Team:Washington/Alkanes/Future/LuxCDE
From 2011.igem.org
Alternative Routes for Fatty Aldehyde Production
Background
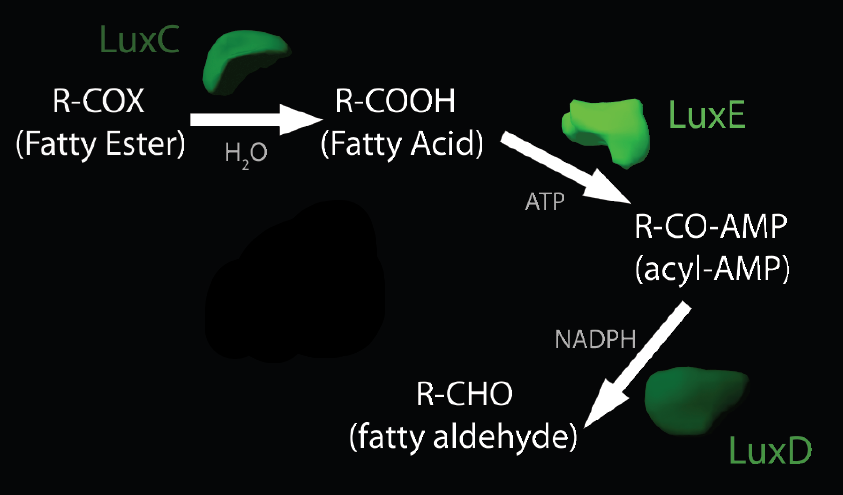
LuxABCDE is an enzyme complex submitted to the iGem registry by the 2010 Cambridge team. Within this complex, LuxAB produces luciferase (used by Cambridge’s EGlowi) using 14-carbon aldehydes (tetradecanal) produced by LuxCDE. Since the aldehyde decarbonylase (ADC) in our project uses aldehydes to synthesize alkanes, we plan to use the LuxCDE complex in conjunction with ADC to produce 13-carbon alkanes (Tridecane).
Methods
PCR and Standard Biobrick Assembly
This method revolved around designing oligos to PCR amplify LuxCD and LuxE off of the LuxBrick; LuxE was not directly adjacent to LuxCD in the Luxbrick, and therefore had to be amplified out in two separate reactions. Homology on the ends of the two fragments would then allow us to obtain a LuxCDE using stitching PCR, then ligate this fragment and ADC into psb1C3. We would then assay for alkanes.
Gibson Assembly
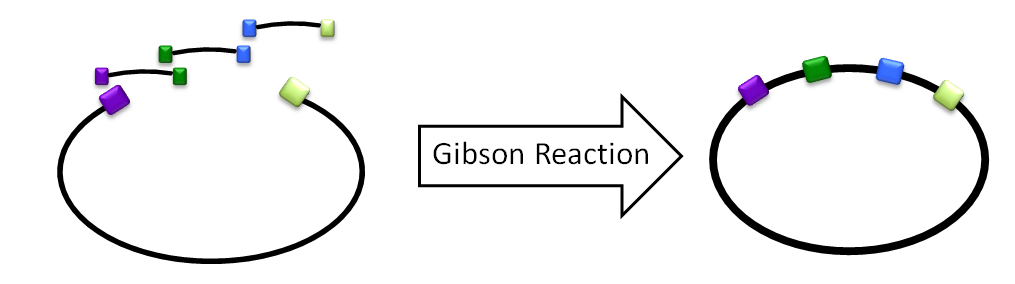
We would design oligos to amplify C, D, and E off of the LuxBrick. Unlike the previous method, the oligos would have sticky ends of [ ] base pairs to prevent misannealing, and the fragment containing C would also contain the vector for ligation. A 3-piece Gibson assembly was then done on C, D, and E, and the product was then transformed into cells.
Gene Synthesis
Instead of using the LuxBrick to obtain C, D, and E, we would re-optimize the codons on C, D, and E to be more cloning friendly, then assemble them from oligonucleotides. PCR was then used to amplify the respective pieces, and each of the three pieces was then ligated into psb1C3. The end result is each of the three genes separated out and with codons that have a normal GC content.
Results
The PCR and Gibson methods failed at multiple steps for uncertain reasons. Since we do not have our LuxCDE construct, no method or assay for testing the hypothetical construct was developed as of this time. We hypothesize that LuxCD's abnormally high AT concentration may have caused misannealing for primers, causing primer-dimer errors and other problems, but no concrete evidence exists for this as of yet. For gene assembly, we have lowered the AT concentration of C, D, and E to more optimal levels for PCR. To be precise, the C, D, and E had an AT content of 66.3, 65.0, and 66.9 percent, which was reduced to 50.2, 50.1, and 51.6 percent respectively.
[Still working on this, will be expanded once all experiments are complete]
Current Status
Attempts to clone LuxCDE using standard PCR/Biobrick Assembly and Gibson Assembly have been abandoned. Gene assembly was successful and each of the three luxCDE genes have been sequenced, biobricked and submitted.
Parts Submitted
[http://partsregistry.org/wiki/index.php?title=Part:BBa_K590059 LuxC], [http://partsregistry.org/wiki/index.php?title=Part:BBa_K590060 LuxD], [http://partsregistry.org/wiki/index.php?title=Part:BBa_K590061 LuxE]
Image(s)
Subsection will be deleted and intergrated into article once images are found.

 "
"