|
|

Wednesday, June 1
- Made agar with 3.7 g / 100 ml DI water
- Made 2 plates from 2 MG1655 strains received yesterday from Misra
- Submitted synthesis requests for RA, DA, and Seq1 from BioBasic
Thursday, June 2
- Meeting with Dr. Chang and her grad students from the School of Life Sciences to explain iGEM project.
- Single sided synthesis with phosphotase direction?
-
- Made liquid culture using LB Broth:
- 5 g peptone
- 2.5 g yeast extract
- 5 g NaCl
- 500 ml water
- Incubated and shook liquid culture overnight
Friday, June 3
- Confirmed placement of synthesis order
- Placed second large order for necessary laboratory materials
- Met with Barrett funding advisor
- Met with Jon, grad student from Misra's lab
- Lab:
- Resuspended part E0840 from well following parts registry protocol
- Followed "competent cells and chemical transformation procedure for DH5 alpha":
- Made 20mM concentration MgCl2 in shaken cells from yesterday
- Shook for 2 hours in 37 C room
Saturday, June 4
- Made 500 mL SOC following OpenWetWare protocol
- Transformed resuspended DNA into E. Coli
- Followed transformation protocol, but did not use water bath
Monday, June 6
- Transformed cells from Saturday, June 4th did not grow yet
- Test if competency procedure killed cells using following procedure:
-
- Repeat transformation using water bath instead of heat block:
- Thaw competent cells on ice
- 50 ul cells + 1 ul resuspended DNA, on ice for 30 minutes
- Heat shock cells at 42 in a water bath for 60 seconds
- Incubate on ice, 5 min
- Add 100 ul SOC to cells
- Shake at 37 C for 2 hours (11 am - 1 pm)
- Plate 20 ul, 200 ul (2 plates)
- Incubate overnight
Tuesday, June 7
- Amp plates did not grow
- Competent cells plated without amp grew
- New transformation using 2 different parts conducted did not work with either part:
-
- Control onto non amp plate to test if transformation killed cells showed that the cells were still viable
Wednesday, June 8
- Autoclaved lab materials to be sterilized
- Made 200 ml new SOB, 50 ml SOC
- New transformation protocols:
- Top10 chemically competent E. Coli from biodesign
- Part: BBa_E0840
- Using top10 protocol
- Plates: # 4 50ul, 5 150ul
- MG1655 plated from plate # 2:
- Plate: # 1
- From: 6-2 plate MG1655
-
Thursday, June 9
- Xiao introduced 3 grad students who can offer advice/assistance throughout the project
- We will meet with them (likely Thursdays @ 10am) to update them on our progress
- #4, 5 have colonies but no glow with UV - no promoter in biobrick part
- Need to add in a promoter
- Today:
- Add in promoter for GFP construct
-
-
- Use Knight restriction protocol
- Cut promoter BBa_J23101 with ECORI, SPEI
- Cut GFP generator BBa_e0840 with ECORI, XHOI
- DNA extraction- use "ethanol precipitation of nucleic acid" procedure
- Ligate restriction products
- Transform ligation products
- Create stock of competent cells
- Order Top10 cells (what strain are these?)
- Make glycerol stock of BioBrick
- Jon/Misra procedure for competent cells and transformation:
- Overnight culture from previous:
- Diluted 1 to 50
- Shook 1 hr in 37 C room
Friday, June 10
- 2 competency procedures (Jon, CCMB80)
- 3 transformation procedures (Jon, CCMB80, top10 from biodesign)
- 12 plates made (see lab notebook)
- Autoclaved glass test tubes
- Dan demonstrated to lab members how to make glycerol stocks
- Bought top10 competent cells
- Got account set up (still need to create Sunrise account)
-
- Met with James Alling, who is a JD-PhD interested in helping us out
- He is very good at speaking and could help with presentation later
- Very attracted to promoting big picture of project
- Kylie determined new primers after noticing that we don't need Cas 1,2,3 for our natural cas construct (from "structural basis for CRISPR")
- This brings Cas construct size down to a total of 3.8kb instead of over 5kb
- We will try both ways, and see if cas 1 and 2 do anything interesting
- We are considering getting primers for each individual Cas gene (Cas A, Cas B, etc...)
- Biobasic is taking twice as long as they advertised (no DNA until june 20?)
- From now on we will go through IDT due to slow turnaround from BioBasic
- We contacted BioBasic about discount, got a synthesis price reduction
Saturday, June 11
HAPPY 22ND BIRTHDAY KEITH!
