Team:UNAM-Genomics Mexico/Project/RhizobialKit
From 2011.igem.org
Contents |
RhizoKit General Overview
Abstract
Expanding our ability to manipulate and engineer biological parts in new organisms is an ever present goal of synthetic biology. With this idea in mind we set out to develop a kit of biological tools to enable working with BioBricks in Rhizobial species. We developed the tools in this kit with our model chassis being Rhizobium etli, a member of the Rhizobiaceae family of nitrogen fixating bacteria, that establishes a symbiotic relationship with legumes within the root nodules.
RhizoKit Description
The RhizoKit will set the ground for the use of Rhizobial species, especially R. etli, as organisms available for the design and construction of novel synthetic biological machines. The kit will be composed of parts with distinct molecular functions that will increase the repertoire of possible design and engineering approaches and possibilities in Rhizobial species.
The parts that we will develop are the following:
- The nifH promoter. This promoter sequence comes from the promoter region of the nifHa gene in R. etli. This promoter region contains a σ54-dependent promoter located between position -24 and -12 relative to the transcription start site, as well as a binding site for NifA, a positive transcriptional regulator. The binding of the NifA activator to the promoter region promotes transcription in conditions of low oxygen concentration, and the -24/-12 conserved bases have been implicated in the context of nitrogen fixation and regulation (Beat Thöny, Hauke Hennecke, 1989; Valderrama, B., et. al., 1996).
- A re-characterization of the Anderson Promoter Collection in R. etli. These constitutive promoters are already characterized in E. coli [1], and our goal is to investigate the strength of these promoters in R. etli as the chasis. This will hopefully give a spectrum of promoters with different strengths that will allow the fine-tuning of the expression of devices in this model.
- We developed a standardized plasmid that is able to integrate BioBrick constructions into Rhizobial species after conjugation with transformed E. coli cells. This plasmid is a modification of the pBBR1MCS-5 plasmid, which is a broad-host-range (bhr) vector designed to assist genetic anaylsis in prokaryotes, specifically for Gram- bacteria that are naturally CmR (chloramphenicol resistant). It is used to transfer genes from E. coli to R. etli by first transforming E. coli S17 competent cells with this vector containing an RFP reporter and later performing a conjugation between the transformed E. coli cells and R. etli CFN42.
- A repC “plasmid replication device” as a standard part will be included. Its purpose is to render a plasmid capable of repC-dependent replication in R. etli. The repC-ctRNA region contains two genes: one encoding the initiator protein RepC (46.8 kDa) and other, an antisense RNA (67 nt). repC is the minimal region required for stable replication of a member of the repC plasmid family.
- A plasmid containing the repC-ctRNA region as part of its backbone is under construction. This plasmid, generated from the pSB1AK3 backbone, will harbor the repC system and the assembly suffix and prefix separately, saving the steps of the repC-ctRNA assembly.
RhizoKit Details
Parts
- nifH promoter. This promoter was amplified from Rhizobium etli CE3 using primers that overlapped with 25 base pairs at the start and at end of the promoter region. The upper, or forward, primer included the prefix sequence, while the lower primer contained the suffix sequence. In this way we achieved standardization of the nifH promoter region for further manipulation.Promoters Logbook.
- pBBR1MCS-5 derived plasmid. This is a general scheme showing the methodology used to standardize the plasmid, the strategy is based on inserting a segment of the pSB1T3 which contains the prefix, suffix and terminators into the pBBR1MCS-5. pBBR1MCS-5 Logbook.
