|
|
| (22 intermediate revisions not shown) |
| Line 1: |
Line 1: |
| - | <!-- -------------------------------------------------From 2009 UQ--------------------------------------------------- -->
| + | {{:Team:UQ-Australia/Template:Header}} |
| | + | {|style="width:100%;" border="0" cellpadding="10" cellspacing="0" |
| | + | |- valign="top" |
| | + | |rowspan="2"|The human circadian rhythm drives many important processes in the body in accordance with the sleep/wake cycle. A characteristic of this biological clock is the periodic oscillation of gene expression. Current parts in the Registry designed to regulate periodic oscillations of gene expression have shown limited success. |
| | + | Here we demonstrate the feasibility of a biological clock being standardised as a set of BioBrick parts. |
| | | | |
| - | __NOTOC__
| + | Our network is controlled by an engineered promoter, Plac/ara, which features both an activator and a repressor domain. This controls the production of downstream genes to activate other inducible promoters, pBAD and GlnAp2, eventually leading to the production of a repressor protein, lacI, which inhibits Plac/ara, resulting in oscillatory expression. This project shows the feasibility of standardising the biological clock in E. coli and grounds further development for applications in regulated drug/hormone delivery and ion channel control. |
| - | <html>
| + | ! [[File:UQ-Australia_logo_2011.png|125x125px|link=https://2011.igem.org/Team:UQ-Australia]] |
| - | <body style="background-color:#772288">
| + | |- |
| | + | | |
| | + | |} |
| | | | |
| - | <style>
| |
| - | h1.firstHeading { display: none; }
| |
| | | | |
| - | p {text-align: justify;}
| + | == <span style="color:#558822">Project Details== |
| | | | |
| - | a:link { color: #603371; text-decoration: none}
| + | The project has been split into categories: |
| - | a:visited { color:#603371; text-decoration: none}
| + | * Development of BioBricks |
| - | a:hover { color:#f29400; text-decoration: none}
| + | ** Experimental methods to be fully recorded in the [[Team:UQ-Australia/Notebook|Notebook]] |
| - | a:active { color:#f29400; text-decoration: none} | + | * Modelling of the circuit |
| | + | ** Modelling of the kinetics of the oscillating cells |
| | + | ** Modelling of the synchronisation of oscillating cells |
| | + | * Thorough evaluation of the safety issues regarding UQ-Autralia's entry in iGEM |
| | + | * Human practices |
| | + | ** Raising awareness of synthetic biology |
| | + | ** Providing a solution to the patenting issue that iGEM is facing |
| | | | |
| - | #bodyContent { padding: 10px auto; width: 910px; margin: auto; clear: none; }
| + | Together, this forms the UQ-Australia project for the 2011 iGEM. |
| | | | |
| - | table#team_members { text-align: justify; border: 0; }
| |
| - | table#team_members h2, table#team_members h3 { clear: both; }
| |
| | | | |
| | + | === <span style="color:#D4A017">Motivation and Background</span> === |
| | | | |
| - | /*-----------------------------------------------------------------------------------------------*/
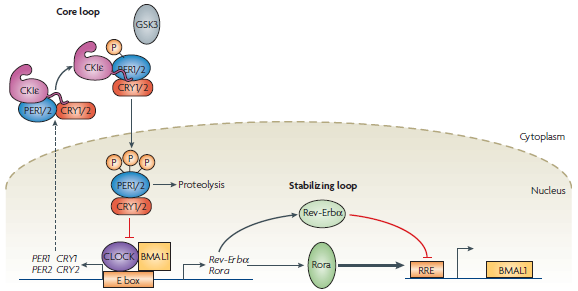
| + | In humans, the circadian rhythm is controlled by several core genes that operate via a series of feedback loops (Figure 1). A transcription–translation negative-feedback loop powers the system, with a delay between the transcription of these genes and the negative feedback being a key factor that allows the system to oscillate [1]. A 'master clock' located in the Suprachiasmatic Nucleus coordinates the timing of the rhythm, but external factors such as light exposure play a large role in regulating the exact |
| - | div.MenuBar ul li ul.DropDownMenu {
| + | |
| - | display: none; /* Hides all drop-down menus. */
| + | |
| | | | |
| - | }
| |
| - | /*
| |
| - | li:hover works in IE7 and FF2.
| |
| - | a:hover works in IE6 and FF2.
| |
| - | a:hover breaks li:hover in FF2.
