Team:UNICAMP-EMSE Brazil/Methodology
From 2011.igem.org
Neshich.iap (Talk | contribs) (→Part descriptions) |
Neshich.iap (Talk | contribs) (→Part descriptions) |
||
| Line 51: | Line 51: | ||
<br clear="all"> | <br clear="all"> | ||
| - | '''<font size=3 | + | <div align=center> |
| - | + | '''<font size=3>In our project, we also used the following biobricks from the 2011´s biobrick collection distribution sent by the iGEM headquarters:</font>''' | |
| + | </div> | ||
<br clear="all"> | <br clear="all"> | ||
[[Image:UNICAMP_EMSE_const.jpg|left|100px]] <br><br>Constitutive promoter, part [http://partsregistry.org/Part:BBa_J23119 BBa_J23119] | [[Image:UNICAMP_EMSE_const.jpg|left|100px]] <br><br>Constitutive promoter, part [http://partsregistry.org/Part:BBa_J23119 BBa_J23119] | ||
Revision as of 19:46, 28 September 2011

| Home | Project | Methods | Results | Data | Team | Notebook | Human Practices | Safety | Profile | Sponsors | Wix |

Contents |
Creating a great WARRIOR!
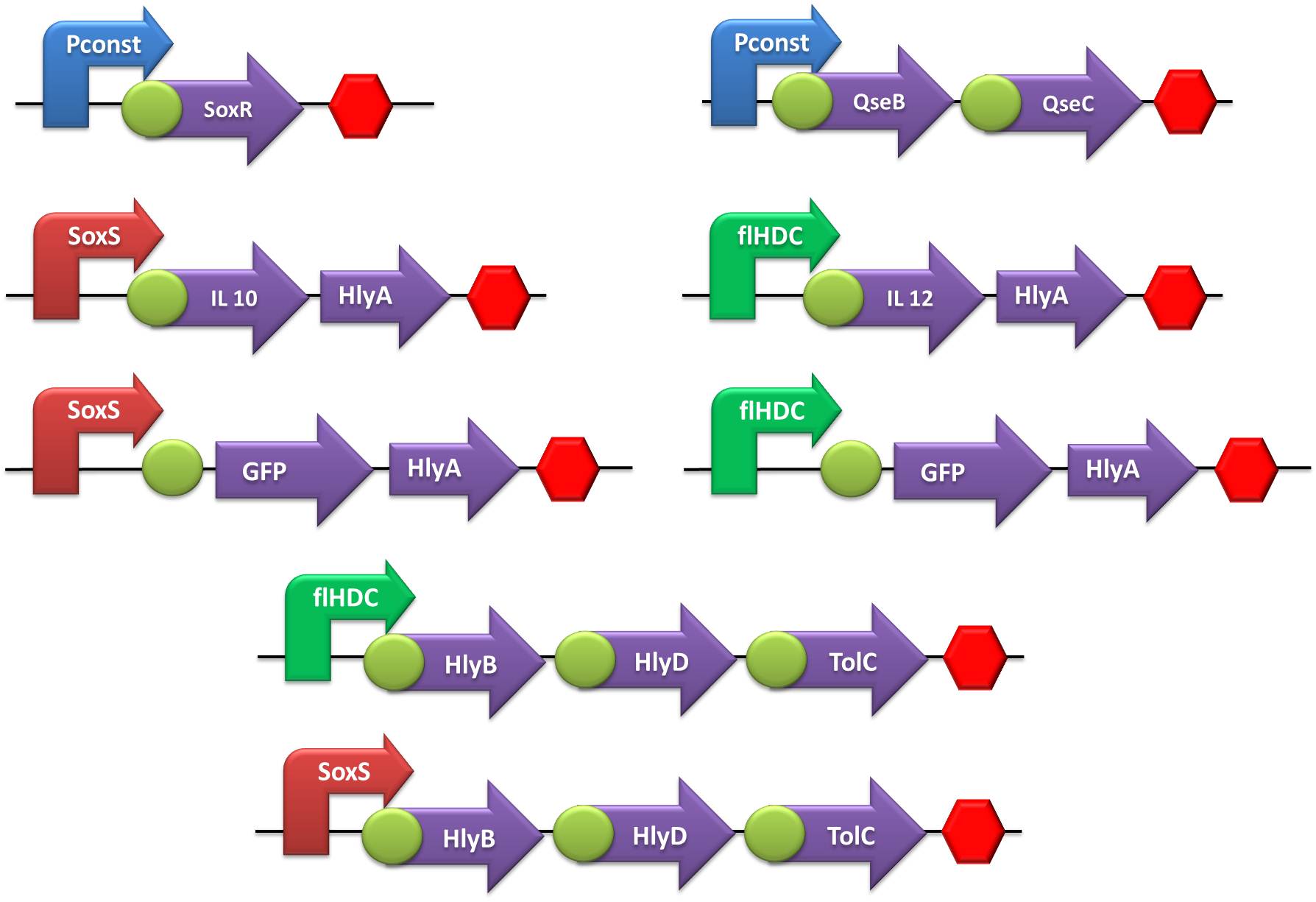
In order to create our “Jedi coli” to fight the STRESS WARS, we trained padawan colis to use their power to sense Nitric Oxide (NO) or Adrenaline and produce IL-10 and IL-12, respectively, to defend or bodyConfederation host against inVader bacteria. By the end of the harsh Jedi training, which included hours of cutting and ligation, cloning and transformation, we aimed to develop the following gene constructions:
Work plan:
Our work was divided in three main steps:
- Biobrick assembly;
- Suit the biobrick for shipment (remove them from the original plasmid and insert them on pSB1C3 plasmid);
- Devices construction and testing.
First step: Biobricks Assembly
Part descriptions
For the reasons explained in the safety proposals, we synthesized the following gene sequences according to assembling standard 23 and in the format of transcriptional units [with RBS ([http://partsregistry.org/Part:BBa_B0030 B0030]) and stop codon, when protein coding:
