Team:UNICAMP-EMSE Brazil/Results
From 2011.igem.org
Neshich.iap (Talk | contribs) (→Device 3 testing, Protein Secretion System) |
Neshich.iap (Talk | contribs) (→Discussion) |
||
| (38 intermediate revisions not shown) | |||
| Line 25: | Line 25: | ||
In order to test the ability of Jedi Bacteria in sensing NO levels and activating genes linked to SoxS promoter, we built a Device testing, with GFP linked to SoxS promoter, as it is shown in the following schema (Figure 1): | In order to test the ability of Jedi Bacteria in sensing NO levels and activating genes linked to SoxS promoter, we built a Device testing, with GFP linked to SoxS promoter, as it is shown in the following schema (Figure 1): | ||
| - | [[Image:UNICAMP_EMSE_GFP_SOX_device.jpg|center| | + | [[Image:UNICAMP_EMSE_GFP_SOX_device.jpg|center|660px]] |
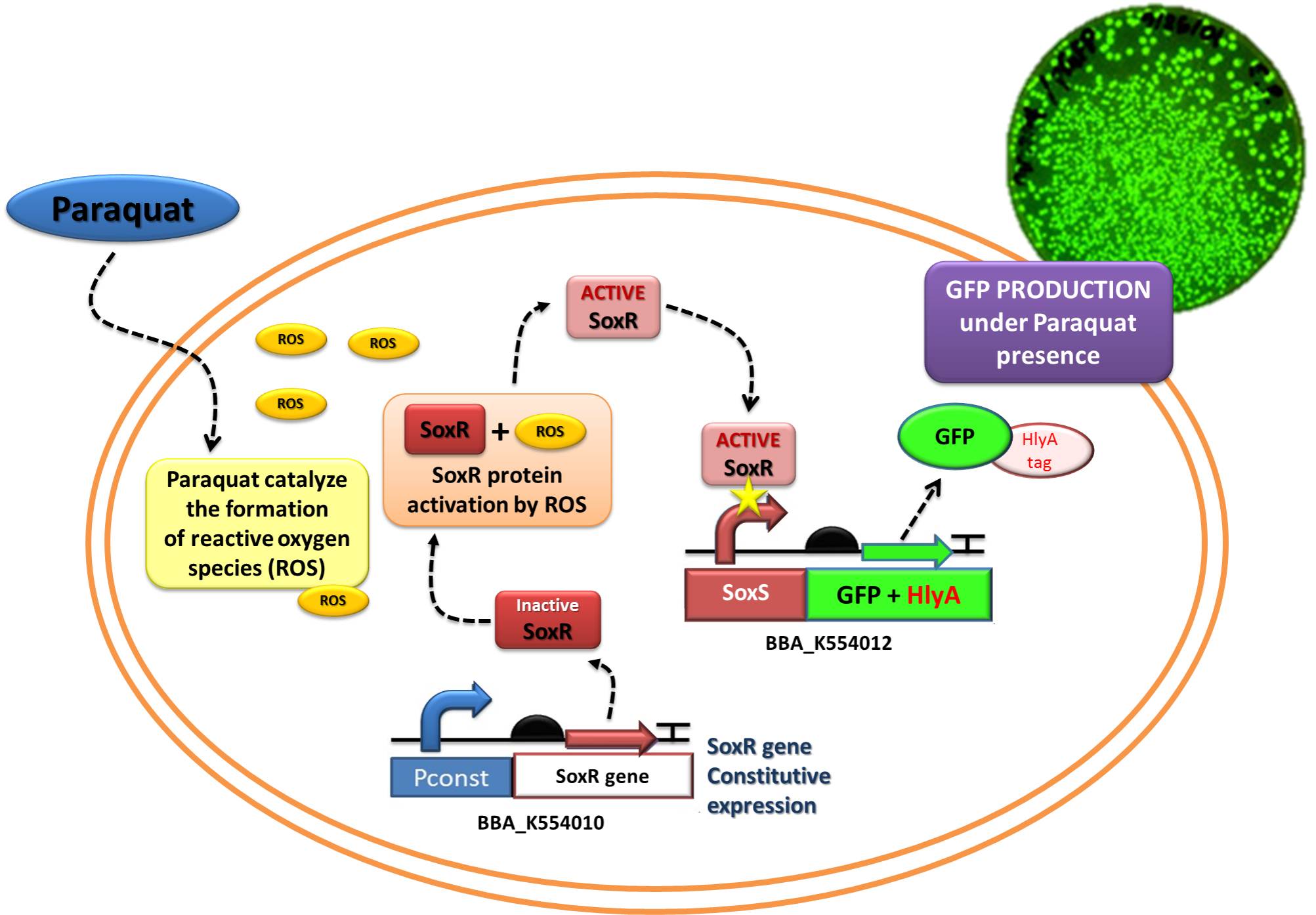
<div align=center> '''Figure 1: Testing [https://2011.igem.org/Team:UNICAMP-EMSE_Brazil/Project#Device_2:_NO_sensor.2FIL-10_producer Device 2] through replacement of IL-10 to GFP. '''</div> | <div align=center> '''Figure 1: Testing [https://2011.igem.org/Team:UNICAMP-EMSE_Brazil/Project#Device_2:_NO_sensor.2FIL-10_producer Device 2] through replacement of IL-10 to GFP. '''</div> | ||
| Line 31: | Line 31: | ||
===Methods=== | ===Methods=== | ||
| - | Competent E. coli DH5α strain cells were transformed with a pSB1A2 vector (Ampicillin resistant) carrying both the sensor (Strong_Constitutive_promoter + RBS + SoxR + Terminator) and the effector (SoxS_pormoter + RBS + GFP + HlyA + Terminator) devices using a chemical shock protocol. Transformed bacteria were plated on solid LB medium with 50 μg/mL Ampicillin and grown at 37ºC overnight. Surviving colonies were grown on liquid LB medium containing 50 μg/mL Ampicillin. Oxidative stress was induced by adding increasing concentrations of | + | Competent E. coli DH5α strain cells were transformed with a pSB1A2 vector (Ampicillin resistant) carrying both the sensor (Strong_Constitutive_promoter + RBS + SoxR + Terminator) and the effector (SoxS_pormoter + RBS + GFP + HlyA + Terminator) devices using a chemical shock protocol. Transformed bacteria were plated on solid LB medium with 50 μg/mL Ampicillin and grown at 37ºC overnight. Surviving colonies were grown on liquid LB medium containing 50 μg/mL Ampicillin. Oxidative stress was induced by adding increasing concentrations of [http://en.wikipedia.org/wiki/Paraquat Paraquat] (Methyl viologen dichloride hydrate - Sigma), an oxidative stress inducer in bacteria. Final Paraquat concentrations were: 0 μM (control), 5 μM, 10 μM, 20 μM, 30 μM and 40 μM. Induction of the designed sensor/effector mechanism was accessed by fluorescence using a fluorometer (SLM – Aminco; 4 nm bandpass and 10 mm) with excitation in 500 nm and emission spectra from 508-550 nm. |
| + | |||
In order to access if increasing concentrations of Paraquat could inhibit culture growth due to it´s toxicity, optical density (OD) levels of the culture were measured during the incubation time with Paraquat. | In order to access if increasing concentrations of Paraquat could inhibit culture growth due to it´s toxicity, optical density (OD) levels of the culture were measured during the incubation time with Paraquat. | ||