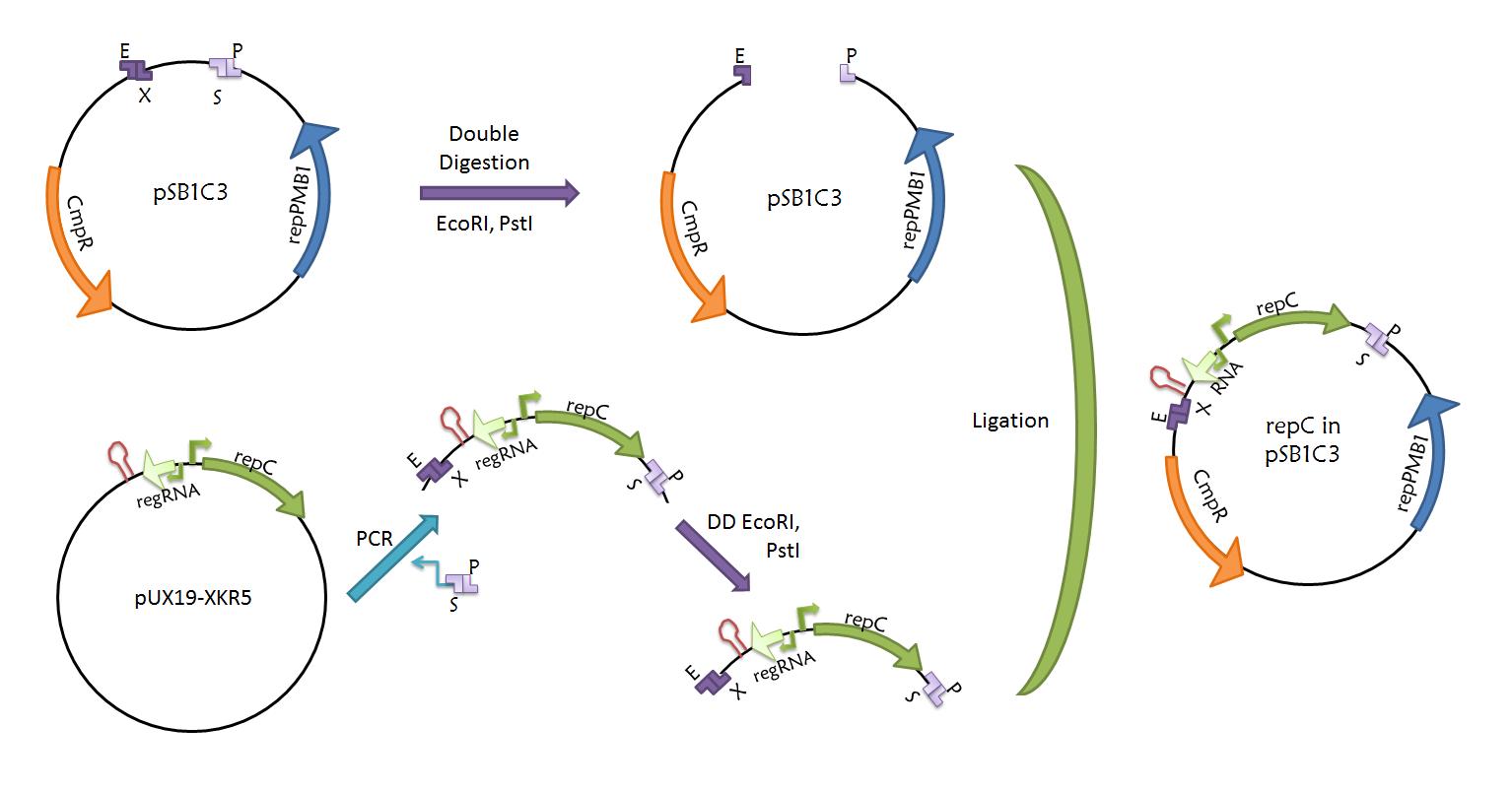
- repC-ctRNA part. The repC-ctRNA region contained in plasmid pUX19-XKR5 was amplified using oligonucleotides with prefix and suffix hangovers in a PCR. We verified that the region contained no standard restriction enzyme sites. The amplification will be double digested with EcoRI and PstI, and inserted into plasmid pSB1C3. RepC logbook.
- repC-ctRNA plasmid. The repC-ctRNA part was digested with EcoRI and SpeI and inserted into pSB1AK3 with B0015 digested with EcoRI and XbaI, forming a composite repC-ctRNA-B0015. Since the ctRNA contains its own transcriptional terminator it is not necessary to add a terminator upstream of the repC-ctRNA part.
A similar procedure was used to compose P1004 (the chloramphenicol resistance) and the B0014 bidirectional terminator in plasmid pSB1AK3. The composites will be fused once more, forming a part with the following ordered subparts: prefix, ctRNA, repC ORF, B0015, P1004, B0014, suffix. This composite will be in turn amplified with primers with XhoI sites as hangovers, skipping both prefix and suffix. The amplification and plasmid pSB1C3 will be digested with XhoI, and ligated to yield a plasmid with chloramphenicol resistance and the regulatory coding region repC-ctRNA with the corresponding transcriptional terminators. RepC logbook.
