Team:Washington/Parts
From 2011.igem.org
(→Diesel Production) |
|||
| Line 5: | Line 5: | ||
<center><gallery caption="An Overview of the 2011 UW iGEM Teams Summer Projects" widths="250px" heights="400px" perrow="3"> | <center><gallery caption="An Overview of the 2011 UW iGEM Teams Summer Projects" widths="250px" heights="400px" perrow="3"> | ||
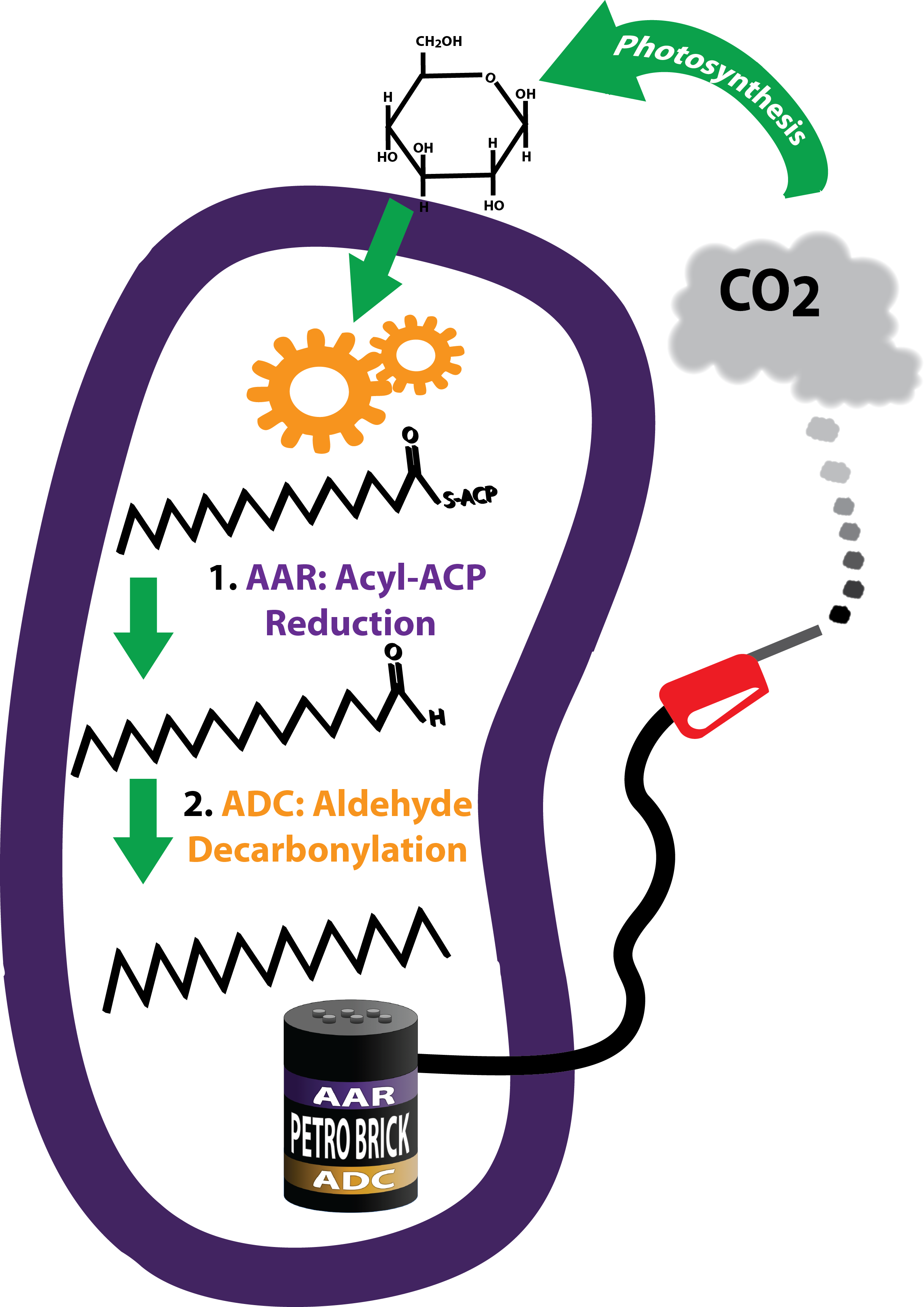
| - | Image:Diesel Production for Wiki.png|'''Make It: Diesel Production'''<br>We | + | Image:Diesel Production for Wiki.png|'''Make It: Diesel Production'''<br>We constructed and tested a modular alkane production BioBrick, the PetroBrick, designed to generate alkanes, the main constituent of diesel. |
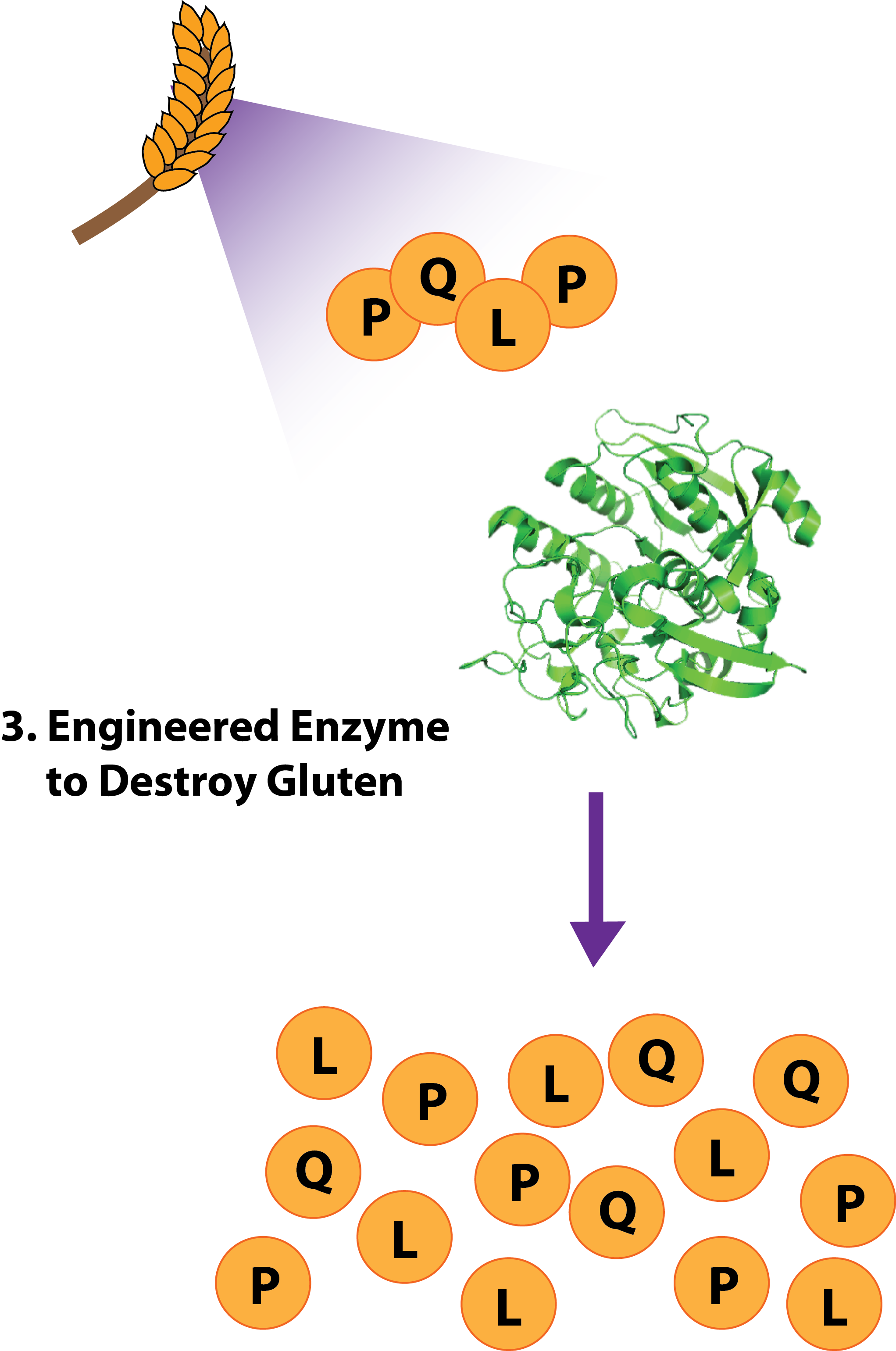
| - | Image:Gluten Destruction for Wiki.png|'''Break It: Gluten Destruction'''<br>Gluten intolerance | + | Image:Gluten Destruction for Wiki.png|'''Break It: Gluten Destruction'''<br>Gluten intolerance stems from an inappropriate immune response to PQLP, the most common motif in the immunogenic peptide. We reengineered a protease, active at low pH, for strongly enhanced activity against PQLP. |
Image:Gibson Assembly and Magnetosomes for Wiki.png|'''The GibsonBricks ToolKit'''<br>We completed two iGEM toolkits. In the first we created five Gibson Cloning versions of traditional biobrick vectors, and in the second we cloned the genes essential for magnetosome formation and transformed two into ''E. coli'' | Image:Gibson Assembly and Magnetosomes for Wiki.png|'''The GibsonBricks ToolKit'''<br>We completed two iGEM toolkits. In the first we created five Gibson Cloning versions of traditional biobrick vectors, and in the second we cloned the genes essential for magnetosome formation and transformed two into ''E. coli'' | ||
