Team:Freiburg/Notebook/9 September
From 2011.igem.org
(Difference between revisions)
SophieCramer (Talk | contribs) (→blue light receptor) |
|||
| (14 intermediate revisions not shown) | |||
| Line 1: | Line 1: | ||
{{:Team:Freiburg/Templates/header}} | {{:Team:Freiburg/Templates/header}} | ||
| + | <html> | ||
| + | <div id="notebook-page-header"> | ||
| + | <div id="notebook-back" width="100px" > | ||
| + | <a href="https://2011.igem.org/Team:Freiburg/Notebook/8_September">Previous entry</a> | ||
| + | </div> | ||
| + | <div id="notebook-title"> | ||
| + | <a href="https://2011.igem.org/Team:Freiburg/Notebook"> 9 September </a> | ||
| + | </div> | ||
| + | <div id="notebook-next"> | ||
| + | <a href="https://2011.igem.org/Team:Freiburg/Notebook/10_September">Next entry</a> | ||
| + | </div> | ||
| + | </div> | ||
| + | </html> | ||
==Commons== | ==Commons== | ||
| Line 7: | Line 20: | ||
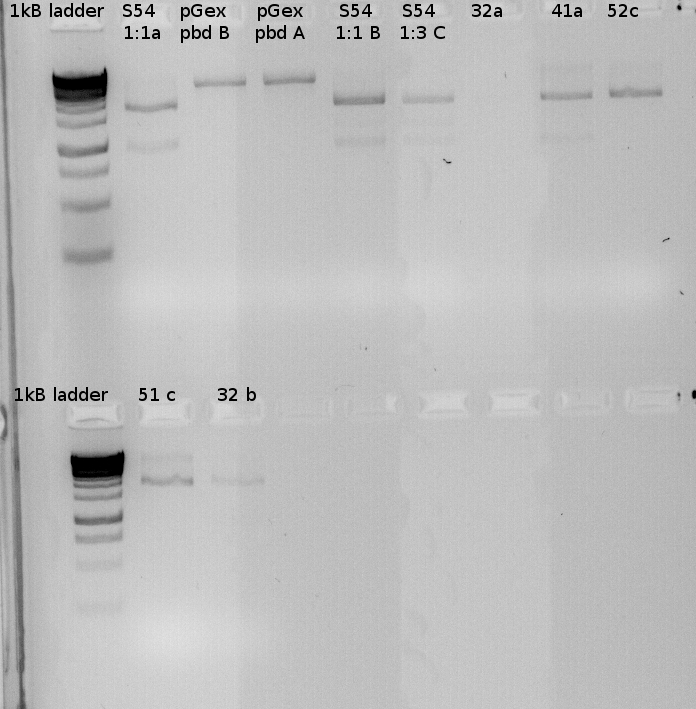
Testdigest of our various minipreps | Testdigest of our various minipreps | ||
| - | [[File:100Testdigests_Freiburg.jpg]] | + | [[File:100Testdigests_Freiburg.jpg |350px|350px]] |
| + | [[File:100Testdigests_Freiburg1.jpg| 350px|350px]] | ||
==<span style="color:green;">green light receptor</span>== | ==<span style="color:green;">green light receptor</span>== | ||
| - | === | + | ===digest for cloning PcpcG infront of CFP/YFP=== |
| - | '''Investigators: | + | '''Investigators:Julia''' |
| + | <br/> | ||
| + | <br/> | ||
| + | 1. Digestion of PCR Product(PcpcG)<br/> | ||
| + | <br/> | ||
| + | 38µl PCR product<br/> | ||
| + | 5µl BSA (10x)<br/> | ||
| + | 5µl NEB buffer 4<br/> | ||
| + | 1µl EcoRI<br/> | ||
| + | 1µl SpeI<br/> | ||
| + | <br/> | ||
| + | 2.Digestion of CFP/YFP Vector | ||
| + | 36,1/34 µl water<br/> | ||
| + | 1.9µl DNA of YFP/ 3.7µl DNA of CFP = 500ng DNA<br/> | ||
| + | 5µl BSA (10x)<br/> | ||
| + | 5µl NEB buffer 4<br/> | ||
| + | 1µl EcoRI<br/> | ||
| + | 1µl XbaI<br/> | ||
| + | |||
| + | Incubation over night. | ||
==<span style="color:blue;">blue light receptor</span>== | ==<span style="color:blue;">blue light receptor</span>== | ||
| - | '''PCR''' | + | |
| + | ===Ligation=== | ||
| + | |||
| + | |||
| + | {| style="border-spacing:0;" | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| Name: Rüdiger | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| Date: 09.09. | ||
| + | |||
| + | |- | ||
| + | | colspan="2" style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| Continue from Date 05.09. Name Rüdiger | ||
| + | |||
| + | Experiment Digestion | ||
| + | |||
| + | |- | ||
| + | | colspan="2" style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| Project Name: Precipitator | ||
| + | |||
| + | |} | ||
| + | '''Procedure''' | ||
| + | |||
| + | |||
| + | PCR tube: | ||
| + | |||
| + | total volume 20 μl | ||
| + | |||
| + | |||
| + | # add H<sub>2</sub>O (17 μl -X-Y-Z) | ||
| + | # add 2 μl Ligase Buffer 10x | ||
| + | # add Insert 1, Insert 2(when proceeding from 3A digestion use 2 μl of each) | ||
| + | # add Vector (20ng needed. When proceeding from 3A digestion use 2 μl) | ||
| + | # Add 1 μl T4-DNA Ligase | ||
| + | # Incubate 10-30 min at room temperature | ||
| + | # heat for 20 minutes at 80°C | ||
| + | # store at -20°C or directly proceed to transformation | ||
| + | |||
| + | |||
| + | {| style="border-spacing:0;" | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| Name of part | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| Ratio Insert:Vector | ||
| + | |||
| + | <nowiki>= 3:1 or 1:1</nowiki> | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| Volume (μl) | ||
| + | |||
| + | |- | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| X insert 1 | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| | ||
| + | |||
| + | |- | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| Y insert 2 | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| | ||
| + | |||
| + | |- | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| Z vector | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| | ||
| + | |||
| + | |- | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| H<sub>2</sub>O | ||
| + | | style="background-color:#bfbfbf;border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| | ||
| + | | style="background-color:#bfbfbf;border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| | ||
| + | |||
| + | |} | ||
| + | '''Documentation:''' | ||
| + | |||
