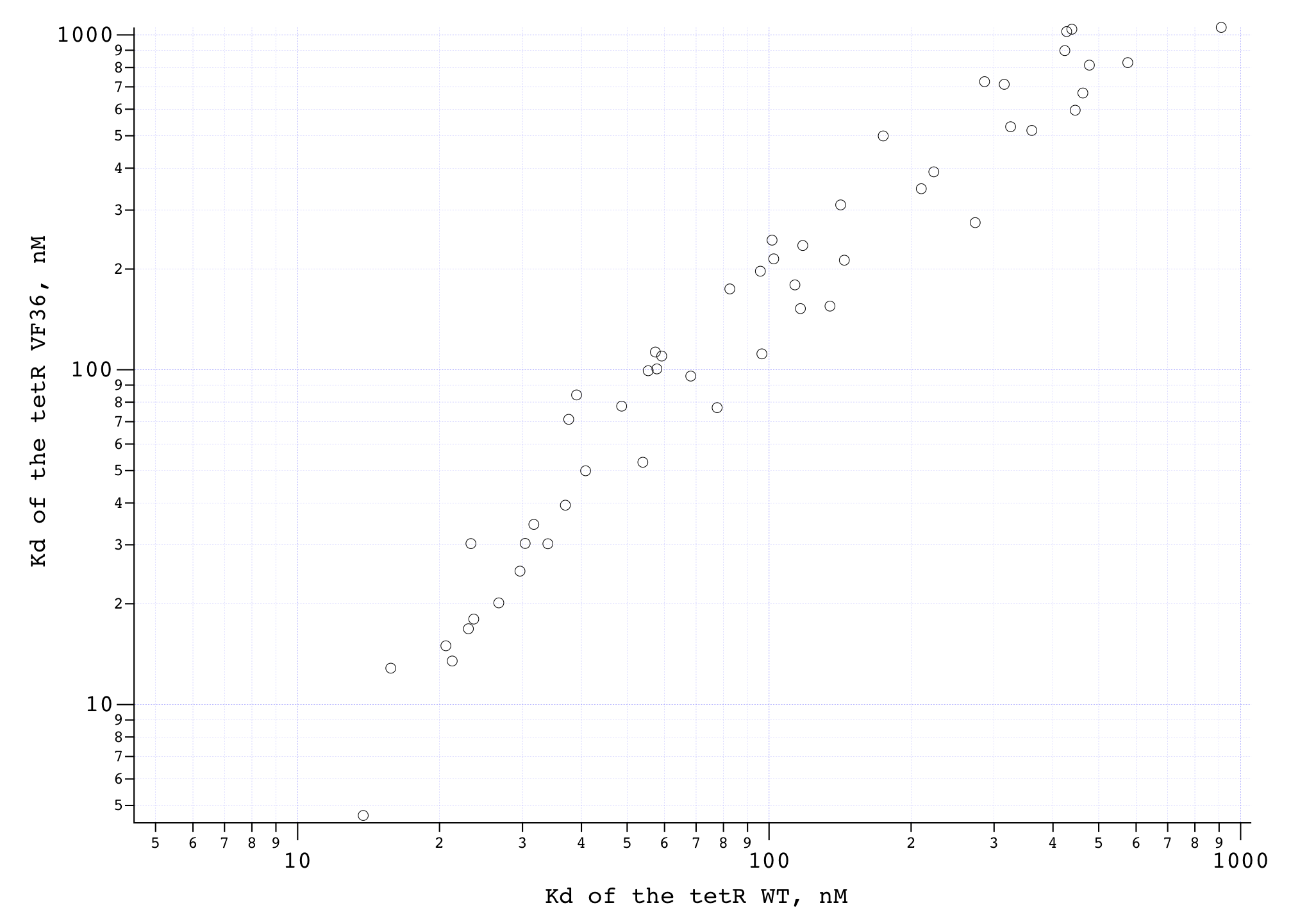
We compared the measured DNA binding affinities of the V36F mutant to the affinities obtained for the wt-TetR and found that V36F mutant interacts with the TetO as strongly as the wild-type variant.
E37A W43S T141A
In Vitro Characterization
Using the MITOMI technique we determined the DNA binding landscape of the TetR E37A W43S T141A mutant. To do so, first we designed and generated the library of double stranded DNA sequences that cover all possible single base substitution within the tetO operator sequence. Based on that library we measured the dissociation constants of the mutant to variable tetO-like sequences and determined the specificity of the mutant to the tet operator sequence (expressed as a PWM).