Revision as of 04:11, 23 September 2011
Data Summary
Data for Favorite New Parts
Diesel Production
A modular and open platform for the biological production of diesel fuel. The Petrobrick consists of AAR and ADC, each behind a standard Elowitz RBS. All of this in psb1C3 regulated by a high constitutive promoter.
Gluten Destruction
An enzyme from the sedolisin family native to Alicyclobacillus sendaiensis with known collagenase activity at low pH and elevated temperatures.
BBa_K590022: Kumamolisin-As_G319S, D358G, D368H
A mutated Kumamolisin-As enzyme aimed to combat gluten intolerance by increased activity with the PQLP peptide, an antigenic epitope in gliadin. This mutant has point mutations at residues 319, 358, and 368 from Glycine to Serine, Aspartate to Glycine, and Aspartate to Histidine, respectively.
BBa_K590023: Kumamolisin-As_N291D
A mutated Kumamolisin-As enzyme aimed to combat gluten intolerance by increased activity with the PQLP peptide, an antigenic epitope in gliadin. This mutant has a point mutation at residue 291 from Asparagine to Aspartate.
BBa_K590024: Kumamolisin-As_S354N, D358G, D368H
A mutated Kumamolisin-As enzyme aimed to combat gluten intolerance by increased activity with the PQLP peptide, an antigenic epitope in gliadin. This mutant has point mutations at residue 354, 358, and 368 from Serine to Asparagine, Aspartate to Glycine, and Aspartate to Histidine, respectively.
Gibson Assembly Toolkit
This is a high copy plasmid backbone which has ampicillin resistance, with pLac promoter and GFP.