The ability to recognize NO (nitric oxide), an inflammation signal molecule, was characterized for SoxR/SoxS sensor, and <font color=red>'''found to be FUNCTIONAL'''</font>. In the section below we present the detailed results. | The ability to recognize NO (nitric oxide), an inflammation signal molecule, was characterized for SoxR/SoxS sensor, and <font color=red>'''found to be FUNCTIONAL'''</font>. In the section below we present the detailed results. | ||
| Line 37: | Line 38: | ||
===Results=== | ===Results=== | ||
Achieved results indicated that the designed sensor/effector device system was capable of inducible production of GFP (or another generic protein controlled by the sensor). In addition, protein induction can be modulated through varying inducer concentration (Figure 2). Higher concentrations of Paraquat exhibited higher fluorescence levels, which indicates increased GFP concentrations. No plateau was achieved using the highest tested concentrations. | Achieved results indicated that the designed sensor/effector device system was capable of inducible production of GFP (or another generic protein controlled by the sensor). In addition, protein induction can be modulated through varying inducer concentration (Figure 2). Higher concentrations of Paraquat exhibited higher fluorescence levels, which indicates increased GFP concentrations. No plateau was achieved using the highest tested concentrations. | ||
| - | + | ||
| - | + | ||
| - | [[Image:UNICAMP_EMSE_GFP_SOX_device_result1.jpg|center| | + | [[Image:UNICAMP_EMSE_GFP_SOX_device_result1.jpg|center|500px]] |
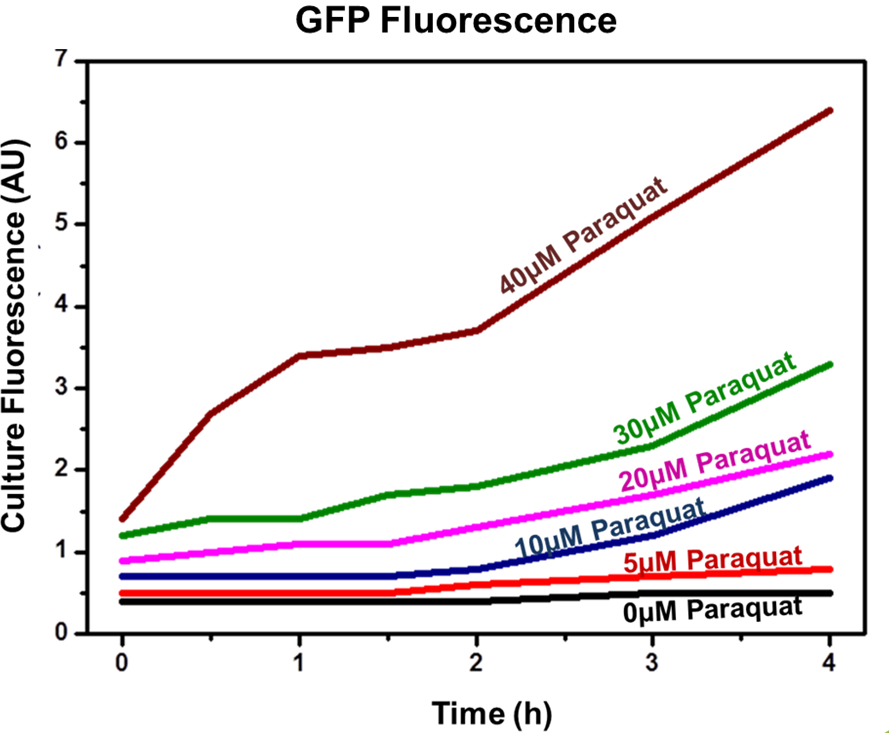
| - | <div align=center>'''Figure 2: | + | <div align=center>'''Figure 2: SoxS/SoxR fluorescence data for concentrations 0, 5, 10, 20, 30 and 40 µM of inducer (Paraquat).'''</div> |
<br> | <br> | ||
| - | |||
| - | |||
| - | |||
Moreover, we tested if Paraquat could be toxic for the bacteria cells, and we found that the experimental concentrations of Paraquat did not show significant differences in cell growth as shown by OD levels in Figure 3. | Moreover, we tested if Paraquat could be toxic for the bacteria cells, and we found that the experimental concentrations of Paraquat did not show significant differences in cell growth as shown by OD levels in Figure 3. | ||
| Line 52: | Line 49: | ||
[[Image:UNICAMP_EMSE_GFP_SOX_device_result2.jpg|center|600px]] | [[Image:UNICAMP_EMSE_GFP_SOX_device_result2.jpg|center|600px]] | ||
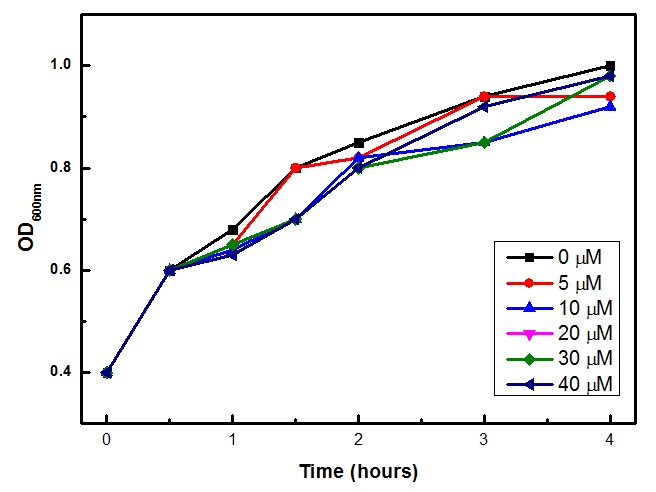
| - | <div align=center>'''Figure 3: SoxS/SoxR optical density data for concentrations 0, 5, 10, 20, 30 and 40 µM of inducer.'''</div> | + | <div align=center>'''Figure 3: SoxS/SoxR optical density data for concentrations 0, 5, 10, 20, 30 and 40 µM of inducer (Paraquat).'''</div> |
| Line 59: | Line 56: | ||
<br> | <br> | ||
| + | |||
