Team:Berkeley/Project
From 2011.igem.org
Gabriellopez (Talk | contribs) |
|||
| Line 58: | Line 58: | ||
<div align="justify"> | <div align="justify"> | ||
| - | The question of how cells detect specific molecules has always been important to biology, but is becoming an increasingly useful question to ask as we | + | The question of how cells detect specific molecules has always been important to biology, but is becoming an increasingly useful question to ask as we begin to engineer life. The applications of biosensing are widespread. There is a demand for biological diagnostic tools in medical or environmental applications, like detecting diseases or environmental contaminants. Sensing also has applications in biosecurity systems, when bioengineers try to develop strains that are stably dependent on the presence certain molecules. Finally, developing sensing systems for specific molecules is invaluable in metabolic engineering, because it allows for construction of metabolite-specific selection schemes. </p> |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | because it allows for construction of specific selection | + | |
<center> | <center> | ||
<p><img src="https://static.igem.org/mediawiki/2011/d/dc/Wiki_-_Biosensors4.png" width="500" align ="middle" /></p> | <p><img src="https://static.igem.org/mediawiki/2011/d/dc/Wiki_-_Biosensors4.png" width="500" align ="middle" /></p> | ||
</center> | </center> | ||
<p> | <p> | ||
| - | Traditionally, | + | Traditionally, most sensing systems in synthetic constructs have been borrowed from nature. However, very often researchers want to detect a ligand that does not have a receptor in the existing list of characterized sensors. Scanning natural systems for proteins that bind to a specific molecules is difficult and time-consuming, when it is not impossible. As synthetic biology realizes more and more unnatural functions, we will want to be able to design cells that can detect molecules that life has never before responded to. This is something we cannot achieve using natural receptors.</p><p> |
<center> | <center> | ||
<img src="https://static.igem.org/mediawiki/2011/6/6b/Wiki_-_modular_biosensor_4.png" width="500" align ="middle" /></p> | <img src="https://static.igem.org/mediawiki/2011/6/6b/Wiki_-_modular_biosensor_4.png" width="500" align ="middle" /></p> | ||
</center> | </center> | ||
| - | <p>Here at Berkeley we were motivated to | + | <p>Here at Berkeley we were motivated to address this problem by constructing a biosensing mechanism that could be readily adapted to the ligand of choice. We wanted to build a modular system that could be modified through the exchange of binding domains. This would provide a standardized method of building biosensors with both natural binding domains, optimized by evolution, and engineered domains that can respond to molecules outside the range of natural receptors.</p><p> |
</div> | </div> | ||
Revision as of 23:28, 27 September 2011


Traditionally, most sensing systems in synthetic constructs have been borrowed from nature. However, very often researchers want to detect a ligand that does not have a receptor in the existing list of characterized sensors. Scanning natural systems for proteins that bind to a specific molecules is difficult and time-consuming, when it is not impossible. As synthetic biology realizes more and more unnatural functions, we will want to be able to design cells that can detect molecules that life has never before responded to. This is something we cannot achieve using natural receptors.

Here at Berkeley we were motivated to address this problem by constructing a biosensing mechanism that could be readily adapted to the ligand of choice. We wanted to build a modular system that could be modified through the exchange of binding domains. This would provide a standardized method of building biosensors with both natural binding domains, optimized by evolution, and engineered domains that can respond to molecules outside the range of natural receptors.
ToxR is a characterized transmembrane transcription factor from Vibrio cholerae with active domains in both the periplasm and cytoplasm. Natively, it is involved in the transcription of a number of virulence factors, including the two subunits of the cholera toxin ctxAB and the TCP pilus. It is activated in trans by ToxS, a membrane-anchored periplasmic protein coded by the gene directly downstream of toxR. Active ToxS in the periplasm stabilizes the dimerization of the periplasmic domain of ToxR. The association transfers through the membrane, coupling the cytoplasmic domains of ToxR to form an active homodimer. The active ToxR drives transcription from the ctx promoter by binding to TTTGAT repeats and recruiting transcription machinery.
ToxR elegantly allows Vibrio cholerae rapid sensing and the activation of pathogenic functions. Virulence factors such as those controlled by this system are often clustered in the genome within pathogenicity islands, remnants from a past horizontal gene transfer. The simplicity and orthogonality of the systems contained in these small virulence cassettes make them an ideal source for modular biosynthetic tools. Indeed, one of the reasons ToxR is so interesting is that it single handedly achieves the task of the standard two-component signal transduction pathway: it is activated in the periplasm and directly promotes transcription in the cytoplasm. The simplicity of this regulation system and the scope of this single protein make it an ideal starting point for synthetic design.
Dimerization of ToxR has been found to activate the ctx promoter in Vibrio cholerae by binding directly to the DNA. DiRita et. al. (1991). The periplasmic domain detects changes in the environment and facilitates the dimerization of the cytoplasmic domain, which activates transcription. These domains are all on one peptide, so signals can be quickly relayed from the periplasm to the cytoplasm. This unique aspect of ToxR makes it a good part for building a biosensor. We aim to make a biosensor by utilizing this dimerization-dependent transcriptional activation feature of the ToxR system.