| |
| - | */
| |
| - | div.MenuBar ul li:hover ul.DropDownMenu li ul.SideMenu,
| |
| - | div.MenuBar ul li a:hover ul.DropDownMenu li a ul.SideMenu {
| |
| - | display: none; /* Hides all side menus. */
| |
| - | }
| |
| - | /*------------------------------------------------------------------------------------- Menu Bar */
| |
| - | div.MenuBar {
| |
| - | font: arial, helvetica, sans-serif;
| |
| - | height: 50px;
| |
| - | width: 910px;
| |
| - | /*width: 100%*/
| |
| - | margin: 0;
| |
| - | border-top: 0;
| |
| - | border-right: 0;
| |
| - | border-left: 0;
| |
| - | padding: 0;
| |
| - | background: black;
| |
| - |
| |
| - | }
| |
| - | div.MenuBar ul {
| |
| - | font: arial, helvetica, sans-serif;
| |
| - | text-align: center;
| |
| - | list-style-type: none;
| |
| - | margin: 0;
| |
| - | border: 0;
| |
| - | padding: 0;
| |
| - | background: black;
| |
| - | }
| |
| - | div.MenuBar ul li {
| |
| - | font: arial, helvetica, sans-serif;
| |
| - | display: block;
| |
| - | padding: 0;
| |
| - | margin: 0;
| |
| - | font-size: 1.3em;
| |
| - | float: left;
| |
| - | background: black;
| |
| - | text-align: center;
| |
| - | width: 90px;
| |
| - | position: relative; /* Sets the positioning context for each drop-down menu. */
| |
| - | }
| |
| | | | |
| - | div.MenuBar ul li a {
| + | <b>Figure 1: The gene network responsible for establishing the circadian rhythm in humans [1]</b> |
| - | font: arial, helvetica, sans-serif;
| + | |
| - | display: block;
| + | |
| - | background: black;
| + | |
| - | height: 40px; /* Keep height + padding-top + padding-bottom sync with the menu bar height. */
| + | |
| - | color: #ffffff;
| + | |
| - | padding-top: 4px;
| + | |
| - | padding-bottom: 4px;
| + | |
| - | padding-left: 1em; /* Sets the left space between top-level items. */
| + | |
| - | padding-right: 1em; /* Sets the right space between top-level items. */
| + | |
| - | text-decoration: none;
| + | |
| - | }
| + | |
| | | | |
| - | /*------------------------------------------------------------------------------ Drop-Down Menus */
| + | [[File:Mammalian.png]] |
| - | div.MenuBar ul li:hover ul.DropDownMenu,
| + | |
| - | div.MenuBar ul li a:hover ul.DropDownMenu {
| + | |
| - | display: block;
| + | |
| - | width: 10em; /* Drop-down menu width.
| + | |
| - | Use MenuTailor.css to customize. */
| + | |
| - | height: 1em;
| + | |
| - | padding: 1px; /* Sets the drop-down menu "effective border" width. */
| + | |
| - | position: absolute;
| + | |
| - | top: 23px; /* Places the drop-down menu under the menu bar.
| + | |
| - | Keep it sync with the menu bar height. */
| + | |
| - | left: 0; /* Aligns the drop-down menu to its top-level item. */
| + | |
| - | background-color: black; /* Selected item. */
| + | |
| - | color: #FFFFFF;
| + | |
| | | | |
| - | }
| |
| - | div.MenuBar ul li:hover ul.DropDownMenu li a,
| |
| - | div.MenuBar ul li a:hover ul.DropDownMenu li a {
| |
| - | width: 10em; /* Keep it sync with the drop-down menu width.
| |
| - | Use MenuTailor.css to customize. */
| |
| - | height: 1em;
| |
| - | padding-left: 0;
| |
| - | padding-right: 0;
| |
| - | background-color: black; /* Selected item. */
| |
| - | color: #FFFFFF;
| |
| - | }
| |
| - | ul.DropDownMenu li a span {
| |
| - | display: block;
| |
| - | padding-left: 0.75em; /* Sets the left space of each drop-down menu item. */
| |
| - | padding-right: 0.25em; /* Sets the right space of each drop-down menu item. */
| |
| - | text-align: right; /* Aligns the >> symbol to the right. */
| |
| - | }
| |
| - | ul.DropDownMenu li a span span {
| |
| - | float: left; /* Aligns the text (back) to the left. */
| |
| - | font: 12px arial, helvetica, sans-serif; /* Required for IE55. */
| |
| - | height: 20px;
| |
| - | color: #FFFFFF;
| |
| - | }
| |
| - | /*----------------------------------------------------------------------------------- Side Menus */
| |
| - | div.MenuBar ul li:hover ul.DropDownMenu li:hover ul.SideMenu,
| |
| - | div.MenuBar ul li a:hover ul.DropDownMenu li a:hover ul.SideMenu {
| |
| - | display: block;
| |
| - | width: 11em; /* Side menu width.
| |
| - | Use MenuTailor.css to customize. */
| |
| - | padding: 1px; /* Sets the side menu "effective border" width. */
| |
| - | position: absolute;
| |
| - | top: -1px; /* Aligns the side menu to its drop-down menu item.
| |
| - | Keep it sync with the side menu "effective border" width. */
| |
| - | left: 13em; /* Places the side menu to the right of the drop-down menu.
| |
| - | Keep it sync with the drop-down menu width.
