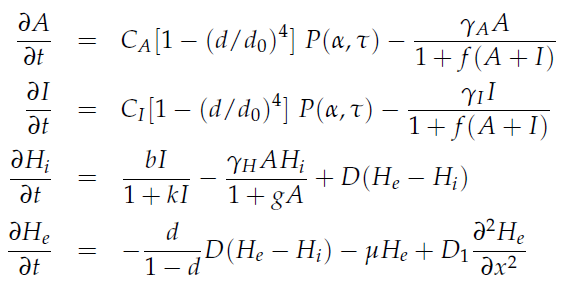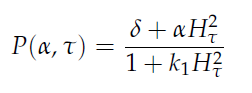Team:Wageningen UR/Project/ModelingProj1
From 2011.igem.org
(→Modeling synchronized oscillations) |
(→Modeling synchronized oscillations) |
||
| Line 35: | Line 35: | ||
[[File:mainproject01.png|center]] | [[File:mainproject01.png|center]] | ||
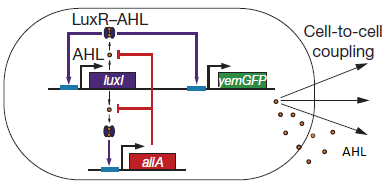
| - | '''Fig.1:''' ''Basic oscillating genetic circuit as published by Danino & Hasty.'' | + | '''Fig.1:''' ''Basic oscillating genetic circuit as published by Danino & Hasty.'' [http://www.nature.com/nature/journal/v463/n7279/abs/nature08753.html] |
The genes luxI and aiiA can be expressed when a AHL-LuxR complex binds to the promoter region. The circuit uses both a positive and negative feedback loop to control the AHL concentration and therefore the expression of the resulting proteins LuxI and AiiA. In the positive loop, LuxI generates more AHL, which, together with LuxR, can form more of the activating AHL-LuxR complex and therefore stimulates even more production of LuxI and consequently AHL. This complex also activates the expression of aiiA. In the negative feedback loop, AiiA degrades the AHL produced. For more detailed information about the circuit read more about the [[Team:Wageningen_UR/Project/CompleteProject1Description#2._Mechanism| mechanism]] in the [[Team:Wageningen_UR/Project/CompleteProject1Description| complete project description.]] | The genes luxI and aiiA can be expressed when a AHL-LuxR complex binds to the promoter region. The circuit uses both a positive and negative feedback loop to control the AHL concentration and therefore the expression of the resulting proteins LuxI and AiiA. In the positive loop, LuxI generates more AHL, which, together with LuxR, can form more of the activating AHL-LuxR complex and therefore stimulates even more production of LuxI and consequently AHL. This complex also activates the expression of aiiA. In the negative feedback loop, AiiA degrades the AHL produced. For more detailed information about the circuit read more about the [[Team:Wageningen_UR/Project/CompleteProject1Description#2._Mechanism| mechanism]] in the [[Team:Wageningen_UR/Project/CompleteProject1Description| complete project description.]] | ||
| Line 77: | Line 77: | ||
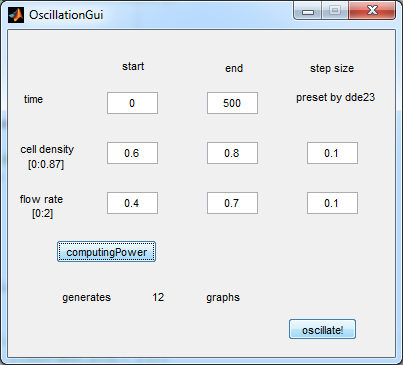
[[File:Oscillation_GUI_WUR.png|center]] | [[File:Oscillation_GUI_WUR.png|center]] | ||
| + | |||
| + | '''Fig.2:''' ''GUI of the matlab modeling tool created by team Wageningen UR'' | ||
| Line 83: | Line 85: | ||
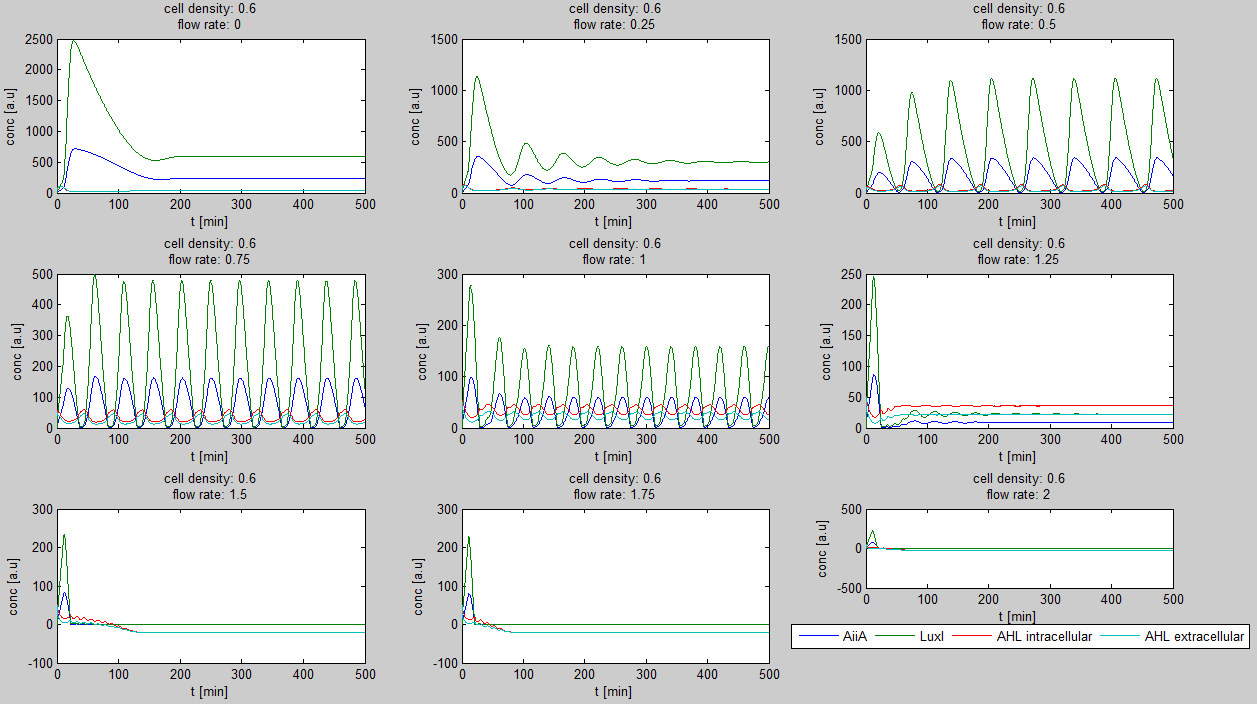
[[File:Example_output_graphs_WUR.png|700px|center]] | [[File:Example_output_graphs_WUR.png|700px|center]] | ||
| + | |||
| + | '''Fig.3:''' ''Example of a '' | ||
| + | |||
Revision as of 23:23, 16 September 2011
 "
"