| + | Why are you doing this experiment? Where are your parts stored? Name the parts for ligation etc. | ||
| + | |||
| + | |||
| + | {| style="border-spacing:0;" | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| | ||
| + | {| style="border-spacing:0;" | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| All vectors from 05.05. digest | ||
| + | |||
| + | |||
| + | |||
| + | {| style="border-spacing:0;" | ||
| + | | style="border-top:0.0007in solid #000000;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| Resistence | ||
| + | | style="border-top:0.0007in solid #000000;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| Name | ||
| + | | style="border-top:0.0007in solid #000000;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| Vector | ||
| + | | colspan="2" style="border:0.0007in solid #000000;padding:0.0382in;"| Insert | ||
| + | |||
| + | |- | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| AMP | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| 1a | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| pGEX 1-3V | ||
| + | | colspan="2" style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:0.0007in solid #000000;padding:0.0382in;"| 1 | ||
| + | |||
| + | |- | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| 2a | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| pGEX 1-3V | ||
| + | | colspan="2" style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:0.0007in solid #000000;padding:0.0382in;"| 2 | ||
| + | |||
| + | |- | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| 3a | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| pGEX 1-3V | ||
| + | | colspan="2" style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:0.0007in solid #000000;padding:0.0382in;"| 3 | ||
| + | |||
| + | |- | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| C3 | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| 4 | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| C3 4,8-10 | ||
| + | | colspan="2" style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:0.0007in solid #000000;padding:0.0382in;"| 4 | ||
| + | |||
| + | |- | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| AMP | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| 5a | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| PET DUET 5-7 | ||
| + | | colspan="2" style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:0.0007in solid #000000;padding:0.0382in;"| 5 | ||
| + | |||
| + | |- | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| 6a | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| PET DUET 5-7 | ||
| + | | colspan="2" style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:0.0007in solid #000000;padding:0.0382in;"| 6 | ||
| + | |||
| + | |- | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| 7a | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| PET DUET 5-7 | ||
| + | | colspan="2" style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:0.0007in solid #000000;padding:0.0382in;"| 7 | ||
| + | |||
| + | |- | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| C3 | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| 8 | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| C3 4,8-10 | ||
| + | | colspan="2" style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:0.0007in solid #000000;padding:0.0382in;"| 8 | ||
| + | |||
| + | |- | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| 9 | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| C3 4,8-10 | ||
| + | | colspan="2" style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:0.0007in solid #000000;padding:0.0382in;"| 9 | ||
| + | |||
| + | |- | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| 10 | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| C3 4,8-10 | ||
| + | | colspan="2" style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:0.0007in solid #000000;padding:0.0382in;"| 10 | ||
| + | |||
| + | |- | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| AMP | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| 1b | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| pGEX 1-3V | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| 1 | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:0.0007in solid #000000;padding:0.0382in;"| Inserts from 31.08. digest | ||
| + | |||
| + | |- | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| 2b | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| pGEX 1-3V | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| 2 | ||