Characterizations
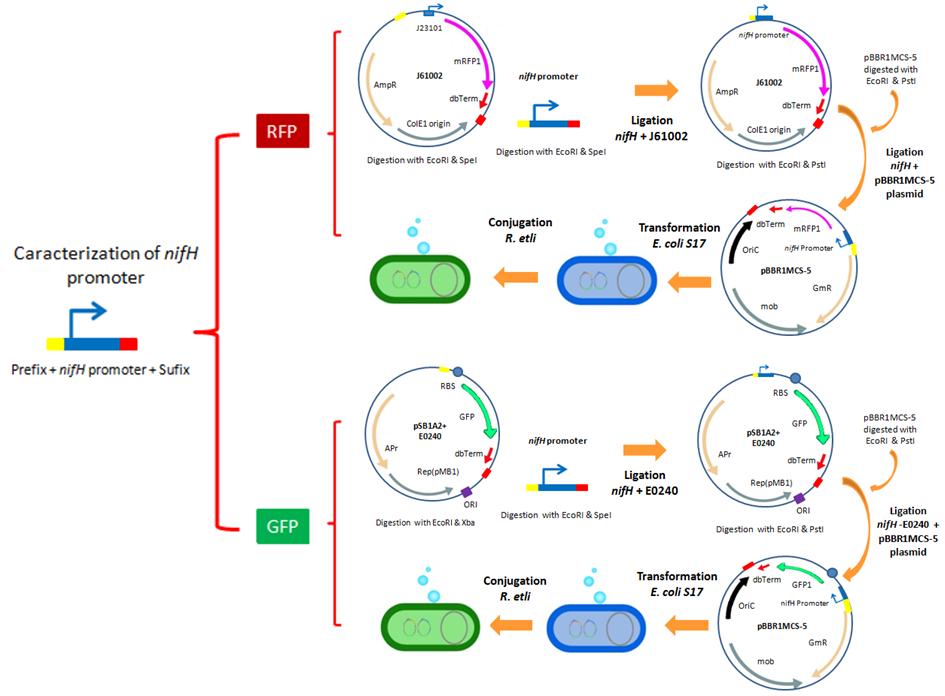
- For the characterization of the nifH promoter as well as the promoters from the Anderson collection, we designed a strategy that involves the use of three reporters: a GFP, RFP and β-Glucuronidase. In the following images, we illustrate the experimental strategies designed for the construction of our GFP and RFP reporter systems.
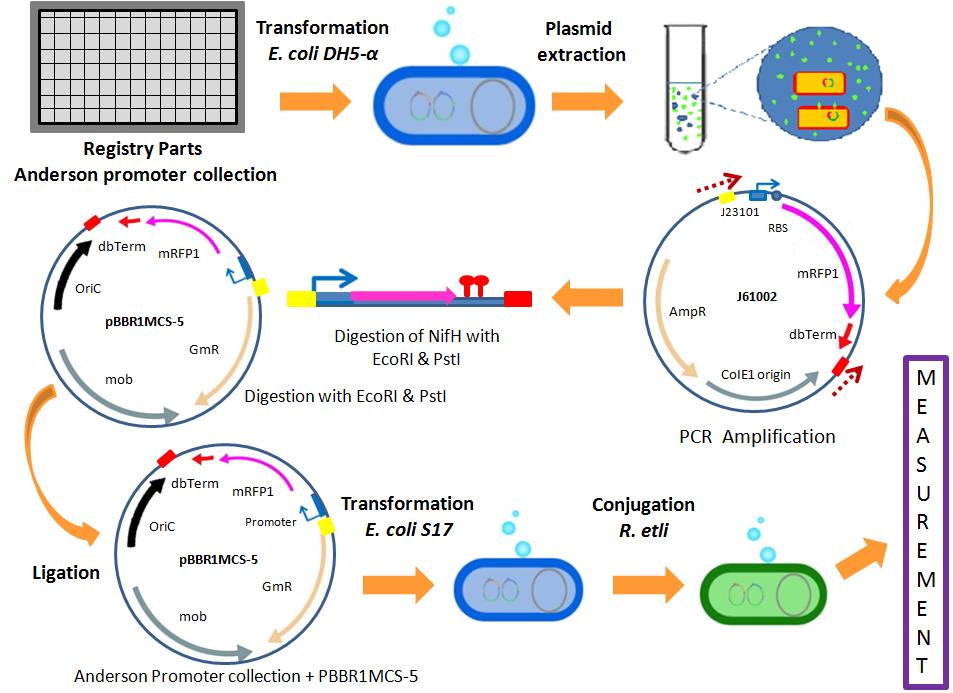
- The integration of the Anderson promoter collection into our standardized was designed as shown in the following image.
