Team:EPF-Lausanne/Tools/MITOMI
From 2011.igem.org
(→MITOMI scans explanation) |
(→MITOMI scans explanation) |
||
| Line 37: | Line 37: | ||
[[File:EPFL2011_MITOMIchip_afterwashDNA_illustration.png|300px| After PBS wash: GFP fluorescence ]] | [[File:EPFL2011_MITOMIchip_afterwashDNA_illustration.png|300px| After PBS wash: GFP fluorescence ]] | ||
| + | For data collection the fluorescence intensity measured under the button is normalised to the background and the DNA/protein ratio is calculated. This ratio is ploted against the intensity of the fluorescence of DNA in solution to make the saturation curves. | ||
| + | |||
| + | [[File:EPFL2011_MITOMI_data_collection_points.JPG|200px]] | ||
{{:Team:EPF-Lausanne/Templates/Footer}} | {{:Team:EPF-Lausanne/Templates/Footer}} | ||
Revision as of 21:37, 21 September 2011
MITOMI
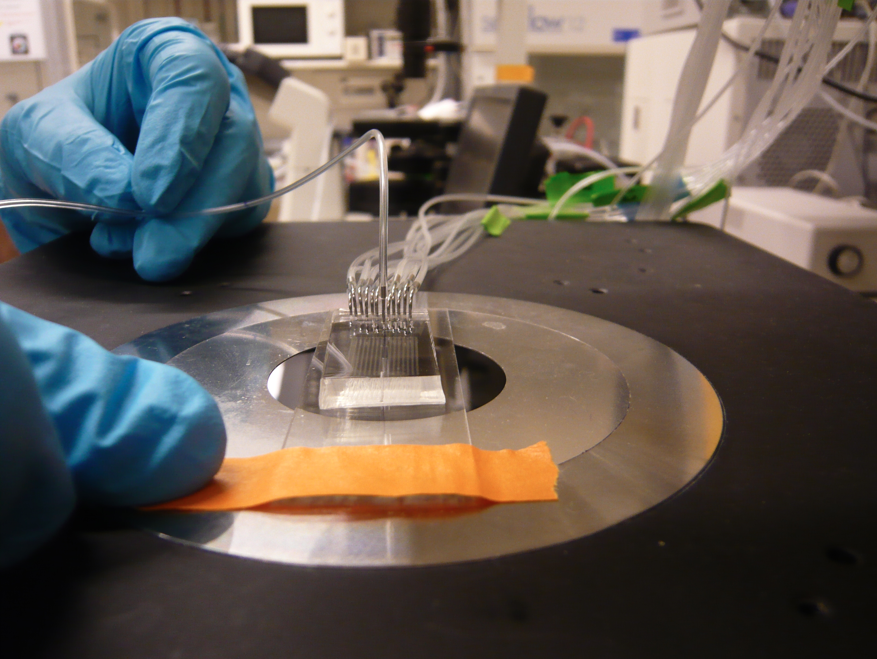
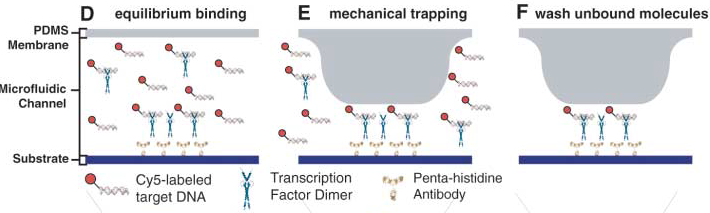
Mitomi is a high troughput microfluidic system designed to mesure low affinity molecular interactions by mechanically trapping them.
Purpose: We applied this technology during our project for characterizing transcription factor binding energy landscapes in absolute terms by determining dissociation constants (Kd) for a comprehensive set of target DNA sequences.
