Team:DTU-Denmark/Background the natural system
From 2011.igem.org
(→The natural chitobiose system) |
(→The natural chitobiose system) |
||
| Line 3: | Line 3: | ||
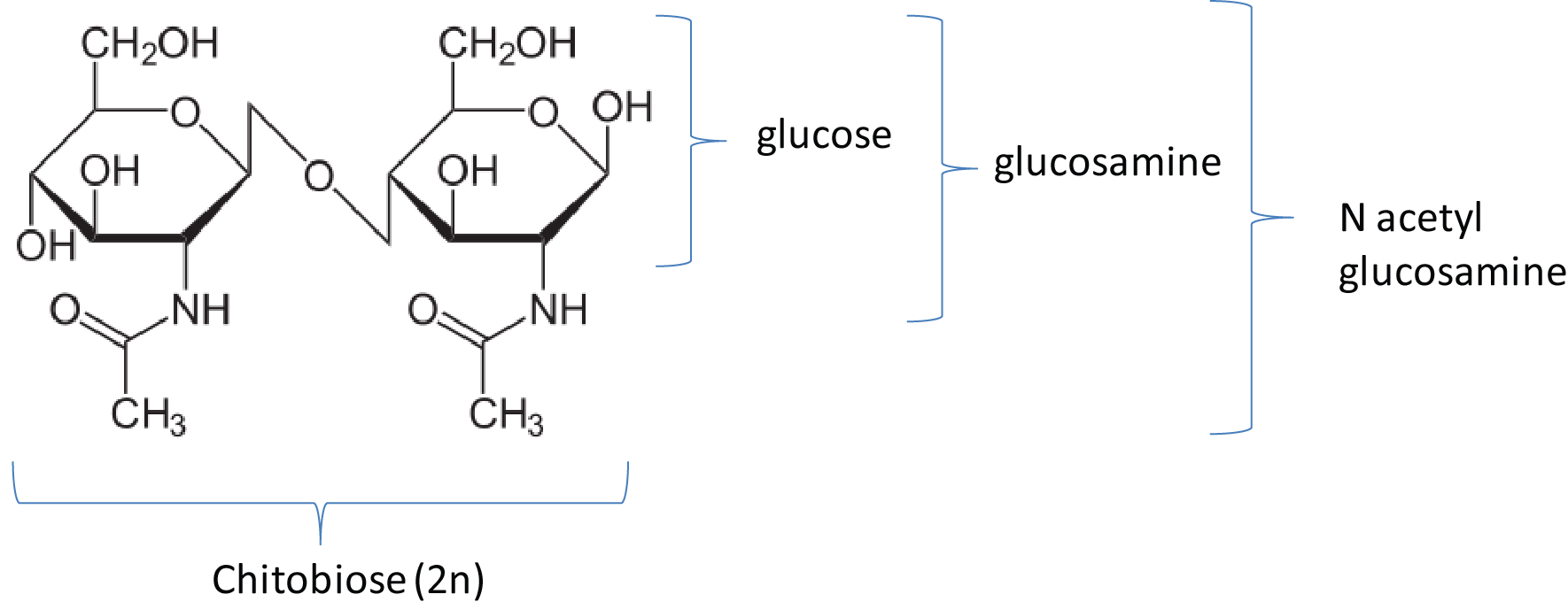
| - | [[File:DTU1_Chitobiose.png| | + | [[File:DTU1_Chitobiose.png|350px|thumb|left|Chitobiose; a disaccharide derived from chitin.]] |
The project is inspired by three components that regulate chitobiose transport in E. coli: (1) chitoporin ''chiP'' (alias ''ybfM'') that facilitates uptake of chitosugars from the medium into the cell; (2) '''sRNA''' ''chiX'' (alias ''sroB'', ''micM'') which regulates ''chiP'' post-transcriptionally; and (3) ''chiXR'' sRNA which is transcribed from intergenic region in ''chbBCARG'' operon and regulates ''chiX'' sRNA. ChiX sRNA regulates chiP expression through binding to the Shine-Dalgarno sequence on the ''chiP'' mRNA and thus inhibits recruitment of 30S ribosome subunit and translation cannot occur. Only when chitobiose is present in the medium can the ''chiXR'' be transcribed and subsequently its transcript can bind to ''chiX'' sRNA, releasing ''chiP'' repression. | The project is inspired by three components that regulate chitobiose transport in E. coli: (1) chitoporin ''chiP'' (alias ''ybfM'') that facilitates uptake of chitosugars from the medium into the cell; (2) '''sRNA''' ''chiX'' (alias ''sroB'', ''micM'') which regulates ''chiP'' post-transcriptionally; and (3) ''chiXR'' sRNA which is transcribed from intergenic region in ''chbBCARG'' operon and regulates ''chiX'' sRNA. ChiX sRNA regulates chiP expression through binding to the Shine-Dalgarno sequence on the ''chiP'' mRNA and thus inhibits recruitment of 30S ribosome subunit and translation cannot occur. Only when chitobiose is present in the medium can the ''chiXR'' be transcribed and subsequently its transcript can bind to ''chiX'' sRNA, releasing ''chiP'' repression. | ||