Our initial step was to validate that a ToxR based two-hybrid system could be expressed in E. Coli. We first truncated ToxR to eliminate the periplasmic domain because it is only the dimerization of the cytoplasmic domain that controls transcription. Next, we attached the constitutively dimerizing proteins IILK to these ToxR truncates such that the constitutively dimerizing proteins were located in the periplasm. IILK is a leucine zipper that is well characterized to be a constitutive dimer.


In order to resolve the toxicity of ToxR, we constructed a new design principle. Stress is a natural regulatory cue in bacteria that dictates changes in behavior or expression levels. We decided to look into the E. coli genome for any type of stress based down regulatory elements. Toxic over expression of ToxR would be down regulated with this negative feedback system. With this method, we are able to express ToxR chimeras at optimal levels without it being toxic to the cell.

Stress is a common regulatory cue that allows organisms to adjust to variable conditions in the environment. As a result, stress-responsive regulatory elements are prevalent motifs in nature. Due to the toxicity caused by the toxR chimeras in E. coli, a method of regulating its expression to produce the maximum amount of ToxR chimera without killing the cell was needed. We hypothesized that the E. coli genome likely contains such stress responsive regulatory systems from which regulatory feedback systems can be constructed. Expression of the ToxR chimera to toxic levels by stress promoters can be downregulated by stress in a feedback loop. This maintains a high but nonlethal level of expression of the toxic gene.

In order to construct a feedback system, we analyzed microarray data of the E. coli genome for regulatory elements downregulated by stress. Moen, et. al. (2009) subjected E. coli to generalized stresses and measured the global mRNA levels of all genome elements for stress response. From their data, we identified 35 stress promoters that were downregulated under all stress conditions tested for further characterization. We isolated the region of the genome upstream of the ORF through PCR and placed the resulting pool of 35 stress promoters in front of GFP.

The construct was transformed into E. coli and subjected to general stress conditions. We then measured the fluorescence level and looked for decreased fluorescence. While not all promoters responded by downregulating GFP expression, a few did respond, with the cold condition yielding the best data. The fluorescence measurements from the Tecan plate reader were confirmed by flow cytometry.


Since some of the stress promoters displayed the desired behavior, we proceeded to incorporate the stress promoter pool into a stress response system with ToxR. In our stress feedback system, expression of the ToxR chimera activates the ctx promoter (Pctx), driving GFP production, and creates cellular stress. The stress promoter responds to stress by downregulating ToxR-IILK expression to a nonlethal level, allowing the cell to maintain a high level of toxR chimera expression without toxicity.

In order to ensure nonlethal expression of the toxR chimera, we created an rbs library for each of the 35 stress promoters to construct a promoter-rbs library in front of ToxR-IILK. We transformed the plasmid into cells containing Pctx.GFP and grew them up to assay the efficacy of the stress responsive feedback system. In our growth assay, we looked for large, healthy colonies as well as strong fluorescence, indicators of robust growth and desired expression levels.

From our combinatorial approach towards the construction of the stress-responsive feedback system, we were able to obtain big, healthy green colonies growing on our plates. The observation of big colonies indicated that our cells were healthy and not experiencing any apparent cellular stress. The high levels of GFP observed in the colonies indicated that cells were expressing enough ToxR chimera to strongly drive transcription of the ctx promoter (Pctx). Since we saw colonies that were big and green, we claim to have successfully expressed our toxic ToxR fusion proteins to the highest levels in order to ensure strong transcriptional output while minimizing cellular stress.