This is a high copy plasmid backbone which has chloramphenicol resistance, with pLac promoter and GFP.
This is a low copy plasmid backbone which has chloramphenicol resistance, with pLac promoter and GFP.
This is a low copy plasmid backbone which has ampicillin resistance, with pLac promoter and GFP .
This is a medium copy plasmid backbone which has kanamycin resistance, with pLac promoter and GFP.
Magnetosome Toolkit
This part consists of mamK gene from Magnetospirillum magneticum strain AMB-1, sfGFP in the backbone of pGA1C3. mamK was previously reported for its importance proper magnetosome chain organization in magnetic bacteria; it is bacterial actin-like cytoskeleton protein required for proper alignment of the magnetosomes in a chain.
This part consists of mamI gene from Magnetospirillum magneticum strain AMB-1, sfGFP in the backbone of pGA1C3. MamI is a membrane-localized protein that localizes magnetic vesicles to the surface of cells, thus forming characteristic magnetosome chains.
Data for Existing Parts
Fill Me In
- Experience - Wintergreen odor enzyme generator, BBa_J45119 (MIT, iGEM 2006): 98 out of 100 volunteer subjects standing up to 5 feet away from the bacterial cultures could distinguish wintergreen-producing bacteria from negative controls.
- Experience - RBS, BBa_J61110 (Arkin Lab, 2007): Of the 5 RBS Parts we tested, this RBS works best for expressing yellow fluorescent protein-tagged BAR
Improved Parts
Fill Me In
- Main Page - Air Freshilizor, BBa_XXXXX: Our mathematical model predicts that the threshold of activation is 10 parts per billion, the concentration of Butanethiol that humans can typically smell
 "
"