| + | |||
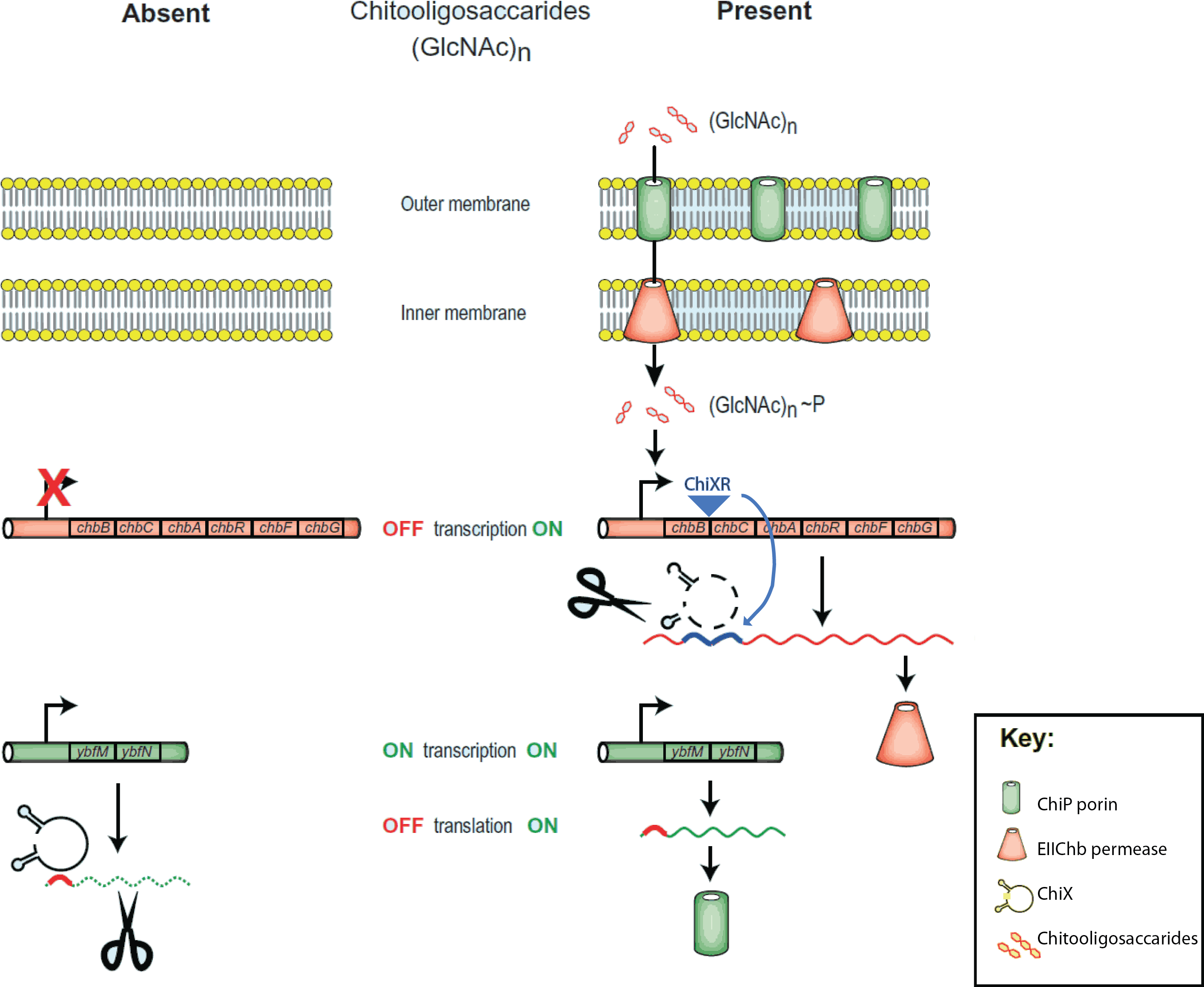
| + | [[File:Chitobiose_system.png|350px|thumb|right|The ''E.coli'' chitobiose system. Figure 5 from<span class="superscript">[[#References|[2]]]</span>]] | ||
| + | |||
Revision as of 13:07, 21 September 2011
Background
The natural chitobiose system
The project is inspired by three components that regulate chitobiose transport in E. coli: (1) chitoporin chiP (alias ybfM) that facilitates uptake of chitosugars from the medium into the cell; (2) sRNA chiX (alias sroB, micM) which regulates chiP post-transcriptionally; and (3) chiXR sRNA which is transcribed from intergenic region in chbBCARG operon and regulates chiX sRNA. ChiX sRNA regulates chiP expression through binding to the Shine-Dalgarno sequence on the chiP mRNA and thus inhibits recruitment of 30S ribosome subunit and translation cannot occur. Only when chitobiose is present in the medium can the chiXR be transcribed and subsequently its transcript can bind to chiX sRNA, releasing chiP repression.
References
[1] Figueroa-Bossi, Nara, Martina Valentini, Laurette Malleret, and Lionello Bossi. “Caught at its own game: regulatory small RNA inactivated by an inducible transcript mimicking its target.” Genes & Development 23, no. 17 (2009): 2004 -2015. http://genesdev.cshlp.org/content/23/17/2004.abstract.
[2] Overgaard, Martin, Jesper Johansen, Jakob Møller‐Jensen, and Poul Valentin‐Hansen. “Switching off small RNA regulation with trap‐mRNA.” Molecular Microbiology 73, no. 5 (September 2009): 790-800. http://onlinelibrary.wiley.com/doi/10.1111/j.1365-2958.2009.06807.x/abstract.
 "
"