| + | Fluorescence Microscopy experiments also corroborates GFP production through Paraquat induction of SoxR/SoxS sensor/effector system. Fluorescence was detected in the cultures containing the sensor and GFP production devices when induced with 40 uM of Paraquat (Figure 5), but not in non-induced cultures(Figure 6). This is an additional evidence that the NO sensor system and Sox driven effector (in this case, GFP) synthesis worked as expected. | ||
| + | |||
| + | |||
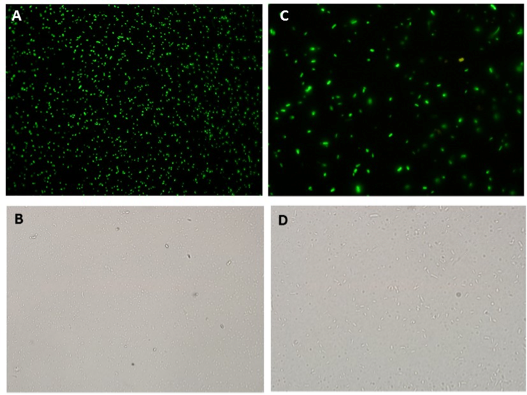
| + | [[Image:picind.png|center|600px]] | ||
| + | <div align=center>'''Figure 5: GFP fluorescence assessed by microscopy in 40 μM Paraquat induced culture. A) Fluorescence microscopy 40X Exp.: 0.478 ms; B) Light microscopy 40X; C) Fluorescence microscopy 100X Exp.: 0.478 ms; D) Light microscopy 100X.'''</div> | ||
| + | |||
| + | |||
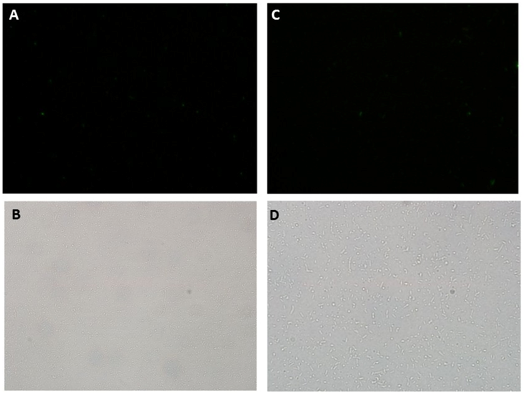
| + | [[Image:picnotind.png|center|600px]] | ||
| + | <div align=center>'''Figure 6: GFP fluorescence assessed by microscopy in non-induced culture. A) Fluorescence microscopy 40X Exp.: 0.478 ms; B) Light microscopy 40X; C) Fluorescence microscopy 100X Exp.: 0.478 ms; D) Light microscopy 100X.'''</div> | ||
===Discussion=== | ===Discussion=== | ||
The SoxR/SoxS system is one of the best characterized redox-sensing mechanisms in bacteria. Under oxidative stress conditions, activation of this system results in a cascade effect that ends in the activation of more than 16 genes that can counteract the harmful effect of superoxide and other oxidative radicals (Pomposiello and Demple 2001). | The SoxR/SoxS system is one of the best characterized redox-sensing mechanisms in bacteria. Under oxidative stress conditions, activation of this system results in a cascade effect that ends in the activation of more than 16 genes that can counteract the harmful effect of superoxide and other oxidative radicals (Pomposiello and Demple 2001). | ||
| - | This device is a modification of Stanford team anti-inflammatory device (iGEM 2009 - https://2009.igem.org/Team:Stanford/ProjectPage), which comprises SoxR gene (BBa_K223047) under control of a Constitutive Promoter (BBa_J23119) and SoxS promoter (BBa_K223001 – deleted part). Although this system has already been designed and used in previously iGEM competitions (iGEM 2009 - https://2009.igem.org/Team:Stanford/ProjectPage), we took advantage of the availability of synthesized sequences to design new parts (SoxR and SoxS) that conforms the common biobrick standards in the iGEM registry. Our team has improved the parts | + | This device is a modification of Stanford team anti-inflammatory device (iGEM 2009 - https://2009.igem.org/Team:Stanford/ProjectPage), which comprises SoxR gene (BBa_K223047) under control of a Constitutive Promoter (BBa_J23119) and SoxS promoter (BBa_K223001 – deleted part). Although this system has already been designed and used in previously iGEM competitions (iGEM 2009 - https://2009.igem.org/Team:Stanford/ProjectPage), we took advantage of the availability of synthesized sequences to design new parts (SoxR and SoxS) that conforms the common biobrick standards in the iGEM registry. Our team has improved the parts as described in [https://2011.igem.org/Team:UNICAMP-EMSE_Brazil/Project#UNICAMP-EMSE_Brazil_Parts_Design:_why_are_they_different_to_Stanford_2009_team_ones.3F Innovation]. |