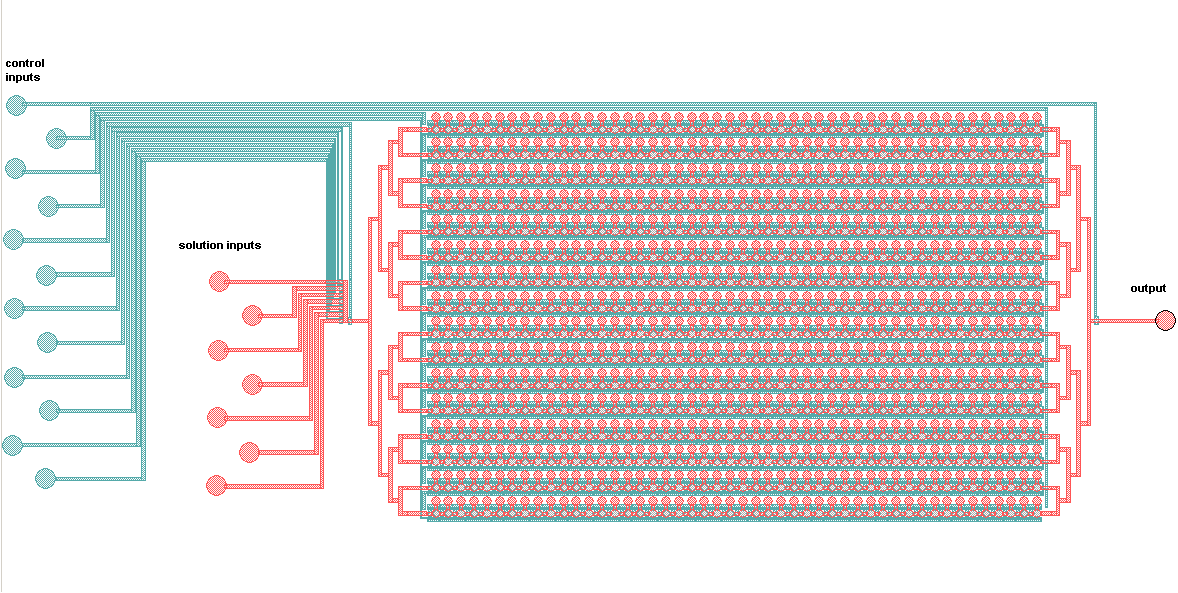
Schematic of the Mitomi platform:
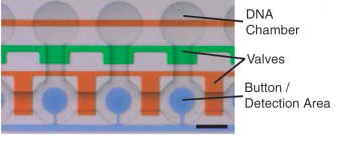
The control inputs control the valves that close the Detailed view of the chambers and of the movable membrane used for mechanical trapping, referred to as the 'Button' .
Advantages: The movable memrane shown in the scheme