| |
| - | Use MenuTailor.css to customize. */
| |
| - | }
| |
| - | div.MenuBar ul li:hover ul.DropDownMenu li:hover ul.SideMenu li a,
| |
| - | div.MenuBar ul li a:hover ul.DropDownMenu li a:hover ul.SideMenu li a {
| |
| - | width: 11em; /* Keep it sync with the side menu width.
| |
| - | Use MenuTailor.css to customize. */
| |
| - | font: 12px arial, helvetica, sans-serif; /* Required for IE55. */
| |
| - | left: 13em; /* Places the side menu to the right of the drop-down menu.
| |
| - | Keep it sync with the drop-down menu width.
| |
| - | Use MenuTailor.css to customize. */
| |
| - | }
| |
| - | div.MenuBar ul li ul.DropDownMenu li ul.SideMenu li a span {
| |
| - | padding-left: 1.5em; /* Sets the left space of each side menu item. */
| |
| - | padding-right: 0.5em; /* Sets the right space of each side menu item. */
| |
| - | text-align: left;
| |
| - | font: 12px arial, helvetica, sans-serif; /* Required for IE55. */
| |
| - | left: 13em; /* Places the side menu to the right of the drop-down menu.
| |
| - | Keep it sync with the drop-down menu width.
| |
| - | Use MenuTailor.css to customize. */
| |
| - | }
| |
| - | /*----------------------------------------------------------------------------- Browser Specific */
| |
| - | * html div.MenuBar ul li a {
| |
| - | float: left; /* Required for IE55 and IE6.
| |
| - | Breaks O9.
| |
| - | Hidden (* html) from non-IE browsers. */
| |
| - | }
| |
| - | * html ul.DropDownMenu li a:hover {
| |
| - | cursor: hand; /* Required for IE55.
| |
| - | Hidden (* html) from non-IE browsers. */
| |
| - | }
| |
| - | ul.DropDownMenu li a:hover {
| |
| - | cursor: pointer; /* Required for IE6 and IE7.
| |
| - | Hidding it (* html) from non-IE browsers breaks IE7!
| |
| - | }
| |
| - | * html div.MenuBar a:hover {
| |
| - | text-decoration: none; /* Required for IE55 and IE6.
| |
| - | Hidden (* html) from non-IE browsers. */
| |
| - | }
| |
| - | * html div.MenuBar ul li table,
| |
| - | * html div.MenuBar ul li table td {
| |
| - | border: 0; /* Required for IE55 and IE6.
| |
| - | Hidden (* html) from non-IE browsers. */
| |
| - | }
| |
| - | /*------------------------------------------------------------------------------- Default Colors */
| |
| - | div.MenuBar {
| |
| - | background-color: Menu;
| |
| - | border-bottom: 1px solid ButtonShadow;
| |
| - | }
| |
| - | div.MenuBar a {
| |
| - | background-color: Menu; /* Top-level unselected items. */
| |
| - | color: MenuText;
| |
| - | }
| |
| - | div.MenuBar ul li:hover a,
| |
| - | div.MenuBar ul li a:hover {
| |
| - | color: #ea7f16;
| |
| - | background-color: Highlight; /* Top-level selected item. */
| |
| - | color: HighlightText;
| |
| - | }
| |
| - | /*...............................................................................................*/
| |
| - | div.MenuBar ul li:hover ul.DropDownMenu,
| |
| - | div.MenuBar ul li a:hover ul.DropDownMenu {
| |
| - | background-color: ButtonShadow; /* Sets the drop-down menu "effective border" color. */
| |
| - | }
| |
| - | div.MenuBar ul li:hover ul.DropDownMenu li a,
| |
| - | div.MenuBar ul li a:hover ul.DropDownMenu li a {
| |
| - | background-color: Menu; Drop-down menu unselected items.
| |
| - | Sets the drop-down menu "effective background" color. */
| |
| - | color: MenuText;
| |
| - | }
| |
| - | div.MenuBar ul li:hover ul.DropDownMenu li:hover a,
| |
| - | div.MenuBar ul li a:hover ul.DropDownMenu li a:hover {
| |
| - | background-color: Highlight; /* Drop-down menu selected item. */
| |
| - | color: HighlightText;
| |
| - | }
| |
| - | /*...............................................................................................*/
| |
| - | div.MenuBar ul li:hover ul.DropDownMenu li:hover ul.SideMenu,
| |
| - | div.MenuBar ul li a:hover ul.DropDownMenu li a:hover ul.SideMenu {
| |
| - | background-color: ButtonShadow; /* Sets the side menu "effective border" color. */
| |
| - | }
| |
| - | div.MenuBar ul li:hover ul.DropDownMenu li:hover ul.SideMenu li a,
| |
| - | div.MenuBar ul li a:hover ul.DropDownMenu li a:hover ul.SideMenu li a {
| |
| - | background-color: Menu; /* Side menu unselected items.