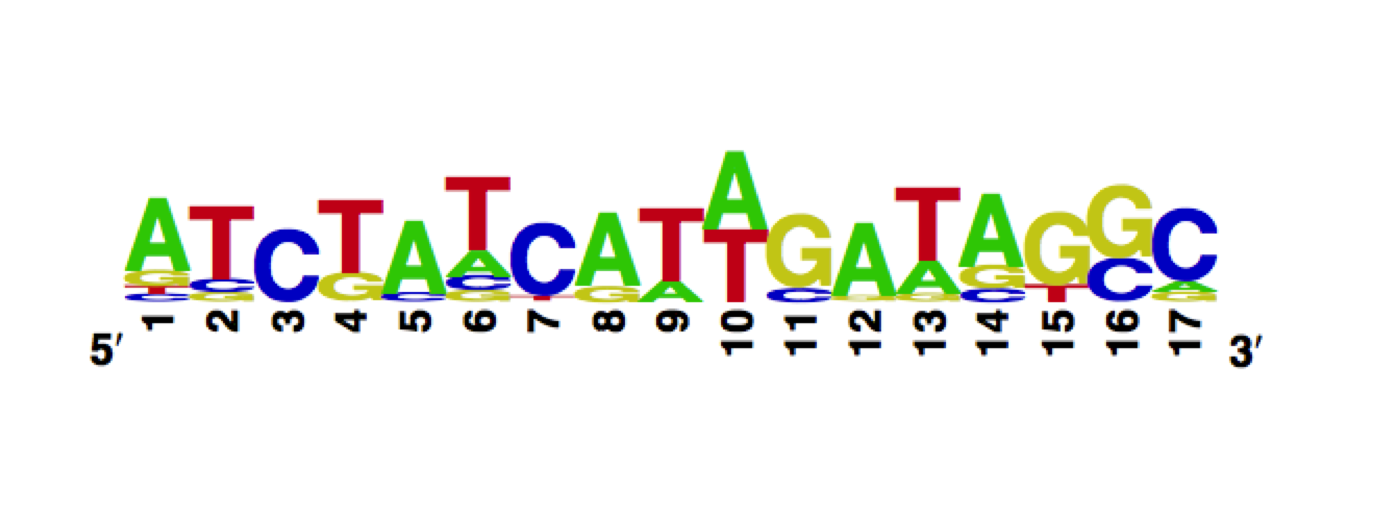
The measured DNA binding affinities of the E37A W43S T141A mutant show that these mutations alter the sequence recognition, while the symmetry and the spacer affinities are conserved.
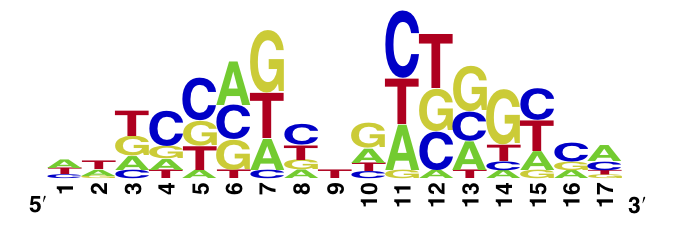
The enoLOGO we obtained for the E37A W43S T141A mutant:
Workman CT, Yin Y, Corcoran DL, Ideker T, Stormo GD, Benos PV. enoLOGOS: a versatile web tool for energy normalized sequence logos. Nucleic Acids Res. 2005 Jul 1;33:W389-92.
Position Weight Matrix
| PO | A | T | C | G |
| 1 | 0.298921 | 0.228558 | 0.136478 | 0.0565634 |
| 2 | 0.217725 | 0.298921 | 0.0365532 | 0.101569 |
| 3 | 0.486164 | 0.884721 | 0.298921 | 0.689301 |
| 4 | 0.381337 | 0.298921 | 1.2275 | 0.443129 |
| 5 | 0.298921 | 0.829256 | 1.48516 | 0.861923 |
| 6 | 1.52008 | 0.298921 | 1.20145 | 1.04388 |
| 7 | 1.08347 | 1.57658 | 0.298921 | 2.1751 |
| 8 | 0.298921 | 0.506289 | 0.723699 | 0.34977 |
| 9 | 0.0627581 | 0.298921 | 0 | 0.0173955 |
| 10 | 0.49991 | 0.298921 | 0.250601 | 0.869584 |
| 11 | 1.58199 | 1.5844 | 2.35356 | 0.298921 |
| 12 | 0.298921 | 1.88623 | 1.34104 | 1.46941 |
| 13 | 1.00088 | 0.298921 | 1.01931 | 1.57583 |
| 14 | 0.298921 | 0.56873 | 0.306484 | 1.91995 |
| 15 | 0.65822 | 1.06676 | 1.10609 | 0.298921 |
| 16 | 0.290169 | 0.067613 | 0.660542 | 0.298921 |
| 17 | 0.557743 | 0.160129 | 0.298921 | 0.134017 |
Each row represents the changes in binding energy, ΔΔG, compared to the reference sequence upon the substitution to the indicated nucleotide at certain position within the target DNA element. Values are indicated in kcal/mol.