below enables measurements of transient interactions, exhibiting sub micromolar affinities which are generally problematic.
Running a MITOMI experiment
MITOMI scans explanation
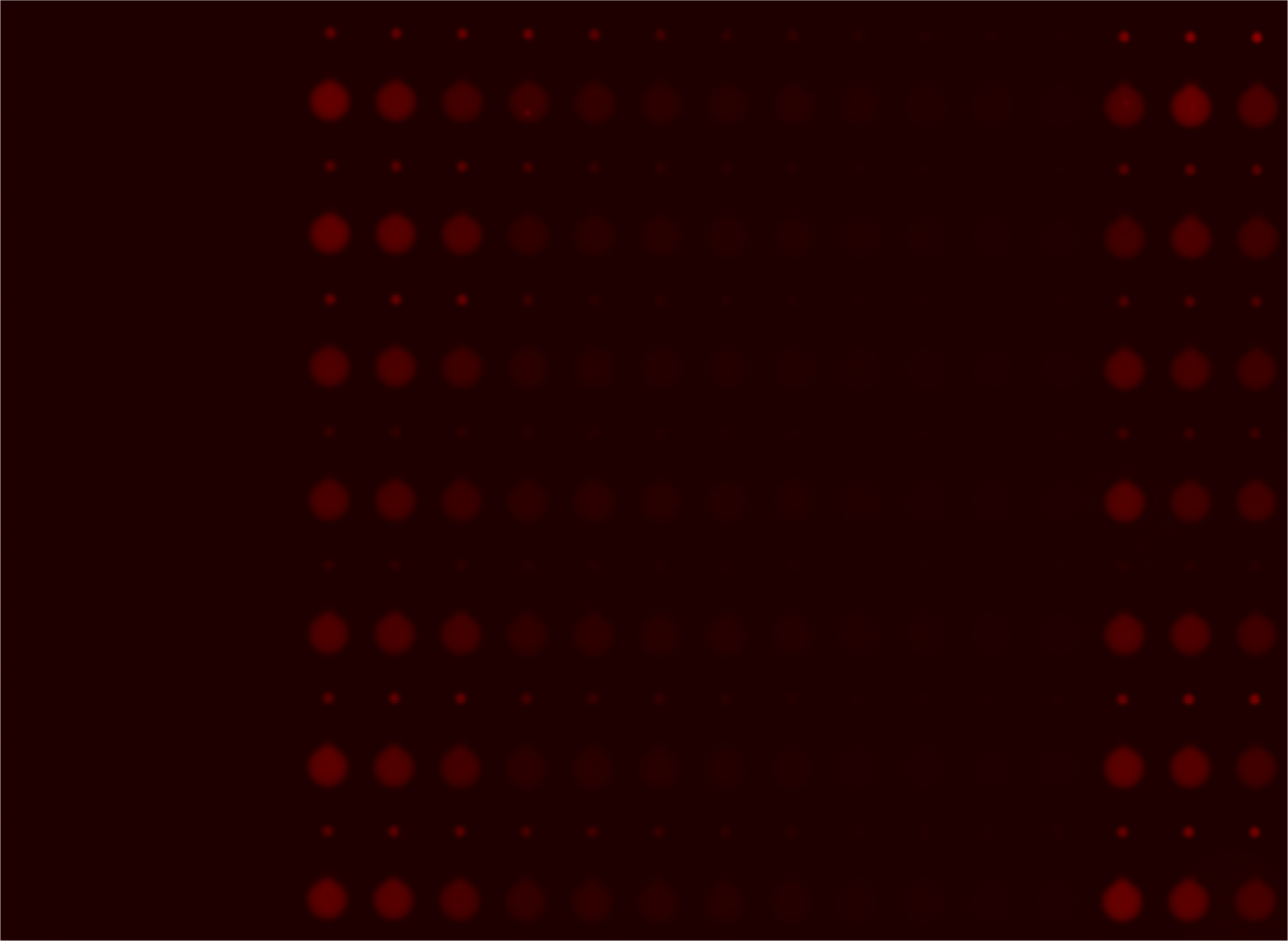
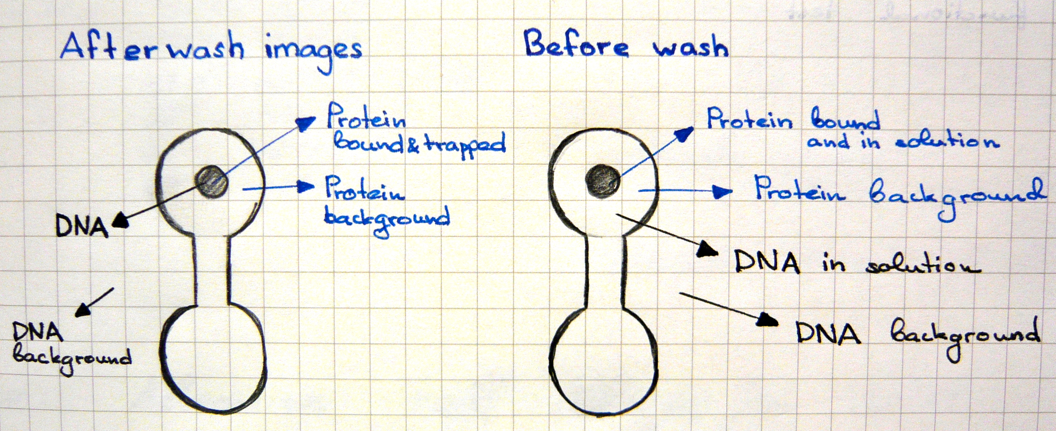
Two scans of the chip are necessary for the analysis of the molecular interactions.
