Commons
Gel
Investigators: Sandra, Sophie
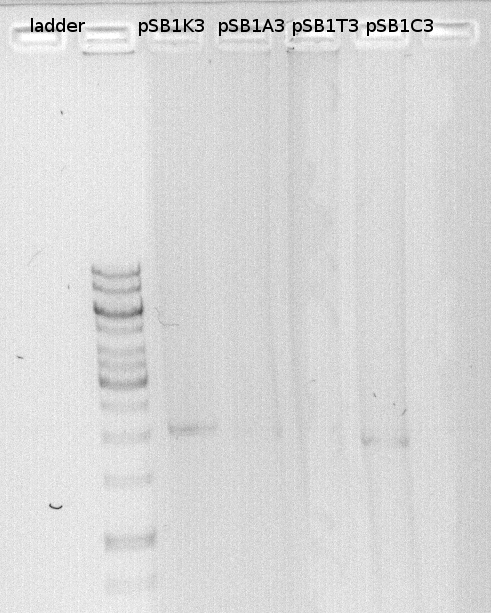
We loaded the PCR products of the vectors on a gel and measured the DNA concentration. DNA concentration were high and we diluted the vectors to get an endconcentration of 25ng/microl.
DNA-concentration measured with nanodrop:
| sample
| DNA concentration (ng/μl)
|
| pSB1C3
| 132.1
|
| pSB1K3
| 101.5
|
| pSB1A3
| 83.9
|
| pSB1T3
| 116.2
|
green light receptor
NAME OF YOUR EXPERIMENT
Investigators:NAME
blue light receptor
PCR of Lov-tap and Not-gate
Investigators: Sandra, Sophie
We repeated the PCR again, with higher temperatures this time.
After PCR we digested with DpnI 37°C for 1 hour and then purified the DNA with PCR purifictaion kit. Afterwards we measured the DNA concentration and loaded 2 microl to a gel.
PCR
| Name:Sandra, Sophie
| Date: 27.07.11
|
| Continue from Experiment: NOT-Gate and LovTAP with Gibson-overhangs (now we use a little bit higher temperature for primer annealing) -(Date): 25.07.11
(Name): Sandra, Sophie
|
| Project Name: Blue Light
|
PCR-Mixture for one Reaction:
For a 50 µl reaction use
| 32,5µl
| H20
| Name
|
| 10µl
| 5x Phusion Buffer
| of Primer
|
| 2.5µl
| Primer fw
| NOT_G_up
LOV_G_up
|
| 2.5µl
| Primer dw
| NOT_G_dw
LOV_G_dw
|
| 1µl
| dNTPs
| of Template DNA
|
| 1µl
| DNA-Template
| S35 (BBa_K322999)
S45 (Bba_Q04400)
|
| 0.5 µl
| Phusion (add in the end)
|
|
What program do you use?
"57°C auf 70°C" (first annealing temperature: 58°C and then after 10 cycles 68°C)
To confirm the PCR-Product has the correct size, load 2 µl of the sample onto an agarose-gel.
How did you label the PCR-Product, where is it stored and what do you do next?
DNA-concentration measured with nanodrop:
| sample
| DNA-concentration (ng/μl)
|
| S35
| 75.1
|
| S45
| 45.1
|
red light receptor
NAME OF YOUR EXPERIMENT
Investigators:NAME
Lysis cassette
NAME OF YOUR EXPERIMENT
Investigators:NAME
Precipitator
NAME OF YOUR EXPERIMENT
Investigators: NAME
 "
"
 Contact
Contact 