| |
| - | Sets the side menu "effective background" color. */
| |
| - | color: MenuText;
| |
| - | }
| |
| - | div.MenuBar ul li:hover ul.DropDownMenu li:hover ul.SideMenu li a:hover,
| |
| - | div.MenuBar ul li a:hover ul.DropDownMenu li a:hover ul.SideMenu li a:hover {
| |
| - | background-color: Highlight; /* Side menu selected item. */
| |
| - | color: HighlightText;
| |
| - | }
| |
| - | /*-----------------------------------------------------------------------------------------------*/
| |
| - | /*Script-Free 3-Level Menu 1.2 Tailor
| |
| - | www.CesarDaniel.info
| |
| - | /*-------------------------------------------------------------------------------------- General */
| |
| - | body {
| |
| - | background: white;
| |
| - | color: black;
| |
| - | margin: 0;
| |
| - | border: 0;
| |
| - | padding: 0;
| |
| - | }
| |
| | | | |
| | + | There has been much effort put into reconstructing and determining the exact nature of this system, as the impact the circadian rhythm has on our lifestyle and cellular processes are still not very well understood. In particular, it is believed the circadian rhythm could exert an effect on everything from body temperature, feeding behaviour and appetite, hormone secretion and metabolism, glucose homeostasis, and cell-cycle progression [2]. |
| | | | |
| - | div.MenuBar {
| + | Consequently, there have been a number of efforts to reconstruct this clock in a mammalian system for further study. In particular, both Hong <i>et al</i>. [3] and Tigges <i>et al</i>. [4] utilize the inducible tTa system to drive the expression and oscillation of genes in their synthetic networks. |
| - | font: 13px arial, helvetica, sans-serif;
| + | |
| - | }
| + | |
| - | div.MenuBar ul {
| + | |
| - | font: 13px arial, helvetica, sans-serif; /* Required for IE55. */
| + | |
| - | }
| + | |
| - | /*--------------------------------------------------------------------------------------- Colors */
| + | |
| - | div.MenuBar {
| + | |
| - | background-color: purple; /* Selected item. */
| + | |
| - | color: #FFFFFF;
| + | |
| - | border-bottom: 1px solid ButtonShadow;
| + | |
| - | }
| + | |
| - | div.MenuBar a,
| + | |
| - | div.MenuBar ul li:hover ul.DropDownMenu li a,
| + | |
| - | div.MenuBar ul li a:hover ul.DropDownMenu li a,
| + | |
| - | div.MenuBar ul li:hover ul.DropDownMenu li:hover ul.SideMenu li a,
| + | |
| - | div.MenuBar ul li a:hover ul.DropDownMenu li a:hover ul.SideMenu li a {
| + | |
| - | background-color: purple; /* Selected item. */
| + | |
| - | color: #FFFFFF;
| + | |
| - | }
| + | |
| - | div.MenuBar ul li:hover a,
| + | |
| - | div.MenuBar ul li a:hover,
| + | |
| - | div.MenuBar ul li:hover ul.DropDownMenu li:hover a,
| + | |
| - | div.MenuBar ul li a:hover ul.DropDownMenu li a:hover,
| + | |
| - | div.MenuBar ul li:hover ul.DropDownMenu li:hover ul.SideMenu li a:hover,
| + | |
| - | div.MenuBar ul li a:hover ul.DropDownMenu li a:hover ul.SideMenu li a:hover {
| + | |
| - | background-color: #603371; /* Selected item. */
| + | |
| - | color: #FFFFFF;
| + | |
| - | }
| + | |
| - | div.MenuBar ul li:hover ul.DropDownMenu,
| + | |
| - | div.MenuBar ul li a:hover ul.DropDownMenu,
| + | |
| - | div.MenuBar ul li:hover ul.DropDownMenu li:hover ul.SideMenu,
| + | |
| - | div.MenuBar ul li a:hover ul.DropDownMenu li a:hover ul.SideMenu {
| + | |
| - | background-color: ButtonShadow; /* Sets the drop-down and side menus "effective border" color. */
| + | |
| - | }
| + | |
| - | /*--------------------------------------------------------------------------------------- Widths */
| + | |
| - | /*
| + | |
| | | | |
| - | /*
| + | It was our initial plan to construct a similar oscillatory system ourselves using standardized parts which could be added to the registry and then utilized by other iGEM teams working in mammalian cells. Such as system could be 'plugged in' to any number of different outputs and would allow for the timely and regular expression of the genes it drives. However, we encountered a number of issues around the intellectual property protecting of certain elements we wanted to use and so decided to switch to a simpler system in <i>E. coli</i>. |
| - | Menu Bar 1
| + | |
| - | Drop-Down Menu #2
| + | |
| - | */
| + | |
| - | div.MenuBar#navi ul li:hover ul.DropDownMenu#MB1-DDM4,
| + | |
| - | div.MenuBar#navi ul li a:hover ul.DropDownMenu#MB1-DDM4,
| + | |
| - | div.MenuBar#navi ul li:hover ul.DropDownMenu#MB1-DDM4 li a,
| + | |
| - | div.MenuBar#navi ul li a:hover ul.DropDownMenu#MB1-DDM4 li a {
| + | |
| - | width: 11em; /* Drop-down menu width. */
| + | |
| - | }
| + | |
| - | div.MenuBar#navi ul li:hover ul.DropDownMenu#MB1-DDM5,
| + | |
| - | div.MenuBar#navi ul li a:hover ul.DropDownMenu#MB1-DDM5,
| + | |
| - | div.MenuBar#navi ul li:hover ul.DropDownMenu#MB1-DDM5 li a,
| + | |
| - | div.MenuBar#navi ul li a:hover ul.DropDownMenu#MB1-DDM5 li a {
| + | |
| - | width: 12em; /* Drop-down menu width. */
| + | |
| - | }
| + | |
| | | | |
| - | /*...............................................................................................*/
| + | === <span style="color:#D4A017">Our System</span> === |
| - | /*
| + | |
| - | Menu Bar 1
| + | |
| - | Drop-Down Menu #2
| + | |
| - | Side Menu #1
| + | |
| - | */
| + | |
| - | div.MenuBar#navi ul li:hover ul.DropDownMenu li:hover ul.SideMenu#MB1-DDM2-SM1,
| + | |
| - | div.MenuBar#navi ul li a:hover ul.DropDownMenu li a:hover ul.SideMenu#MB1-DDM2-SM1 {
| + | |
| - | left: 15.5em !important; /* Places the side menu to the right of the drop-down menu.