- Created idea web for CRISPR in MindNode
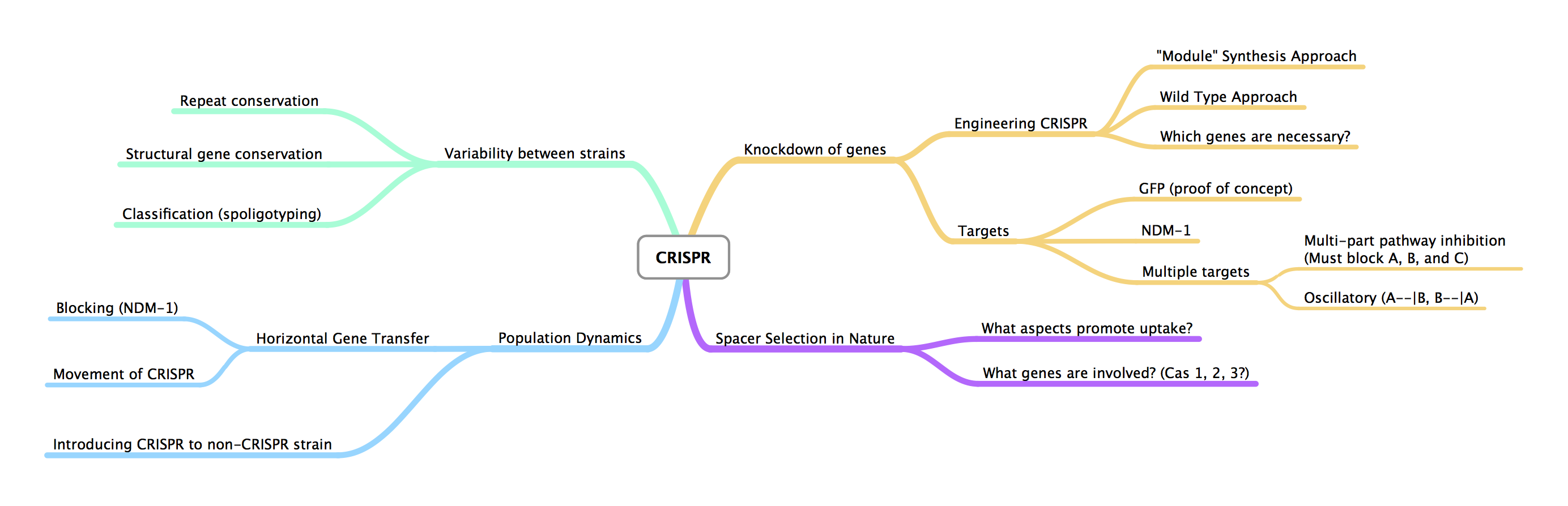
Tuesday, June 14 - Friday, June 17
- Synthetic Biology 5.0 conference
Monday, June 20
- Prepared overnight culture x 2 (LB) for DNA miniprepping
- Prepared overnight culture x 2 (LB, SOC) for culture
-
- Carry out genome prep for K12 genome from MG1655
- Begin PCR amplification of Cas genes
- Create and test competent cells
Tuesday, June 21
-
- Ruben and Ethan carried out this competency procedure
- Made 250 ml SOB
- 9 210ul tubes placed into -80 C fridge
- plates made:
- 1. LB, transformation: DA
- 2. LB + amp, transformation: DA
- 3. LB + amp, transformation: PUC19
- TSS procedure w/ BL21 cells
- Madeline, Juan, and Keith carried out this competency procedure
- plates made:
- 4. LB + amp, PUC19, burned
- 5. LB + amp, PUC19, unburned
- 6. LB + amp, DA
- 7. LB + amp, DA
- 8. LB + +amp, DA
- NEB top10 competency test:
-
- 9. LB + amp, PU19
- 10. LB + amp, PUC19
- 11. LB + amp, DA
-
- First PCR conducted overnight of attempting to amplify CAS genes A-E and 3
Wednesday, June 22
- Made new LB + amp stock
- Made 200 ml LB + amp broth
- Results from plates made yesterday:
- Ampicillin stock integrity in question
- 1: normal growth (no distinct colonies)
- 2: colonies
- 3: no growth
- 4: no growth
- 5: no growth
- 6: colonies
- 7: colonies
- 8: colonies
- 9: very heavy colonies
- 10: very heavy colonies
- 11: light colonies
- Overnight cultures made of 2, 6, 7, 8, 11 (3 each in LB + amp broth)
- Ran a gel of PCR product
- New plates:
- 1. CCMB80, LB, PUC19
- 2. CCMB80, LB, DB
- 3. CCMB80, LB, SEQ1
- 4. CCMB80, LB + amp, no plasmid
- 5. CCMB80, LB + amp, PUC19
- 6. CCMB80, LB + amp, DB
- 7. CCMB80, LB + amp, SEQ1
- 8. NEB, LB, PUC19
- 9. NEB, LB, DB
- 10. NEB, LB, SEQ1
- 11. NEB, LB, no plasmid
- 12. NEB, LB + amp, no plasmid
- 13. NEB, LB + amp, PUC19
- 14. NEB, LB + amp, DB
- 15. NEB, LB + amp, SEQ1
- 16. TSS, LB, PUC19
- 17. TSS, LB, DB
- 18. TSS, LB, SEQ1
- 19. TSS, LB, no plasmid
- 20. TSS, LB + amp, no plasmid
- 21. TSS, LB + amp, PUC19
- 22. TSS, LB + amp, SEQ1
- 23. TSS, LB + amp, DB
Thursday, June 23
- Plates from yesterday worked completely as expected
Plate 6:
Plate 15:
- Overnight culture in amp grew
- Today:
- Glycerol stock made of DA
- Overnight cultures made of DB, sEQ1 from plates
- Ran gel of PCR from last night
- DNA extraction 2x (elution)- verified using nanodrop, did not get enough to be successful
- Another overnight PCR using different settings
- Designed new primers for casA-E + cas3
Friday, June 24
- Ran gel from PCR carried out last night
-
- DNA extraction using spin method from Miniprep protocol(DB, SEQ1)
-
- We went through hassle of ordering new Cas primers from IDT
- Ordered a pair of primers for each Cas gene - this way we can customize and perhaps PCR out in sections