We picked big green colonies and grew them in the same conditions as we had grown the pBAD versions of the parts, measuring OD600 and RFU with a Tecan plate reader every five minutes. The improved fluorescence is noticeable and cells harboring this plasmid behave much more predictably.
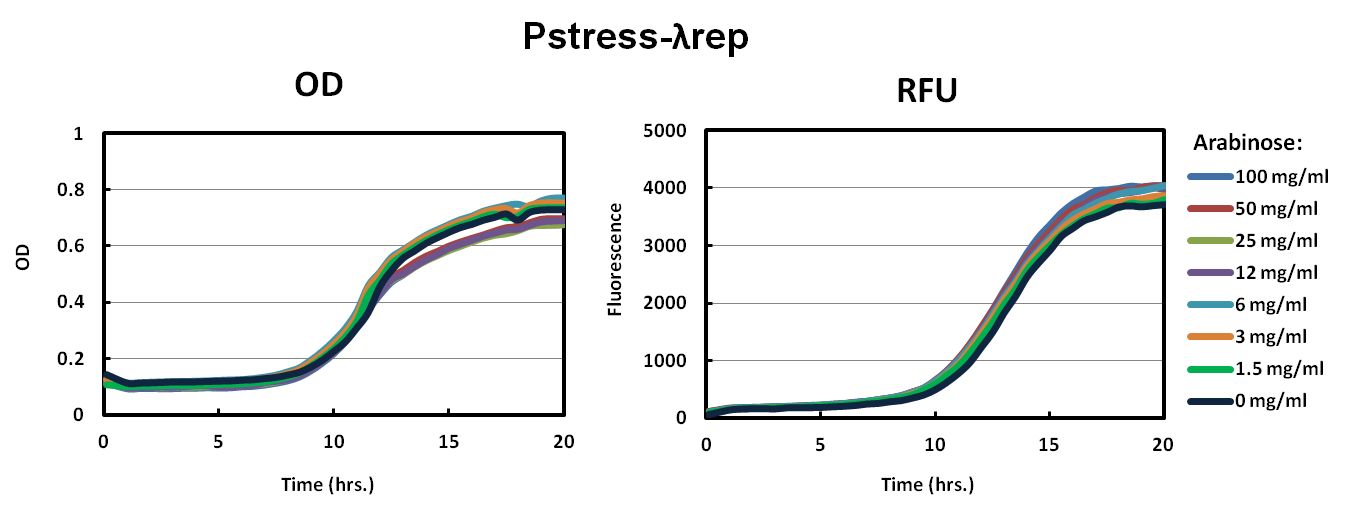
From our results thus far, we were able to:
-Characterize a library of stress promoters from the E. coli genome, providing data that they function as stress-responsive downregulators-Create an off-the-shelf negative feedback expression system using our stress promoters that enables the expression of toxic proteins to their highest non-toxic levels, solving our problem of ToxR chimera toxicity
-Demonstrated our proof of principle biosensing system by creating constitutively dimerizing ToxR chimeras that drive transcription off of the ctx promoter (Pctx)
Having achieved our goal of testing our proof of principle, we were ready to construct inducible homodimerizing systems.
Once we had successfully expressed a ToxR chimera that demonstrated constitutive transcription off Pctx we were ready to design a system that could be inducible, and thus a more useful biosensor. We set out to find a ligand dependent homodimer, and after an extensive literature search, we decided to use the Estrogen Receptor. We chose the Estrogen receptor because it fulfilled the criteria of being a homodimer and dependant on Estradiol for dimerization. Furthermore, it has been previously used in a two-hybrid assay as a chimera attached to GAL4 parts to measure ß-galactosidase activity (Peters 1999). In addition to its structural and mechanical advantages it would also make a useful biosensor since estradiol poses numerous hazards to the environment. Therefore the ability to sense it cheaply and effectively would have great implications.
The Estrogen Receptor is made up of 6 domains: A, B, C, D, E, and F. Domains A and B serve as a transcriptional factor, domain C recognizes and binds to specific DNA sequences and is also involved in dimerization (Peters 1999). Domain E is the Ligand Binding Domain (LBD); when it binds to estrogen it homodimerizes to another Estrogen Receptor LBD. Domain D’s function is unknown. Domain F has been shown to inhibit dimerization and/or binding of estrogen and is thus undesirable for our project. In a wild type system these domains interact to bind Estrogen with Domain E, confer conformation so that C can bind a specific DNA sequence, dimerize with another estrogen receptor and act as a transcription factor to promote transcription downstream of its binding.

After studying the function of each of the domains as well as the findings of Peters 1999 paper we began by making two different ToxR chimeras with two Estrogen Receptor truncations: ER∆F (no F domain) and the ER-LBD alone. Both of these estrogen receptor truncations were attached to the 3' end of ToxR and tested under various Estradiol levels. However, these truncations were non-responsive with and without estradiol. We hypothesized that different estrogen receptor truncations may be unable to express in the membrane, and the LBD by itself does not homodimerize as it didn't in the Peters' 1999 study. So, in order to mitigate both possibilities we made 8 more truncations that spanned the range between Domain A and Domain C.

Preliminary data suggests that truncations 5, 6, and 8 all showed constitutive ToxR dimerization and resulting fluorescence. However, none seemed to be inducible in response to Estrogen. Nevertheless, we believe we are getting very close to successfully expressing a chimera that will be inducible. After further reading in the literature several more strategies have been identified to make this system inducible. We are going to try several more truncations, as well as various linkers, and the co-expression of different estrogen receptor truncations as was shown to work in the Peters paper.
All of the composite parts submitted to the registry and experiments discussed on this wiki were created and performed by the 2011 iGEM team Berkeley undergraduates. Some of the basic parts (RBSes, etc.) were obtained through the Anderson lab, for which they have our thanks.
 "
"