These newly designed parts were capable of sensing different concentrations of the inducer, resulting in an increased production of the protein under control of the SoxS promoter (in this case, GFP). | These newly designed parts were capable of sensing different concentrations of the inducer, resulting in an increased production of the protein under control of the SoxS promoter (in this case, GFP). | ||
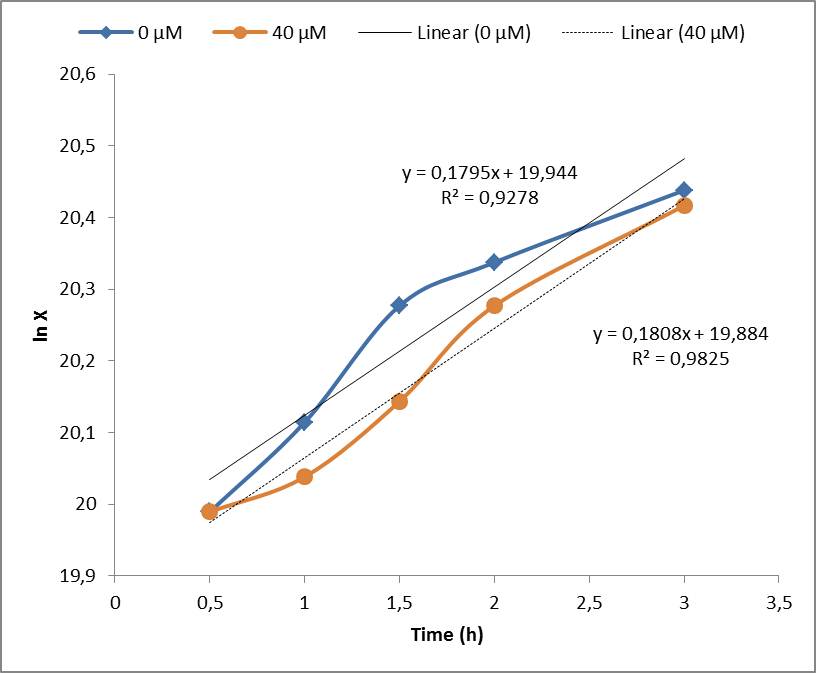
| - | Under the experimental conditions, no cell growth inhibition was observed and confirmed by the calculations of the specific growth rate (μ). No significant difference was found in control experiment (0 μM of Paraquat) and cell induced with 40 μM of Paraquat, both presented μ= 0,18 h-1. | + | Under the experimental conditions, no cell growth inhibition was observed and confirmed by the calculations of the specific growth rate (μ). No significant difference was found in control experiment (0 μM of Paraquat) and cell induced with 40 μM of Paraquat, both presented μ= 0,18 h-1. Stanford 2009 team showed that Paraquat concentrations between 60 μM and 80 μM can inhibit ''E. coli'' cell growth due to enhanced toxicity, but these concentrations were not included in our tests. |
With this procedure were able to validate the following parts: SoxS promoter ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K554000 BBa_K554000]), SoxR gene ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K554003 BBa_K554003]), SoxR device ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K554012 BBa_K554012]), SoxS GFP HlyA device [http://partsregistry.org/wiki/index.php?title=Part:BBa_K554012 BBa_K554012] | With this procedure were able to validate the following parts: SoxS promoter ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K554000 BBa_K554000]), SoxR gene ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K554003 BBa_K554003]), SoxR device ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K554012 BBa_K554012]), SoxS GFP HlyA device [http://partsregistry.org/wiki/index.php?title=Part:BBa_K554012 BBa_K554012] | ||
| Line 73: | Line 80: | ||
==Device 3 testing, Protein Secretion System== | ==Device 3 testing, Protein Secretion System== | ||
| - | The assembled devices related to secretion system (Hemolysin secretion system under control of SoxS – SoxS-HlyB-HlyD-TolC) were tested under laboratory conditions using GFP as reporter | + | The assembled devices related to secretion system (Hemolysin secretion system under control of SoxS – SoxS-HlyB-HlyD-TolC) were tested under laboratory conditions using GFP as reporter. |
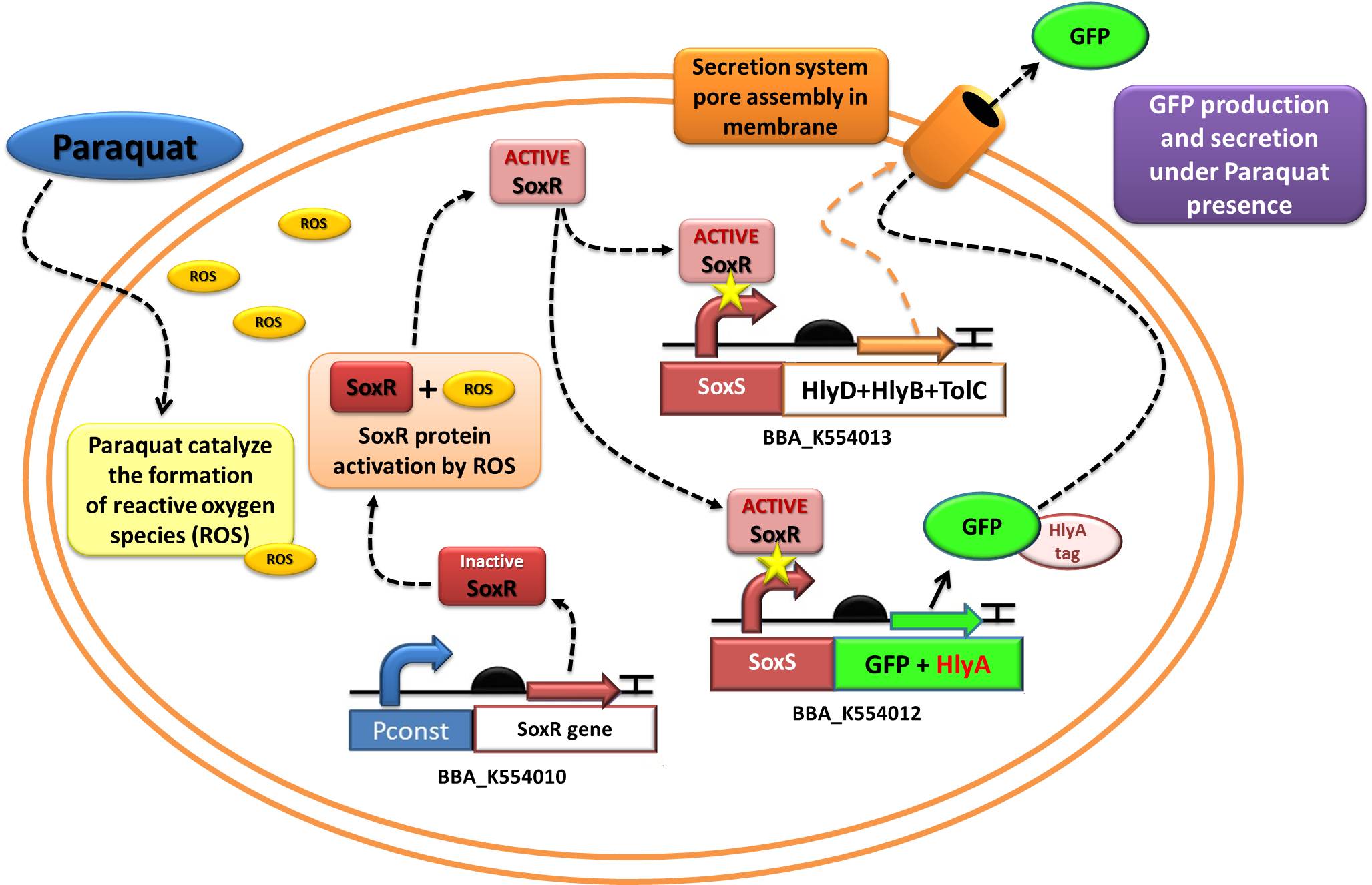
[[Image:UNICAMP_EMSE_secretion_device_schema3.jpg|center|730px]] | [[Image:UNICAMP_EMSE_secretion_device_schema3.jpg|center|730px]] | ||
| Line 89: | Line 96: | ||
*Sensor/Effector - Secretion system: 40µM Paraquat (non-secretion control) | *Sensor/Effector - Secretion system: 40µM Paraquat (non-secretion control) | ||