Saturday, June 25
- Transformations of DA, DB, and Seq1 into the BioBrick ampR vector (pSB1A3) into TSS and NEB cells was successful
- Made overnight liquid culture to miniprep tomorrow
Sunday, June 26
- Made LB amp plates
- Conducted restriction digest...
- We used the wrong enzymes! used EX and EP instead of EX and ES
- Didn't linearize plasmid before running results on a gel
Monday, June 27
- We identified the Top 10 lab techniques to learn and love
- Dan emphasized that we need to be independent and know these!
- Two methods for restriction: Ginkgo bioworks (two bricks into desired plasmid) and traditional (EX and ES)
- DA: ES, EX
- Seq1: ES, XP
- PSB1A3: EX, EP
- 1) did not let gel dry completely before removing comb
- 2) too much voltage caused gel deformation
- Lesson learned: don't use it directly! must grow it up first
- Ordered more from iGEM HQ
- We rehydrated cells and let culture grow overnight in tryptic soy broth
- Made overnight cultures of Seq1, DA, DB, and E0840
- Overall message: Not a great day in terms of results, but many tough lessons learned.
Tuesday, June 28
- B. halodurans developments
- Cells retrieved from overnight culture
- Made 4 plates on tryptic soy media, as well as 7 more tryptic soy plates
- Made one glycerol stock
- Conducted Genomic prep + PCR using primers R1 and R2
- Nanodrop new record! 220ng/ul template DNA
- "Ode to Trinette" Haiku by Joseph Flay
- PCR is hard
- Trinette, you are so thermal
- Thanks for the fun times
- Conducted miniprep of Seq1, DA, DB, E0840
- Nanodrop results (see Kylie's notebook)
- Further restriction digests
- Made "restriction supermix" of water, BSA, NEB4 (1x, enough for 30 digests)
- Seq 1: ES, EX
- DA: ES, EX
- DB: ES, EX
- E0840: EP
- Made and ran a large gel
- Problem: used wrong hyperladder (used I instead of II)
- Successfully isolated: Seq 1 (ES), Seq 1 (EX), DB (EX), E0840 (insert), E0840 (vector)
- Unsuccessful: DA (ES), DA (EX), DB (ES)
- Transformed RA, RB into NEB cells (no control)
- Replated BL21DE3 x 1 and MG1655 x 1 on LB Agar
- Moved plates from 4 degree room to small fridge in lab because they are fixing the room tomorrow (and got rid of some old plates)
- Took lab inventory (mostly)
- Made more overnight cultures:
-
- Other notes: Paul Johnson sent us a nice message basically saying that as long as we can justify it, iGEM is here to stay
- (meaning they will keep funding the team in the coming years)).
- We also talked about getting FURI and SOLUR funding for next year's team.
- An REU proposal was discussed, but ultimately abandoned because a majority of the team's students would need to be from outside ASU, which we don't want.
- Overall: people kept very busy, we worked well in teams, however we need to make sure we are really paying attention to what we do - mistakes cost time and money!