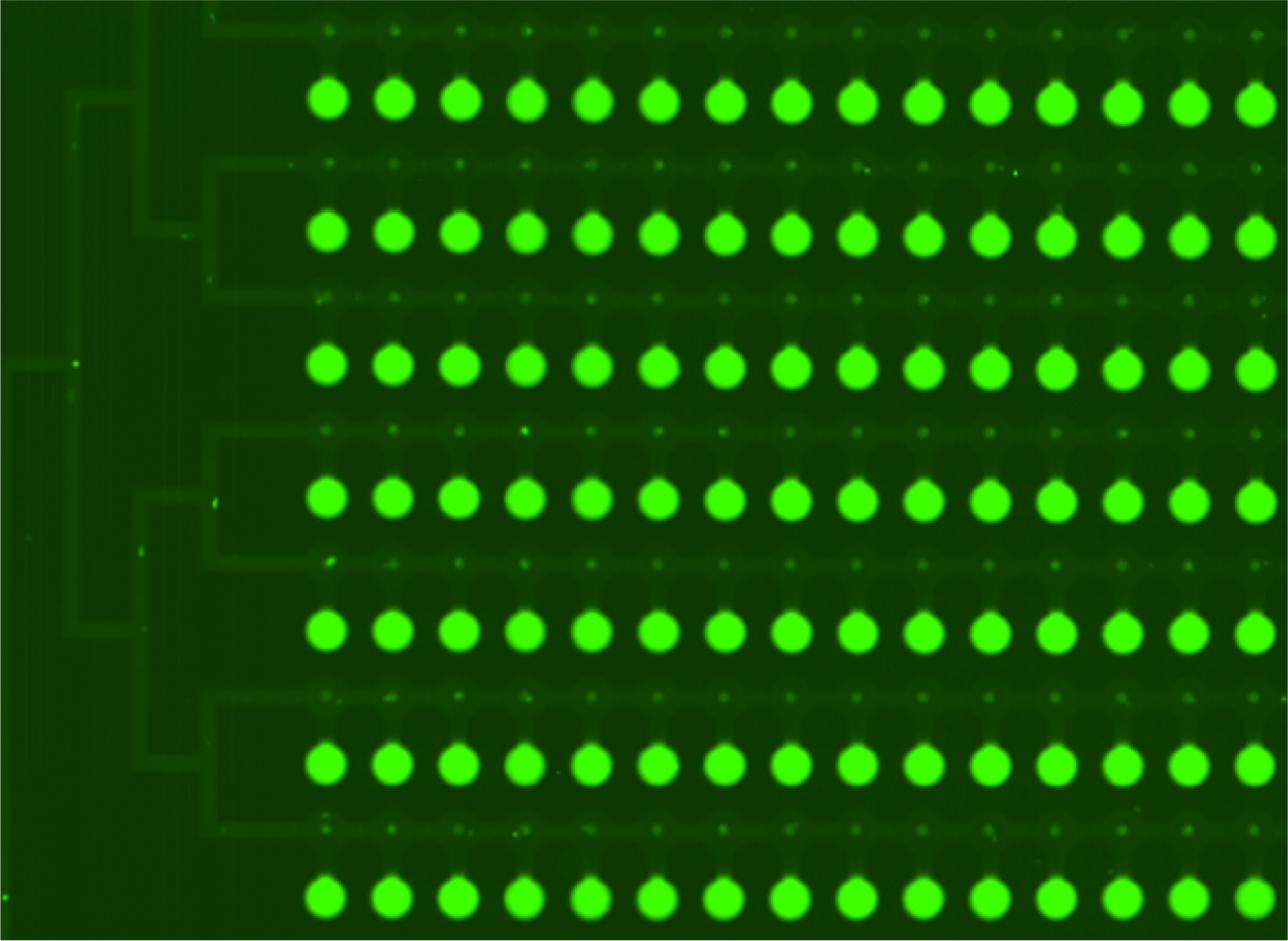
- First scan is taken after 30-60 minutes of incubation, when the protein-DNA interactions reach thermodynamic stability. At this point the chamber containing DNA is open to allow diffusion and the valves separating each of the 756 units are closed, which prevents contamination.
- Second scan is taken after 10-15 minutes of wash with PBS.
For data collection the fluorescence intensity measured under the button is normalised to the background and the DNA/protein ratio is calculated. This ratio is ploted against the intensity of the fluorescence of DNA in solution to make the saturation curves.
 "
"