flhDC promoter, part [http://partsregistry.org/wiki/index.php?title=Part:BBa_K554001 BBa_K554001]
RBS+SoxR gene, part [http://partsregistry.org/wiki/index.php?title=Part:BBa_K554003 BBa_K554003]
RBS+IL-12 gene, part [http://partsregistry.org/wiki/index.php?title=Part:BBa_K554005 BBa_K554005]
RBS+Hemolysin D(HlyD) gene, part [http://partsregistry.org/wiki/index.php?title=Part:BBa_K554008 BBa_K554008]
RBS+QseB gene+QseC gene, part [http://partsregistry.org/wiki/index.php?title=Part:BBa_K554006 BBa_K554006]
In our project, we also used the following biobricks from the 2011´s biobrick collection distribution sent by the iGEM headquarters:
Constitutive promoter, part [http://partsregistry.org/Part:BBa_J23119 BBa_J23119]
Double terminator, part [http://partsregistry.org/Part:BBa_B0014 BBa_B0014]
Description of the assembly process:
- NO and Adrenaline sensors:
- The first step was to ligate both SoxR and QseB/QseC transcriptional units with the terminator, followed by the ligation of these constructs to the constitutive promoter.
- IL-10, IL-12 and GFP expressing units:
- Terminator sequence was ligated in the Hemolysin A sequence, followed by the fusion of this construction with the IL-10, IL-12 or GFP transcriptional units (GFP one was first coupled with RBS sequence). The final biobrick was obtained after the coupling of interleukin or GFP transcription units (+ Hemolysin A + Terminator) with the respective promoters.
- Secretion systems:
- The first round of ligation was accomplished by coupling the Terminator with the TolC transcriptional unit. The subsequent ligation round comprised the ligation of HlyD in the TolC+Terminator construct, while HlyB was ligated in the promoters. The last assembling step was accomplished by coupling the SoxS or flHDC promoter+HlyB with the previously assembled construction HlyD+TolC+T.
 "
"