- Tomorrow: plan on ligation, check PCR results, run a gel for PCR results, order primers for B. halodurans, try restriction of DA again, miniprep and try restriction of RA/RB
Wednesday, June 29
- Plates from last night (see pictures):
- LB + AMP + RA
- LB + AMP + RA
- LB + AMP + RB
- LB + AMP + RB
- Restriction digest of DA, DB (2x)
- Run a gel: CMR product from BH PCR
- Conducted gel extraction, submitted extracted DNA for sequencing
- Very low yield (~20 ng/uL)
Thursday, June 30
- New primers arrived from IDT for second round of attempts at getting the cas genes out of MG1655
- After successful isolation of what looks like the CMR genes from Bacillus halodurans, a second attempt was run overnight
- Got our sequence data from last night for CMR genes: looks like we successfully amplified CMR!
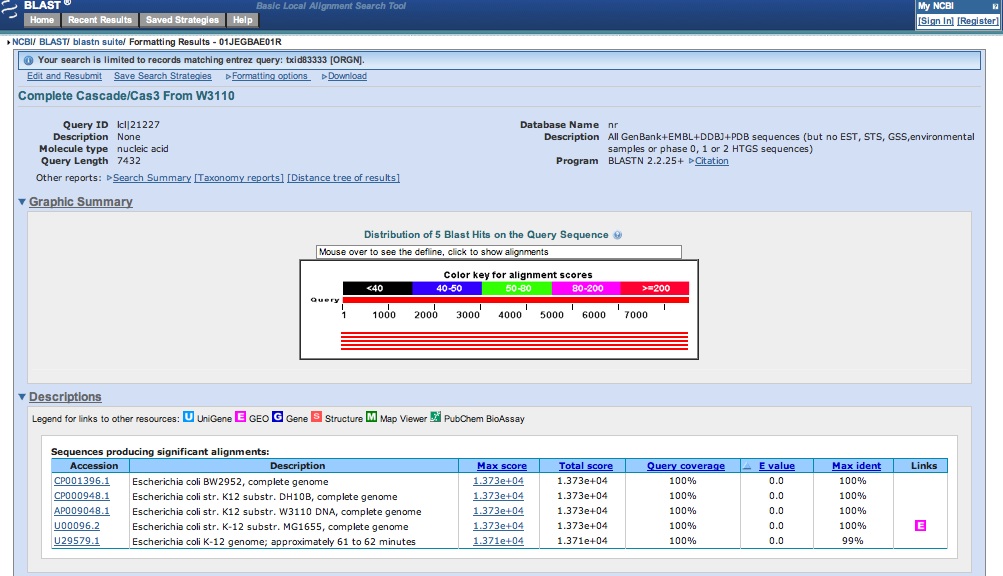
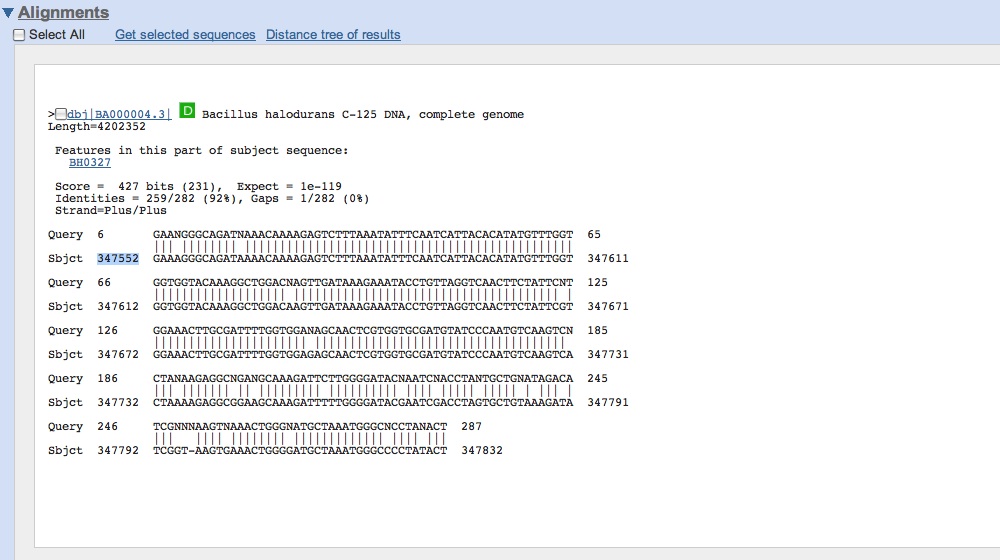
- Cultures of RA/RB grew well
- Conducted miniprep of RA x2, RB x2
- DA ES EX (2x)
- DB ES XP (2x)
- RA ES EX XP (2x)
- RB ES EX XP (2x)
|