| - | After 3 hours of induction, samples were collected, centrifuged (4000 rpm / 10 min; to avoid cell lysis) and the supernatant was collected and centrifuged again (13000 rpm / 10 min; to remove remaining cells). The supernatant fluorescence was measured in fluorometer (SLM – Aminco; 4 nm bandpass and 10 mm) with excitation in 500 nm and emission spectra from 508-550 nm. | + | After 3 hours of induction, samples were collected, centrifuged (4000 rpm / 10 min; to avoid cell lysis) and the supernatant was collected and centrifuged again (13000 rpm / 10 min; to remove remaining cells). The supernatant fluorescence was measured in fluorometer (SLM – Aminco; 4 nm bandpass and 10 mm) with excitation in 500 nm and emission spectra from 508-550 nm. |
| + | <div align=left> | ||
===Results=== | ===Results=== | ||
| - | GFP secretion was confirmed by fluorescence emission and estimated to be approximately 10% of total protein, according to fluorescence levels. Significant levels of GFP fluorescence were found only in the supernatant of Paraquat induced cultures containing both the sensor/effector and the secretion systems but not in the non-induced cultures and in the cultures containing only the sensor/effector system | + | GFP secretion was confirmed by fluorescence emission and estimated to be approximately 10% of total protein, according to fluorescence levels found in bacterial cultures (See figure 2, from Device 2 testing). Significant levels of GFP fluorescence were found only in the supernatant of Paraquat induced cultures containing both the sensor/effector and the secretion systems but not in the non-induced cultures and in the cultures containing only the sensor/effector system. |
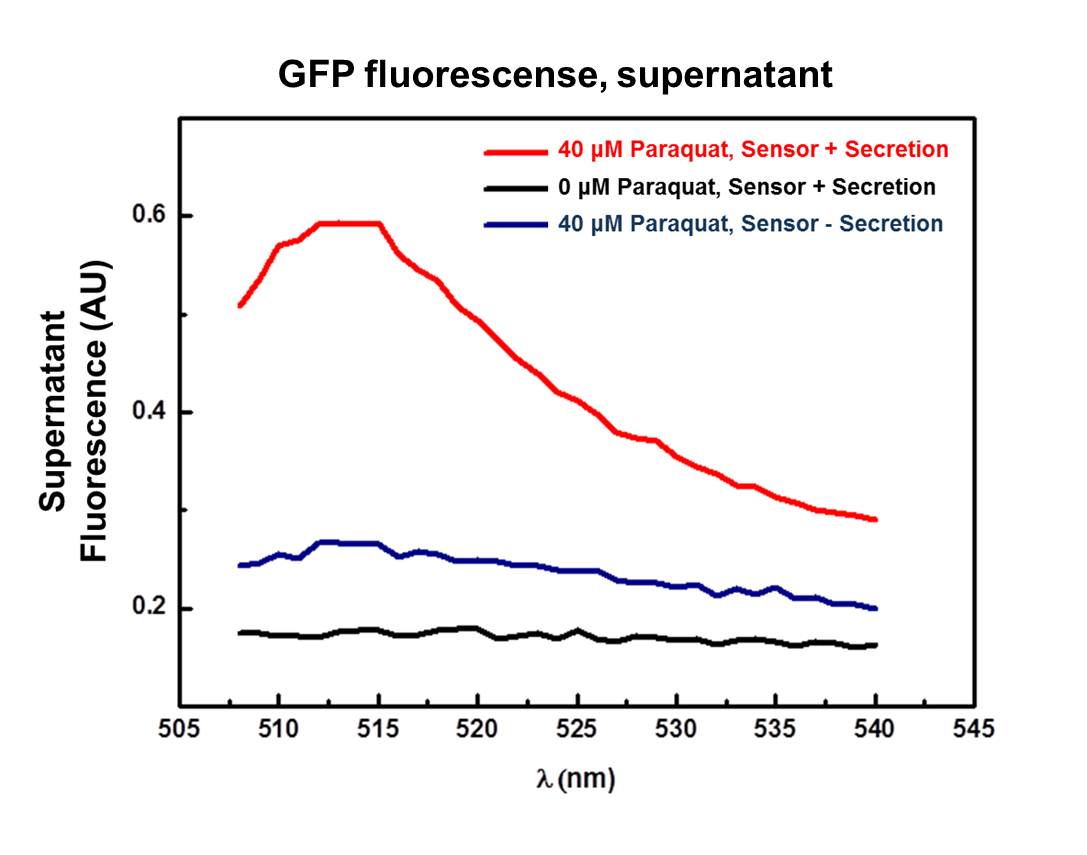
[[Image:Unicamp-emse-graph-secr.png|center|600px]] | [[Image:Unicamp-emse-graph-secr.png|center|600px]] | ||
| - | <div align=center>'''Figure | + | <div align=center>'''Figure 2: Supernatant fluorescence emission from 508 to 540 nm, excitation in 500 nm. Red line (40 μM Paraquat): sensor/effector with secretion system, showing a clear peak at 511 nm on fluorescence spectrum; Black line (0 μM): non-induced control, sensor/effector with secretion system, non-induced; Blue line (40 µM Paraquat): non-secretion control, sensor/effector without secretion system. |
| + | <div align=left> | ||
| + | |||
| + | |||
| + | According to the graphic (Figure 2), the supernatant of the culture harboring the complete secretion system and induced with 40 µM of Paraquat (Red line) exhibited a well-defined fluorescence peak between 508 and 520 nm, which indicates the presence of GFP in the supernatant probably due to secretion. | ||
| + | Validating this result, there is no clear fluorescence peak both in the supernatant of non-induced control spectrum (Black line - complete secretion system with 0 µM of Paraquat) and non-secretion control spectrum (Blue line – sensor/effector without secretion system). The aforementioned controls corroborates the result of GFP secretion though the secretion system, eliminating false positive results due to cell lysis for example. | ||