| + | |||
| + | |- | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| 3b | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| pGEX 1-3V | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| 3 | ||
| + | |||
| + | |- | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| 5b | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| PET DUET 5-7V | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| 6 | ||
| + | |||
| + | |} | ||
| + | |||
| + | |||
| + | |} | ||
| + | |||
| + | |||
| + | |} | ||
| + | |||
| + | |||
| + | |||
| + | ===Gibson assembly=== | ||
| + | {| style="border-spacing:0;" | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| Name | ||
| + | |||
| + | Sandra, Sophie | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| Date: | ||
| + | |||
| + | 26.07.2011 | ||
| + | |||
| + | |- | ||
| + | | colspan="2" style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| Continue from Experiment (Date) from PCR (Lovtap 3000bp and Not-Gate 980bp) from today. | ||
| + | |||
| + | (Name) Sophie | ||
| + | |||
| + | |- | ||
| + | | colspan="2" style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| Project Name: Blue light receptor: Ligation of Lovtap and Not-Gate | ||
| + | |||
| + | |} | ||
| + | Gibson-Assembly | ||
| + | |||
| + | |||
| + | 1. Prepare 5X ISO buffer. Six ml of this buffer can be prepared by combining the following: | ||
| + | |||
| + | 3 ml of 1 M Tris-HCl pH 7.5150 μl of 2 M MgCl<sub>2</sub>60 μl of 100 mM dGTP 60 μl of 100 mM dATP 60 μl of 100 mM dTTP60 μl of 100 mM dCTP300 μl of 1 M DTT 1.5 g PEG-8000300 μl of 100 mM NADAdd water to 6 ml Aliquot 100 μl and store at -20 °C | ||
| + | |||
| + | 2. Prepare an assembly master mixture. This can be prepared by combining the following: | ||
| + | |||
| + | 320 μl 5X ISO buffer0.64 μl of 10 U/ μl T5 exo20 μl of 2 U/μl Phusion pol160 μl of 40 U/μl Taq ligAdd water to 1.2 ml | ||
| + | |||
| + | Aliquot 15 μl and store at -20 °C. This assembly mixture can be stored at -20 °C for at least one year. The enzymes remain active following at least 10 freeze-thaw cycles. This is ideal for the assembly of DNA molecules with 20-150 bp overlaps. For DNA molecules overlapping by larger than 150 bp, prepare the assembly mixture by using 3.2 μl of 10 U/ μl T5 exo. | ||
| + | |||
| + | 3. Thaw a 15 μl assembly mixture aliquot and keep on ice until ready to be used. | ||
| + | |||
| + | 4. Add 5 μl of DNA to be assembled to the master mixture. The DNA should be in equimolar amounts. Use 10-100 ng of each ~6 kb DNA fragment. For larger DNA segments, increasingly proportionate amounts of DNA should be added (e.g. 250 ng of each 150 kb DNA segment). | ||
| + | |||
| + | 5. Incubate at 50 °C for 15 to 60 min (60 min is optimal). | ||
| + | |||
| + | 6. If cloning is desired, electroporate 1 μl of the assembly reaction into 30 μl electrocompetent ''E. coli''. | ||
| + | |||
| + | '''Documentation:''' | ||
| + | |||
| + | Why are you doing this experiment? Name the parts for the Gibson-Assembly. | ||
| + | |||
| + | |||
| + | {| style="border-spacing:0;" | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| Parts for Gibson-Assembly: G-♥ and G-NOT | ||
| + | |||
| + | |} | ||
| + | How did you label your samples and where are they stored? | ||
| + | |||
| + | |||
| + | {| style="border-spacing:0;" | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| Labelled G-♥-NOT and G-♥-NOT 50 | ||
| + | |||
| + | |} | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | ===PCR=== | ||
| Line 95: | Line 360: | ||
stored in blue light box | stored in blue light box | ||
| - | + | ||
| + | |||
| + | ===Digestion=== | ||
| Line 225: | Line 492: | ||
|} | |} | ||
| - | |||
| - | === | + | ===Ligation=== |
| - | |||
| + | {| style="border-spacing:0;" | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| Name: Sophie | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| Date: 9.9.11 | ||
| + | |- | ||
| + | | colspan="2" style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| Continue from Date:9.9.11 Name: Sophie | ||
| + | Experiment: Digestion | ||
| + | |||
| + | |- | ||
| + | | colspan="2" style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| Project Name: G-♥-NOT in PR-vector | ||
| + | |||
| + | |} | ||
| + | '''Procedure''' | ||
| + | |||
| + | |||
| + | PCR tube: | ||
| + | |||
| + | total volume 20 μl | ||
| + | |||
| + | |||
| + | # add H<sub>2</sub>O (17 μl -X-Y-Z) | ||