| + | |
| - | Keep it sync with the drop-down menu width. */
| + | |
| - | }
| + | |
| - | div.MenuBar#navi ul li:hover ul.DropDownMenu li:hover ul.SideMenu#MB1-DDM2-SM1,
| + | |
| - | div.MenuBar#navi ul li a:hover ul.DropDownMenu li a:hover ul.SideMenu#MB1-DDM2-SM1,
| + | |
| - | div.MenuBar#navi ul li:hover ul.DropDownMenu li:hover ul.SideMenu#MB1-DDM2-SM1 li a,
| + | |
| - | div.MenuBar#navi ul li a:hover ul.DropDownMenu li a:hover ul.SideMenu#MB1-DDM2-SM1 li a {
| + | |
| - | width: 10em; /* Side menu width. */
| + | |
| - | }
| + | |
| - | /*...............................................................................................*/
| + | |
| - | /*
| + | |
| - | Menu Bar 1
| + | |
| - | Drop-Down Menu #2
| + | |
| - | Side Menu #2
| + | |
| - | */
| + | |
| - | div.MenuBar#navi ul li:hover ul.DropDownMenu li:hover ul.SideMenu#MB1-DDM2-SM2,
| + | |
| - | div.MenuBar#navi ul li a:hover ul.DropDownMenu li a:hover ul.SideMenu#MB1-DDM2-SM2 {
| + | |
| - | left: 15.5em !important; /* Places the side menu to the right of the drop-down menu.
| + | |
| - | Keep it sync with the drop-down menu width. */
| + | |
| - | }
| + | |
| - | div.MenuBar#navi ul li:hover ul.DropDownMenu li:hover ul.SideMenu#MB1-DDM2-SM2,
| + | |
| - | div.MenuBar#navi ul li a:hover ul.DropDownMenu li a:hover ul.SideMenu#MB1-DDM2-SM2,
| + | |
| - | div.MenuBar#navi ul li:hover ul.DropDownMenu li:hover ul.SideMenu#MB1-DDM2-SM2 li a,
| + | |
| - | div.MenuBar#navi ul li a:hover ul.DropDownMenu li a:hover ul.SideMenu#MB1-DDM2-SM2 li a {
| + | |
| - | width: 10em; /* Side menu width. */
| + | |
| - | }
| + | |
| - | /*...............................................................................................*/
| + | |
| - | /*
| + | |
| - | Menu Bar 1
| + | |
| - | Drop-Down Menu #2
| + | |
| - | Side Menu #3
| + | |
| - | */
| + | |
| - | div.MenuBar#navi ul li:hover ul.DropDownMenu li:hover ul.SideMenu#MB1-DDM2-SM3,
| + | |
| - | div.MenuBar#navi ul li a:hover ul.DropDownMenu li a:hover ul.SideMenu#MB1-DDM2-SM3 {
| + | |
| - | left: 15.5em !important; /* Places the side menu to the right of the drop-down menu.