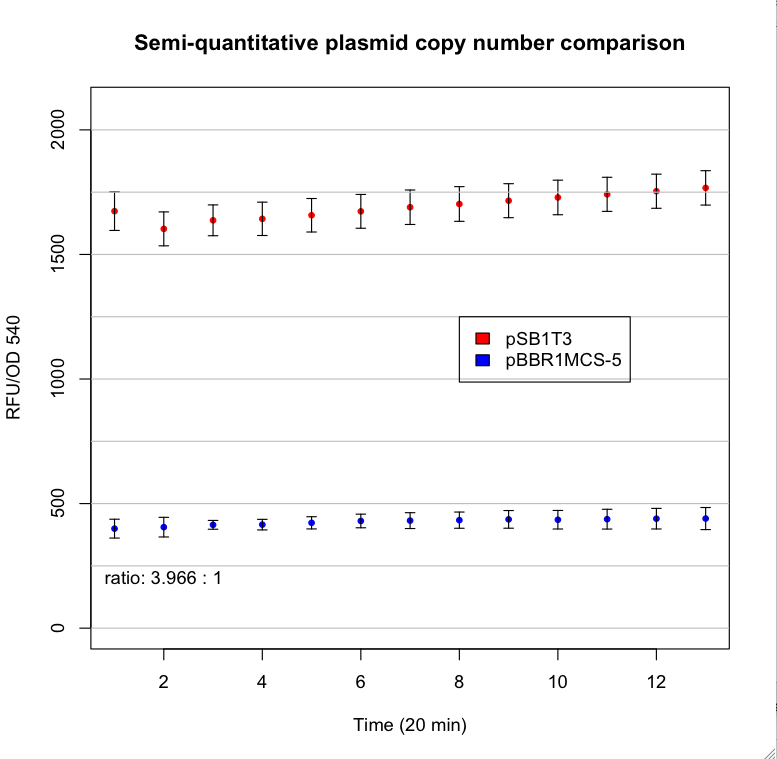
This are the results:
See also the promoters Logbook
Justification
The main goal of our project is to produce molecular hydrogen as an energy transporter in R. etli cells in a auto-sustainable fashion. While pursuing this objective we faced the necessity of generating basic standard tools to systematically characterize the components of our system. It became clear that expanding the repertoire of standard parts available for their use in our model system, R. etli, was not only beneficial to our causes, but touched on a basic purpose of synthetic biology: broadening the range of organisms in which rationalized engineering approaches help develop and characterize biological devices. Our model chassis and related species stand out for their nitrogen fixation capacity, and the transcriptional regulation dependent on anaerobic or hypoxic conditions. Thus, with the development of these bioengineering tools, we are extending the reach of synthetic biology to new crowds, questions, and horizons.
References
- Brenda Valderrama, Araceli Dávalos, Lourdes Girard, Enrique Morett. 1996. Regulatory proteins and cis-acting elements involved in the transcriptional control of Rhizobium etli reiterated NifH genes. Journal of Bacteorology, 178, 3119-3126.
- Beat Thöny, Hauke Hennecke. 1989. The -24/-12 promoter comes of age. Microbiology Reviews 63, 341-358.
- Michael E. Kovach, Philip H. Elzer, D. Steven Hill, Gregory T. Robertson, Michael A. Farris,R. Martin Roop, Kenneth M. Peterson (1995) Four new derivatives of the broad-host-range cloning vector pBBR1MCS, carrying different antibiotic-resistance cassettes. Gene 166, 175-176
- Javier Izquierdo. 2005. An antisense RNA plays a central role in the replication control of a repC plasmid. Plasmid 54, 259–277
 "
"