Team:Harvard/Results
From 2011.igem.org
(→Selection Strain) |
|||
| Line 19: | Line 19: | ||
=Selection Strain= | =Selection Strain= | ||
| + | We designed a one-hybrid metabolic system that was entirely genome-based. Using multiplex automated genome engineering (MAGE) and lambda red, we knocked out HisB, PyrF, and rpoZ; inserted a kanamycin cassette-zinc finger binding site-His3-URA3 construct into the 1529620 locus; and changed the zinc finger binding site directly on the genome. The strain was fully characterized and was sensitive enough to recognize a valid zinc finger when diluted as much as one into one million of negative controls. | ||
=Zinc Finger Binders= | =Zinc Finger Binders= | ||
</div> | </div> | ||
Revision as of 20:23, 25 September 2011
Overview | MAGE | Lambda Red| Chip-Based Library | One-Hybrid Selection | Zinc Finger Binders | Biobricks
Bioinformatics
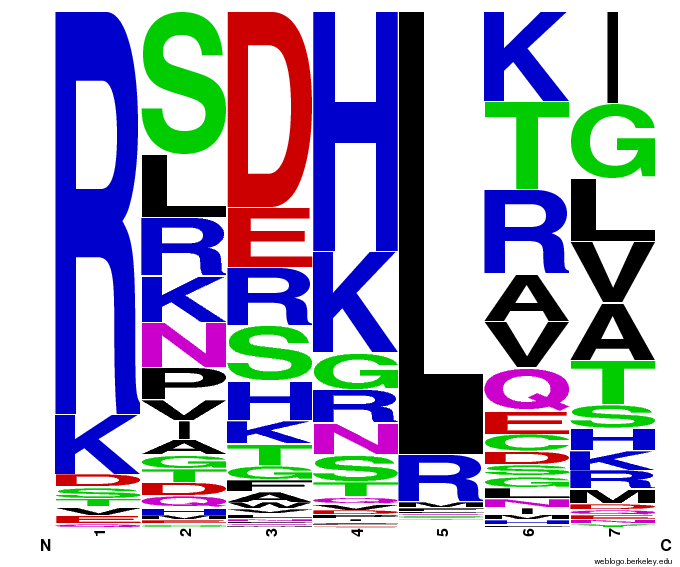
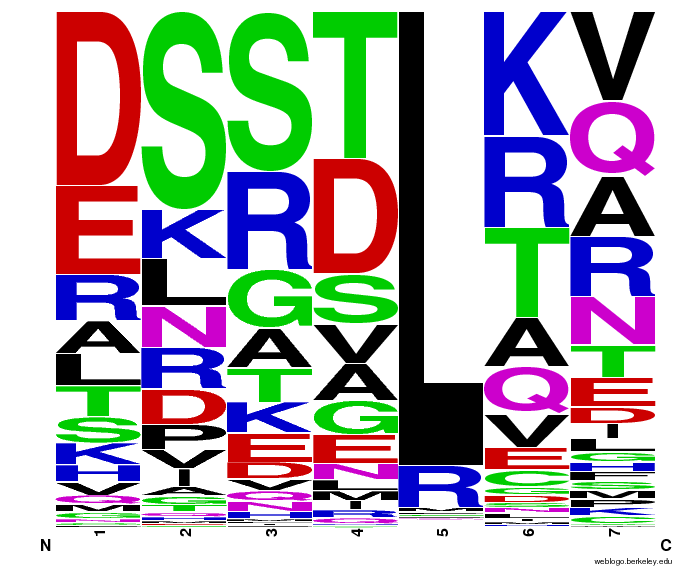
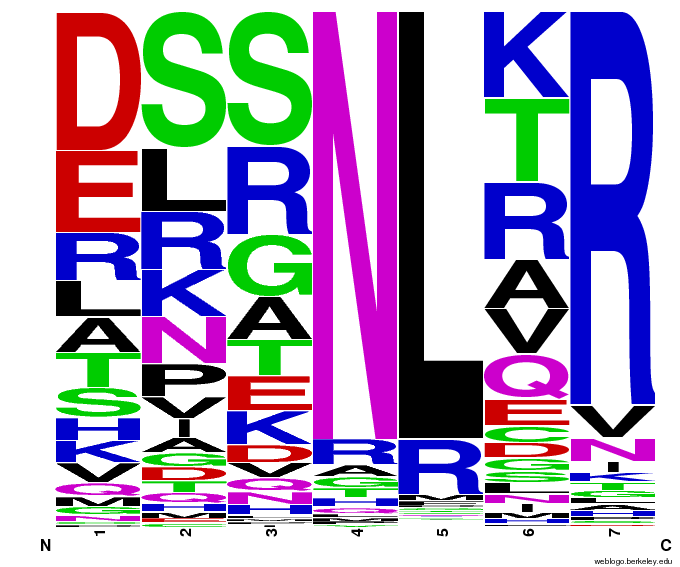
55,000 Possible Zinc Fingers
We made 55,000 sequences, distributed evenly among 6 DNA target triplets. That's 9150 per target.
Because our program's output changes dramatically based on the input triplet, no two sets of sequences are the same:
Selection Strain
We designed a one-hybrid metabolic system that was entirely genome-based. Using multiplex automated genome engineering (MAGE) and lambda red, we knocked out HisB, PyrF, and rpoZ; inserted a kanamycin cassette-zinc finger binding site-His3-URA3 construct into the 1529620 locus; and changed the zinc finger binding site directly on the genome. The strain was fully characterized and was sensitive enough to recognize a valid zinc finger when diluted as much as one into one million of negative controls.
 "
"