| + | |
| - | Keep it sync with the drop-down menu width. */
| + | |
| - | }
| + | |
| - | div.MenuBar#navi ul li:hover ul.DropDownMenu li:hover ul.SideMenu#MB1-DDM2-SM3,
| + | |
| - | div.MenuBar#navi ul li a:hover ul.DropDownMenu li a:hover ul.SideMenu#MB1-DDM2-SM3,
| + | |
| - | div.MenuBar#navi ul li:hover ul.DropDownMenu li:hover ul.SideMenu#MB1-DDM2-SM3 li a,
| + | |
| - | div.MenuBar#navi ul li a:hover ul.DropDownMenu li a:hover ul.SideMenu#MB1-DDM2-SM3 li a {
| + | |
| - | width: 10em; /* Side menu width. */
| + | |
| - | }
| + | |
| - | /*...............................................................................................*/
| + | |
| | | | |
| - | </style> | + | In prokaryotic systems, there are a number of tools available for synthetic biologists to construct gene networks such as the circadian clock we hope to replicate. In particular, we are going to make use of the inducible promoters glnAp2 and pBAD and their inducers glnG and araC. These components have been used in synthetic oscillators from both Atkinson <i>et al</i>. [5] and Stricker <i>et al</i>. [6]. In addition, we plan on using an engineered promoter denoted as Plac/ara, which features a pBAD activation domain as well as several lacO sites, meaning it is both inducible and repressible depending on the input. This promoter was first engineered by Lutz & Bujard [7]. |
| | | | |
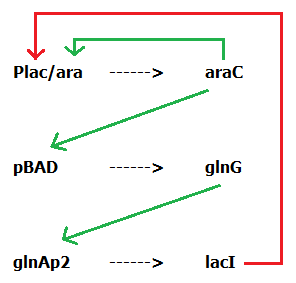
| | + | Based on the work from these previous studies, we designed our own system using components that have been previously utilized to construct other synthetic clocks (Figure 2). We decided to make a 3 component system rather that a dual component system as the issues encountered by some other synthetic clocks suggest that there might not have been enough time between the transcription of various components [8]. As such, including this extra step to add a delay should ensure our network is able to reach an oscillatory expression pattern. |
| | | | |
| - | <body> | + | <b>Figure 2: The genetic system we are aiming to construct in order to produce an oscillating gene network. An engineered promoter, Plac/ara, which features both an activator and a repressor domain controls the production of downstream genes to activate other inducible promoters, pBAD and GlnAp2. These eventually lead to the production of a repressor protein, lacI, which inhibits Plac/ara, resulting in oscillatory expression.</b> |
| - | <div id="header"><img src=" https://static.igem.org/mediawiki/2009/a/a6/UQ_banner.jpg" /></div>
| + | |
| - | <div class='MenuBar' id="navi">
| + | |
| - | <ul>
| + | |
| - | <li>
| + | |
| - | <a href="https://2011.igem.org/Team:UQ-Australia" style="color: white">Home
| + | |
| - | <!--[if gt IE 6]><!--></a><!--<![endif]-->
| + | |
| - | <!--[if lt IE 7]><table border="0" cellpadding="0" cellspacing="0"><tr><td><![endif]-->
| + | |
| - | <!--[if lte IE 6]></td></tr></table></a><![endif]-->
| + | |
| - | </li>
| + | |
| - | <li>
| + | |
| - | <a href="https://2011.igem.org/Team:UQ-Australia/Team" style="color: white">Team
| + | |
| - | <!--[if gt IE 6]><!--></a><!--<![endif]-->
| + | |
| - | <!--[if lt IE 7]><table border="0" cellpadding="0" cellspacing="0"><tr><td><![endif]-->
| + | |
| - | <!--[if lte IE 6]></td></tr></table></a><![endif]-->
| + | |
| - | </li>
| + | |
| - | <li>
| + | |
| - | <a href="https://2011.igem.org/Team:UQ-Australia/Project" style="color: white">Project
| + | |
| - | <!--[if gt IE 6]><!--></a><!--<![endif]-->
| + | |
| - | <!--[if lt IE 7]><table border="0" cellpadding="0" cellspacing="0"><tr><td><![endif]-->
| + | |
| - | <!--[if lte IE 6]></td></tr></table></a><![endif]-->
| + | |
| | | | |
| - | </li>
| + | [[File:Oursystem.png]] |
| - | <li>
| + | |
| | | | |
| - | <a href="https://2011.igem.org/Team:UQ-Australia/Parts" style="color: white">Parts
| |
| - | <!--[if gt IE 6]><!--></a><!--<![endif]-->
| |
| - | <!--[if lt IE 7]><table border="0" cellpadding="0" cellspacing="0"><tr><td><![endif]-->
| |
| - | <!--[if lte IE 6]></td></tr></table></a><![endif]-->
| |
| - | </li>
| |
| - | <li>
| |
| | | | |
| - | <a href="https://2011.igem.org/Team:UQ-Australia/Data" style="color: white">Data
| + | Once our system has been constructed, we hope to be able to make it even more stable by integrating the necessary components into the <i>E coli</i> chromosome. In this manner, we will be able to remove any interfering factors from our system, and will also be able to control the gene dosage level of our system. This has been a particular problem for other synthetic oscillators, with researchers being unable to control for the uptake of plasmid and therefore having cell-to-cell differences in the rate of oscillation [4, 6]. |
| - | <!--[if gt IE 6]><!--></a><!--<![endif]-->
| + | |