| + | # add 2 μl Ligase Buffer 10x | ||
| + | # add Insert 1, Insert 2(when proceeding from 3A digestion use 2 μl of each) | ||
| + | # add Vector (20ng needed. When proceeding from 3A digestion use 2 μl) | ||
| + | # Add 1 μl T4-DNA Ligase | ||
| + | # Incubate 10-30 min at room temperature | ||
| + | # heat for 20 minutes at 80°C | ||
| + | # store at -20°C or directly proceed to transformation | ||
| + | |||
| + | |||
| + | {| style="border-spacing:0;" | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| Name of part | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| Ratio Insert:Vector | ||
| + | |||
| + | <nowiki>= 3:1 or 1:1</nowiki> | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| Volume (μl) | ||
| + | |||
| + | |- | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| X insert 1 | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| G-♥-NOT insert a,b | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| | ||
| + | |||
| + | |- | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| Z vector | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| D39 -D44 | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| | ||
| + | |||
| + | |- | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| H<sub>2</sub>O | ||
| + | | style="background-color:#bfbfbf;border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| | ||
| + | | style="background-color:#bfbfbf;border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| | ||
| + | |||
| + | |} | ||
| + | '''Documentation:''' | ||
| + | |||
| + | Why are you doing this experiment? Where are your parts stored? Name the parts for ligation etc. | ||
| + | |||
| + | |||
| + | {| style="border-spacing:0;" | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| Ligated parts labeled L39 -L44 (red pencil) | ||
| + | |||
| + | stored in blue light box | ||
| + | |||
| + | |} | ||
==<span style="color:orange;">Lysis cassette</span>== | ==<span style="color:orange;">Lysis cassette</span>== | ||
| - | === | + | ===Troubleshooting of the modified Lysis genes K124017=== |
| - | '''Investigators: | + | '''Investigators:Theo''' |
| + | The Ligated parts (M48+1 and M48+7) could not be transformed so it was decided to try again and wait until Monday for the sequencing.<br> | ||
| + | In the meantime, | ||
| + | <br> | ||
| + | |||
| + | ===M48 is in an Amp Vector, part has to be sent to registry=== | ||
| + | The M48 stock was inoculated so that it could be mini-preped on Saturday | ||
| + | |||
| + | <br> | ||
| + | <br> | ||
==<span style="color:grey;">Precipitator</span>== | ==<span style="color:grey;">Precipitator</span>== | ||
| + | |||
| + | ===Transformation=== | ||
| + | |||
| + | |||
| + | {| style="border-spacing:0;" | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| Name: Rüdiger | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| Date: 09.09. | ||
| + | |||
| + | |- | ||
| + | | colspan="2" style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| Continue from Date 09.09. Name Rüdiger | ||
| + | |||
| + | Experiment Ligation | ||
| + | |||
| + | |- | ||
| + | | colspan="2" style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| Project Name: Precipitator | ||
| + | |||
| + | |} | ||
| + | Procedure | ||
| + | |||
| + | |||
| + | # take cells from -80°C freezer and put them on ice! (every eppi contains about 400 μl cells) | ||
| + | # thaw cells on ice 20 minutes | ||
| + | # pipette 50 μl cells and 2 μl DNA into eppi still on ice! | ||
| + | # Incubate for 30 minutes on ice | ||
| + | # Heat at 42°C for 60 sec | ||
| + | # Incubate on ice for 5 minutes | ||
| + | # Add 200 μl LB Broth | ||
| + | # Incubate for 2 hours at 37°C (cells with lysis cassette at 30°C!!) | ||
| + | # Plate 50 μl and 200μl on two different LB/Agar plates with appropriate antibiotic resistance | ||
| + | |||
| + | '''Documentation:''' | ||
| + | |||
| + | Why are you doing this experiment? Name of the sample? Where are they stored? Name the vector with inserts, antibiotika resistance etc. | ||
| + | |||
| + | |||
| + | Had no C3 plates, took AMP plates only | ||
| + | |||
| + | |||
| + | Picked cells from yesterday's Transformation | ||
| + | |||
| + | |||
| + | 4a/b 1-4 | ||
| + | |||
| + | 8a/b 1-4 | ||
| + | |||
| + | 9a/b 1-4 | ||
| + | |||
| + | 10a/b 1-4 | ||
| + | |||
===NAME OF YOUR EXPERIMENT=== | ===NAME OF YOUR EXPERIMENT=== | ||
Latest revision as of 00:24, 22 September 2011
 "
"
 Contact
Contact