===Discussion=== | ===Discussion=== | ||
| - | + | This device is a modification of Stanford team 2009 secretion system composite [http://partsregistry.org/Part:BBa_K223062 BBa_K223062]. The secretion system was improved according to biobrick assembly standard 23, and modified by deletion of Hemolysin C (HlyC) which encodes a 170 aminoacids protein that is related to pathogenesis and has no secretion function. HlyC protein facilitates the activation of Hemolysin A protein (do not confuse with HlyA secretion signal peptide, used in fusion to GFP, and IL’s) (Fath et al, 1993). When HlyA protein is activated by HlyC enzyme it has cytotoxic and cytolytic effects, then HlyC is related only to pathogenesis and not secretion. | |
| - | + | We could show that this new secretion device (composed by secretion signal peptide HlyA, HlyD, HlyB and TolC) was able to secrete GFP fusioned to HlyA secretion peptide signal in response to Paraquat induction of the sensor/effector/secretion system, as showed by supernatant fluorescence. This also reinforces that HlyC is not necessary for secretion. | |
| - | + | ||
| + | By this experimental approach, we were able to validate the parts HlyB ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K554007 BBa_K554007]), HlyD ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K554008 BBa_K554008]), TolC ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K554009 BBa_K554009]), HlyA ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K554002 BBa_K554002] and SoxS HlyB HlyD TolC device ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K554013 K554013]). | ||
| + | You can find out more details regarding the differences between our device and 2009 Stanford's in a new section of our projects page, by clicking [https://2011.igem.org/Team:UNICAMP-EMSE_Brazil/Project#UNICAMP-EMSE_Brazil_Parts_Design:_why_are_they_different_to_Stanford_2009_team_ones.3F HERE!!]. | ||
| + | ==References== | ||
| + | Pomposiell P.J., Demple B. (2001). Redox-operated genetic switches: the SoxR and OxyR transcription factors. Trends in Biotechnology. 19(3):109-114. [http://www.ncbi.nlm.nih.gov/pubmed/11179804 Link to PubMed] | ||
| + | Fath MJ, Kolter R. ABC transporters: bacterial exporters. Microbiol Rev. 1993 Dec;57(4):995-1017. Review. [http://www.ncbi.nlm.nih.gov/pubmed/8302219 Link to PubMed] | ||
| + | <br><br><br><br><br> | ||
<font size="3" color=olive align=center>'''Special acknowledgement:'''</font><br><div align="center"><br> | <font size="3" color=olive align=center>'''Special acknowledgement:'''</font><br><div align="center"><br> | ||
<div align="left"> | <div align="left"> | ||
Latest revision as of 00:28, 29 October 2011

| Home | Project | Methods | Results | Data | Team | Notebook | Human Practices | Safety | Profile | Sponsors | Wix |

Contents |
Overview
Functional tests
The students assembled the devices subparts: the SoxR/SoxS sensor (Device 2, NO sensor device/ IL-10 producer), the Adrenaline sensor (Device 1, CA/AI-3 sensor device/ IL-12 producer), and the hemolysin secretion system (Device 3, Protein Secretion System).