P39K
In Vitro Characterization
Using the MITOMI technique we determined the DNA binding landscape of the TetR P39K mutant. To do so, first we designed and generated the library of double stranded DNA sequences that cover all possible single base substitution within the tetO operator sequence. Based on that library we measured the dissociation constants of the mutant to variable tetO-like sequences and determined the specificity of the mutant to the tet operator sequence (expressed as a PWM).
The enoLOGO we obtained for the P39K mutant:
The strong difference of the binding affinities between the P39K mutant and the wtTetR, might be due to the altered recognition of the P39K. This mutant was shown to have a new recognition specificity for the tetO-4C operator in [http://www.sciencedirect.com/science/article/pii/S0022283697915400 Helbl et al, 1998].
Workman CT, Yin Y, Corcoran DL, Ideker T, Stormo GD, Benos PV. enoLOGOS: a versatile web tool for energy normalized sequence logos. Nucleic Acids Res. 2005 Jul 1;33:W389-92.
Helbl, V., and Hillen, W. (1998). Stepwise selection of TetR variants recognizing tet operator 4C with high affinity and specificity. J Mol Biol 276, 313-318.
Position Weight Matrix
| PO | A | T | C | G |
| 1 | 0.259286 | 0.242355 | 0.296487 | 0 |
| 2 | 0.13749 | 0.259286 | 0 | 0.444867 |
| 3 | 0.149356 | 0.0391642 | 0.259286 | 0 |
| 4 | 0.000432676 | 0.259286 | 0.250803 | 0.09369 |
| 5 | 0.259286 | 0 | 0.183728 | 0.411598 |
| 6 | 0.104299 | 0.259286 | 0 | 0.157345 |
| 7 | 0.287302 | 0.0195324 | 0.259286 | 0.0244544 |
| 8 | 0.259286 | 0.286228 | 0.256377 | 0.0991247 |
| 9 | 0 | 0.259286 | 0.317281 | 0.536557 |
| 10 | 0 | 0.259286 | 0 | 0.31848 |
| 11 | 0.582859 | 0 | 0 | 0.259286 |
| 12 | 0.259286 | 0.184012 | 0.330234 | 0 |
| 13 | 0.0462643 | 0.259286 | 0.242912 | 0.357439 |
| 14 | 0.259286 | 0.011556 | 0.0598842 | 0.120511 |
| 15 | 0.228422 | 0.0270438 | 0.00125937 | 0.259286 |
| 16 | 0.102476 | 0.214565 | 0.0872813 | 0.259286 |
| 17 | 0.127319 | 0.112242 | 0.259286 | 0.299159 |
Each row represents the changes in binding energy, ΔΔG, compared to the reference sequence upon the substitution to the indicated nucleotide at certain position within the target DNA element. Values are indicated in kcal/mol.
Y42F
In Vitro Characterization
Using the MITOMI technique we determined the DNA binding landscape of the TetR Y42F mutant. To do so, first we designed and generated the library of double stranded DNA sequences that cover all possible single base substitution within the tetO operator sequence. Based on that library we measured the dissociation constants of the mutant to variable tetO-like sequences and determined the specificity of the mutant to the tet operator sequence (expressed as a PWM). For the Y42F mutant we observed the decrease of the specificity compared to the wild-type tetR sequence.
The enoLOGO we obtained for the Y42F mutant:
The measured binding affinities show no symmetry conservation and the contribution of the outer bases equals that of the inner. These experiment will have to be repeated in order to get a clearer idea of the binding affinities.
Workman CT, Yin Y, Corcoran DL, Ideker T, Stormo GD, Benos PV. enoLOGOS: a versatile web tool for energy normalized sequence logos. Nucleic Acids Res. 2005 Jul 1;33:W389-92.