| - | <!--[if lt IE 7]><table border="0" cellpadding="0" cellspacing="0"><tr><td><![endif]-->
| + | |
| - | <!--[if lte IE 6]></td></tr></table></a><![endif]-->
| + | |
| - | </li>
| + | |
| - | <li>
| + | |
| - | <a href="https://2011.igem.org/Team:UQ-Australia/Modeling" style="color:white">Modelling
| + | |
| - | <!--[if gt IE 6]><!--></a><!--<![endif]-->
| + | |
| - | <!--[if lt IE 7]><table border="0" cellpadding="0" cellspacing="0"><tr><td><![endif]-->
| + | |
| - | <!--[if lte IE 6]></td></tr></table></a><![endif]-->
| + | |
| - | </li>
| + | |
| - | <li>
| + | |
| - | <a href="https://2011.igem.org/Team:UQ-Australia/Notebook" style="color: white">Notebook
| + | |
| - | <!--[if gt IE 6]><!--></a><!--<![endif]-->
| + | |
| - | <!--[if lte IE 6]></td></tr></table></a><![endif]-->
| + | |
| - |
| + | |
| | | | |
| - | </li>
| + | Kuhlman & Cox [9] have developed a method for integrating large synthetic constructs into the bacterial chromosome, and kindly provided us with the necessary plasmids should we reach this stage. Furthermore, Dr. Alex Ninfa [5] provided us with <i>E. coli</i> strain 3.300L*G, which has both glnG and lacI knocked out so as not to interfere with the network we construct. |
| - | <li>
| + | |
| - | <a href="https://2011.igem.org/Team:UQ-Australia/Safety" style="color: white">Safety
| + | |
| - | <!--[if gt IE 6]><!--></a><!--<![endif]-->
| + | |
| - | <!--[if lt IE 7]><table border="0" cellpadding="0" cellspacing="0"><tr><td><![endif]-->
| + | |
| - | <!--[if lte IE 6]></td></tr></table></a><![endif]-->
| + | |
| - | </li>
| + | |
| - | <li>
| + | |
| - | <a href="https://2011.igem.org/Team:UQ-Australia/Human_Practices" style="color: white">Human Practices
| + | |
| - | <!--[if gt IE 6]><!--></a><!--<![endif]-->
| + | |
| - | <!--[if lt IE 7]><table border="0" cellpadding="0" cellspacing="0"><tr><td><![endif]-->
| + | |
| - | <!--[if lte IE 6]></td></tr></table></a><![endif]-->
| + | |
| - | </li>
| + | |
| | | | |
| - | <li>
| |
| - | <a href="https://2011.igem.org/Team:UQ-Australia/Attributions" style="color: white">Attributions</a>
| |
| - | </li>
| |
| - | </ul>
| |
| | | | |
| - | </div>
| |
| - | <div id="header_bottom"><img src="https://static.igem.org/mediawiki/2008/e/ef/Navi_bg.gif" alt="" / ></div>
| |
| - | </body>
| |
| - | </html>
| |
| | | | |
| | + | Details on the modelling behind out circuit can be found on the [[Team:UQ-Australia/Modeling|Modelling]] page. |
| | | | |
| - | <!-- ----------------------------------------------------text-------------------------------------------------------- --> | + | === <span style="color:#D4A017">Experimental Outline & Goals</span> === |
| | | | |
| | + | Our main aim for the experimental work is to produce each component as a standardized and characterised BioBrick part that can be re-used by other teams in future to build their own oscillators. In the long run, we would hope to produce an <i>E. Coli</i> strain incorporating the three main 'modules' of our oscillator. In this manner, other teams could use this strain and simply transform the bacteria with a plasmid driven by one of our inducible promoters in order to obtain regular, oscillating expression. |
| | | | |
| | + | To achieve these goals, we must first source all the components we require and standardize them to include BioBrick sites. Standard BioBrick assembly can then be used to construct our main components, before we will utilize a pre-determined system and protocols [9] to attempt to integrate our modules into the chromosome. In the process of developing our parts and constructing the system, we hope to characterise all our parts. In particular, confirming the inducibility and kinetics of our promoters will allow us to refine our models. |
| | | | |
| | + | === <span style="color:#D4A017">Other activities</span> === |
| | | | |
| | + | Our safety analysis is on the [[Team:UQ-Australia/Safety|Safety]] page, while the Human Practices discussion is in the [[Team:UQ-Australia/Human_Practices|Human Practices]] section. |
| | | | |
| | | | |
| | + | === <span style="color:#D4A017">References</span>=== |
| | | | |
| - | {|align="justify"
| + | [1] Gallego, M & Virshup, DM 2007. "Post-translational modifications regulate the ticking of the circadian clock", <i>Molecular Cell Biology</i>, vol. 8, pp. 139-148. |
| - | |Inspired by the circadian clock in humans which regulates a number of very important processes, we are trying to replicate this biological clock in a bacterial system. We are aiming to construct a network of genes that oscillates in a similar fashion to the 24 hour system in humans. If we are successful, we will be able to put different genes into our system so that we can make the bacteria perform a particular process periodically – a simple example of this would be to make them flash on and off consistently.