The assembled devices related to oxidative-stress sensing ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K554010 SoxR transcription factor under control of a strong constitutive promoter - BBa_J23119]) and effector function in response to oxidative stress ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K554012 a generic protein fusioned with the C-terminal tail of Hemolysin A - HlyA -under control of the SoxS promoter, recognized by the SoxR transcription factor]) were tested under laboratory conditions using GFP as reporter.
Device 2 testing: SoxR/SoxS system regulating GFP production
The Device 2 is a modification of Stanford team anti-inflammatory device (iGEM 2009 - https://2009.igem.org/Team:Stanford/ProjectPage), which comprises SoxR gene (BBa_K223047) under control of a Constitutive Promoter (BBa_J23119) and SoxS promoter (BBa_K223001 – deleted part), coupled to human IL-10 gene (sequence designed by the team, improved for bacterial expression).
In order to test the ability of Jedi Bacteria in sensing NO levels and activating genes linked to SoxS promoter, we built a Device testing, with GFP linked to SoxS promoter, as it is shown in the following schema (Figure 1):
Methods
Competent E. coli DH5α strain cells were transformed with a pSB1A2 vector (Ampicillin resistant) carrying both the sensor (Strong_Constitutive_promoter + RBS + SoxR + Terminator) and the effector (SoxS_pormoter + RBS + GFP + HlyA + Terminator) devices using a chemical shock protocol. Transformed bacteria were plated on solid LB medium with 50 μg/mL Ampicillin and grown at 37ºC overnight. Surviving colonies were grown on liquid LB medium containing 50 μg/mL Ampicillin. Oxidative stress was induced by adding increasing concentrations of [http://en.wikipedia.org/wiki/Paraquat Paraquat] (Methyl viologen dichloride hydrate - Sigma), an oxidative stress inducer in bacteria. Final Paraquat concentrations were: 0 μM (control), 5 μM, 10 μM, 20 μM, 30 μM and 40 μM. Induction of the designed sensor/effector mechanism was accessed by fluorescence using a fluorometer (SLM – Aminco; 4 nm bandpass and 10 mm) with excitation in 500 nm and emission spectra from 508-550 nm.
In order to access if increasing concentrations of Paraquat could inhibit culture growth due to it´s toxicity, optical density (OD) levels of the culture were measured during the incubation time with Paraquat. The ability to recognize NO (nitric oxide), an inflammation signal molecule, was characterized for SoxR/SoxS sensor, and found to be FUNCTIONAL. In the section below we present the detailed results.
Results
Achieved results indicated that the designed sensor/effector device system was capable of inducible production of GFP (or another generic protein controlled by the sensor). In addition, protein induction can be modulated through varying inducer concentration (Figure 2). Higher concentrations of Paraquat exhibited higher fluorescence levels, which indicates increased GFP concentrations. No plateau was achieved using the highest tested concentrations.
Moreover, we tested if Paraquat could be toxic for the bacteria cells, and we found that the experimental concentrations of Paraquat did not show significant differences in cell growth as shown by OD levels in Figure 3.
Fluorescence Microscopy experiments also corroborates GFP production through Paraquat induction of SoxR/SoxS sensor/effector system. Fluorescence was detected in the cultures containing the sensor and GFP production devices when induced with 40 uM of Paraquat (Figure 5), but not in non-induced cultures(Figure 6). This is an additional evidence that the NO sensor system and Sox driven effector (in this case, GFP) synthesis worked as expected.
Discussion
The SoxR/SoxS system is one of the best characterized redox-sensing mechanisms in bacteria. Under oxidative stress conditions, activation of this system results in a cascade effect that ends in the activation of more than 16 genes that can counteract the harmful effect of superoxide and other oxidative radicals (Pomposiello and Demple 2001).
This device is a modification of Stanford team anti-inflammatory device (iGEM 2009 - https://2009.igem.org/Team:Stanford/ProjectPage), which comprises SoxR gene (BBa_K223047) under control of a Constitutive Promoter (BBa_J23119) and SoxS promoter (BBa_K223001 – deleted part). Although this system has already been designed and used in previously iGEM competitions (iGEM 2009 - https://2009.igem.org/Team:Stanford/ProjectPage), we took advantage of the availability of synthesized sequences to design new parts (SoxR and SoxS) that conforms the common biobrick standards in the iGEM registry. Our team has improved the parts as described in Innovation.
These newly designed parts were capable of sensing different concentrations of the inducer, resulting in an increased production of the protein under control of the SoxS promoter (in this case, GFP).