Position Weight Matrix
| PO | A | T | C | G |
| 1 | 0.385241 | 0.85619 | 0 | 0.693793 |
| 2 | 1.30965 | 0.385241 | 0.629213 | 1.2542 |
| 3 | 1.18851 | 1.22909 | 0.385241 | 1.09825 |
| 4 | 1.1084 | 0.385241 | 1.57045 | 1.20206 |
| 5 | 0.385241 | 1.3955 | 1.10171 | 1.2069 |
| 6 | 1.80251 | 0.385241 | 1.37974 | 1.23728 |
| 7 | 1.25285 | 1.54821 | 0.385241 | 1.43937 |
| 8 | 0.385241 | 0.475161 | 0.93954 | 0.488479 |
| 9 | 0.341937 | 0.385241 | 0 | 0.053757 |
| 10 | 0.973293 | 0.385241 | 0.426141 | 1.06563 |
| 11 | 1.1587 | 1.50073 | 1.5542 | 0.385241 |
| 12 | 0.385241 | 1.30762 | 1.11351 | 1.39762 |
| 13 | 1.23638 | 0.385241 | 1.11282 | 1.24361 |
| 14 | 0.385241 | 1.09074 | 0.75983 | 0.933921 |
| 15 | 0.679315 | 1.33124 | 1.09937 | 0.385241 |
| 16 | 0 | 0.467396 | 1.08277 | 0.385241 |
| 17 | 0.664957 | 0.130021 | 0.385241 | 0 |
Each row represents the changes in binding energy, ΔΔG, compared to the reference sequence upon the substitution at the indicated nucleotide at a certain position within the target DNA element. Values are indicated in kcal/mol.
P39Q Y42M
In Vitro Characterization
Using the MITOMI technique we determined the DNA binding landscape of the TetR P39Q Y42M mutant. To do so, first we designed and generated the library of double stranded DNA sequences that cover all possible single base substitution within the tetO operator sequence. Based on that library we measured the dissociation constants of the mutant to variable tetO-like sequences and determined the specificity of the mutant to the tet operator sequence (expressed as a PWM). For the P39Q Y42M mutant we observed the decrease of the specificity compared to the wild-type tetR sequence.
The enoLOGO we obtained for the P39Q Y42M mutant:
enoLOGO reference:Workman CT, Yin Y, Corcoran DL, Ideker T, Stormo GD, Benos PV. enoLOGOS: a versatile web tool for energy normalized sequence logos. Nucleic Acids Res. 2005 Jul 1;33:W389-92.
Position Weight Matrix
| PO | A | T | C | G |
| 1 | 0.455977 | 0.0513378 | 0.0469535 | 0.0942038 |
| 2 | 0 | 0.455977 | 0.0808971 | 0.0550434 |
| 3 | 0.024486 | 0 | 0.455977 | 0 |
| 4 | 0 | 0.455977 | 0 | 0.175581 |
| 5 | 0.455977 | 0 | 0.0549319 | 0 |
| 6 | 0.162363 | 0.455977 | 0.0897636 | 0.0689181 |
| 7 | 0 | 0.02975 | 0.455977 | 0.0170034 |
| 8 | 0.455977 | 0 | 0 | 0.0975597 |
| 9 | 0.134079 | 0.455977 | 0 | 0 |
| 10 | 0.486152 | 0.455977 | 0.0220095 | 0 |
| 11 | 0 | 0.0246322 | 0.0847586 | 0.455977 |
| 12 | 0.455977 | 0 | 0 | 0.0321177 |
| 13 | 0.20916 | 0.455977 | 0 | 0.0482279 |
| 14 | 0.455977 | 0 | 0.0837475 | 0.141681 |
| 15 | 0 | 0.107215 | 0 | 0.455977 |
| 16 | 0 | 0 | 0.269667 | 0.455977 |
| 17 | 0.0634094 | 0 | 0.455977 | 0.0582705 |
Each row represents the changes in binding energy, ΔΔG, compared to the reference sequence upon the substitution to the indicated nucleotide at certain position within the target DNA element. Values are indicated in kcal/mol.
Uncharacterized Mutant TetRs
Unfortunately we did not have time to finish the characterization of the following parts:
Y42F K108E
V36F W43S
P39Q Y42M L197S
P39Q Y42M L52P
E37A P39K
E37A P39Q Y42F
 "
"