| + | |
| - | |[[Image:UQ-Australia_logo.png|200px|right|frame]]
| + | |
| - | |-
| + | |
| - | |
| + | |
| - | To achieve this oscillatory behaviour we will utilise a gene network with a series of inducible promoters that generate the production of other activating proteins, all driven by a constitutively active promoter. This promoter features an engineered repression domain (the inhibitor of this promoter being the output of the final step in the network). If everything goes as planned, these linked activations and repression will produce fluctuating levels of the proteins in question, which could then be used to drive our output function (initially just GFP production and a timed fluorescence). Ultimately, we hope our system could be used to drive the timed release of drugs or other biological factors.
| + | |
| - | |[[Image:UQ-Australia_team.png|right|frame|Your team picture]]
| + | |
| - | |-
| + | |
| - | |
| + | |
| - | |align="center"|[[Team:UQ-Australia | Team Example]]
| + | |
| - | |}
| + | |
| - | | + | |
| - | <!--- The Mission, Experiments --->
| + | |
| - | | + | |
| - | | + | |
| - | == <span style="color:#558822">Project Details==
| + | |
| - | | + | |
| - | The project has been split into categories:
| + | |
| - | * Development of BioBricks
| + | |
| - | ** Experimental methods to be fully recorded in the [[Team:UQ-Australia/Notebook|Notebook]]
| + | |
| - | * Modelling of the circuit
| + | |
| - | ** Modelling of the kinetics of the oscillating cells
| + | |
| - | ** Modelling of the synchronisation of oscillating cells
| + | |
| - | * Thorough evaluation of the safety issues regarding UQ-Autralia's entry in iGEM
| + | |
| - | * Human practices
| + | |
| - | ** Raising awareness of synthetic biology
| + | |
| - | ** Providing a solution to the patenting issue that iGEM is facing
| + | |
| - | | + | |
| - | Together, this forms the UQ-Australia project for the 2011 iGEM.
| + | |
| - | | + | |
| - | | + | |
| - | | + | |
| - | | + | |
| - | === <span style="color:#883622">Experimental Work</span> ===
| + | |
| - | | + | |
| - | Outcomes of experimental work are to be recorded here.
| + | |
| | | | |
| - | === <span style="color:#883622">Modelling</span> ===
| + | [2] Takahashi, JS, Hong, HK, Ko, CH & McDearmon, EL 2008. "The genetics of mammalian circadian order and disorder: implications for physiology and disease", <i>Genetics</i>, vol. 9, pp. 764-775. |
| | | | |
| | + | [3] Hong, HK, Chong, JL, Song, W, Song, EJ & Jyawook, AA <i>et al</i>. 2007. "Inducible and reversible <i>Clock</i> gene expression in brain using the tTa system for the study of circadian behaviour", <i>PLoS Genetics</i>, vol. 3, no. 2, 324-338. |
| | | | |
| - | Details of this is on the [[Team:UQ-Australia/Modeling|Modelling]] page.
| + | [4] Tigges, M, Marques-Lago, TT, Stelling, J & Fusseneger, M 2009. "A tunable synthetic mammalian oscillator", <i>Nature</i>, vol. 457, pp. 309-312. |
| | | | |
| - | === <span style="color:#883622">Safety</span> ===
| + | [5] Atkinson, MR, Savageau, MA, Myers, JT & Ninfa, AJ 2003. "Development of genetic circuitry exhibiting toggle switch or oscillatory behaviour in Escherichia coli", <i>Cell</i>, vol. 113, pp. 597-607. |
| | | | |
| - | Details of this is on the [[Team:UQ-Australia/Safety|Safety]] page.
| + | [6] Stricker, J, Cookson, S, Bennett, MR, Mather, WH, Tsimring, LS & Hasty, J 2008. "A fast, robust and tunable synthetic oscillator", <i>Nature</i>, vol. 456, pp. 516-520. |
| | | | |
| | + | [7] Lutz, R & Bujard H 1997. "Independent and tight regulation of transcriptional units in Escherichia coli via the LacR/O, the TetR/O and araC/I1-I2 regulatory elements", <i>Nucleic Acids Research</i>, vol. 25, no. 6, pp. 1203-1210. |
| | | | |
| - | === <span style="color:#883622">Human Practices </span>===
| + | [8] Purcell, O, Savery, NJ, Grierson, CS & di Bernardo, M 2010. "A comparative analysis of synthetic genetic oscillators", <i>Journal of the Royal Society</i>, vol. 7, pp. 1503-1524. |
| | | | |
| - | The Human Practices section is on the [[Team:UQ-Australia/Human_Practices|Human Practices]]
| + | [9] Kuhlman, TE & Cox, EC 2010. "Site-specific chromosomal integration of large synthetic constructs", <i>Nucleic Acids Research</i>, vol. 38, no. 6, pp. e92. |
 "
"