Under the experimental conditions, no cell growth inhibition was observed and confirmed by the calculations of the specific growth rate (μ). No significant difference was found in control experiment (0 μM of Paraquat) and cell induced with 40 μM of Paraquat, both presented μ= 0,18 h-1. Stanford 2009 team showed that Paraquat concentrations between 60 μM and 80 μM can inhibit E. coli cell growth due to enhanced toxicity, but these concentrations were not included in our tests.
With this procedure were able to validate the following parts: SoxS promoter ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K554000 BBa_K554000]), SoxR gene ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K554003 BBa_K554003]), SoxR device ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K554012 BBa_K554012]), SoxS GFP HlyA device [http://partsregistry.org/wiki/index.php?title=Part:BBa_K554012 BBa_K554012]
Device 3 testing, Protein Secretion System
The assembled devices related to secretion system (Hemolysin secretion system under control of SoxS – SoxS-HlyB-HlyD-TolC) were tested under laboratory conditions using GFP as reporter.
Methods
To access GFP secretion, competent E. coli DH5α strain cells were transformed simultaneously with a pSB1C3 vector (Chloramphenicol resistant) carrying both the sensor (Strong_Constitutive_promoter + RBS + SoxR + Terminator) and the effector (SoxS_promoter + RBS + GFP + HlyA + Terminator), and pSB1AK3 (Ampicillin resistant) carrying the secretion system (SoxS_promoter + RBS + HlyB + HlyD + TolC + Terminator). As a non-secretion control, E. coli harboring only the pSB1C3 vector with both sensor and effector systems was used. Oxidative stress was induced by adding [http://en.wikipedia.org/wiki/Paraquat Paraquat] (Methyl viologen dichloride hydrate - Sigma), an oxidative stress inducer in bacteria, according to the following conditions:
- Sensor/Effector + Secretion system: 0µM Paraquat
- Sensor/Effector + Secretion system: 40µM Paraquat
- Sensor/Effector - Secretion system: 40µM Paraquat (non-secretion control)
After 3 hours of induction, samples were collected, centrifuged (4000 rpm / 10 min; to avoid cell lysis) and the supernatant was collected and centrifuged again (13000 rpm / 10 min; to remove remaining cells). The supernatant fluorescence was measured in fluorometer (SLM – Aminco; 4 nm bandpass and 10 mm) with excitation in 500 nm and emission spectra from 508-550 nm.
Results
GFP secretion was confirmed by fluorescence emission and estimated to be approximately 10% of total protein, according to fluorescence levels found in bacterial cultures (See figure 2, from Device 2 testing). Significant levels of GFP fluorescence were found only in the supernatant of Paraquat induced cultures containing both the sensor/effector and the secretion systems but not in the non-induced cultures and in the cultures containing only the sensor/effector system.
According to the graphic (Figure 2), the supernatant of the culture harboring the complete secretion system and induced with 40 µM of Paraquat (Red line) exhibited a well-defined fluorescence peak between 508 and 520 nm, which indicates the presence of GFP in the supernatant probably due to secretion.
Validating this result, there is no clear fluorescence peak both in the supernatant of non-induced control spectrum (Black line - complete secretion system with 0 µM of Paraquat) and non-secretion control spectrum (Blue line – sensor/effector without secretion system). The aforementioned controls corroborates the result of GFP secretion though the secretion system, eliminating false positive results due to cell lysis for example.
Discussion
This device is a modification of Stanford team 2009 secretion system composite [http://partsregistry.org/Part:BBa_K223062 BBa_K223062]. The secretion system was improved according to biobrick assembly standard 23, and modified by deletion of Hemolysin C (HlyC) which encodes a 170 aminoacids protein that is related to pathogenesis and has no secretion function. HlyC protein facilitates the activation of Hemolysin A protein (do not confuse with HlyA secretion signal peptide, used in fusion to GFP, and IL’s) (Fath et al, 1993). When HlyA protein is activated by HlyC enzyme it has cytotoxic and cytolytic effects, then HlyC is related only to pathogenesis and not secretion.
We could show that this new secretion device (composed by secretion signal peptide HlyA, HlyD, HlyB and TolC) was able to secrete GFP fusioned to HlyA secretion peptide signal in response to Paraquat induction of the sensor/effector/secretion system, as showed by supernatant fluorescence. This also reinforces that HlyC is not necessary for secretion.
By this experimental approach, we were able to validate the parts HlyB ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K554007 BBa_K554007]), HlyD ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K554008 BBa_K554008]), TolC ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K554009 BBa_K554009]), HlyA ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K554002 BBa_K554002] and SoxS HlyB HlyD TolC device ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K554013 K554013]).
You can find out more details regarding the differences between our device and 2009 Stanford's in a new section of our projects page, by clicking HERE!!.
References
Pomposiell P.J., Demple B. (2001). Redox-operated genetic switches: the SoxR and OxyR transcription factors. Trends in Biotechnology. 19(3):109-114. [http://www.ncbi.nlm.nih.gov/pubmed/11179804 Link to PubMed]
Fath MJ, Kolter R. ABC transporters: bacterial exporters. Microbiol Rev. 1993 Dec;57(4):995-1017. Review. [http://www.ncbi.nlm.nih.gov/pubmed/8302219 Link to PubMed]
We would like to thank the following people for the support given in the testing experiments
- Msc. Viviane Cristina Heinzen da Silva, Laboratório de Bioquímica de Proteínas (LBqP) - IQ/Unicamp
- Msc. Gleidson Silva Teixeira, Laboratório de Genômica e Expressão (LGE) - IB/Unicamp
- Dr. Joan Grande Barau, Laboratório de Genômica e Expressão (LGE) - IB/Unicamp

 "
"