Team:Washington/Magnetosomes/GibsonVectors
From 2011.igem.org
(→References) |
|||
| (13 intermediate revisions not shown) | |||
| Line 3: | Line 3: | ||
<center><big><big><big><big>'''iGEM Toolkits: Gibson Assembly Vectors'''</big></big></big></big></center><br><br> | <center><big><big><big><big>'''iGEM Toolkits: Gibson Assembly Vectors'''</big></big></big></big></center><br><br> | ||
| - | === About Gibson | + | === About Gibson Assembly=== |
| - | Gibson | + | Gibson assembly is a new synthetic biology tool that allows scar-free assembly of multiple gene-inserts in one isothermal reaction by using an exonuclease, polymerase, and heat-stable ligase to chew back, anneal, and repair gaps from homologous DNA fragments. This method is gaining popularity as it tends to be more efficient than standard restriction/ligation assembly, which can save a great amount of time in the cloning process. Overall, Gibson assembly allows teams to built large gene constructs with ease. |
<br/> [[File:Igem2011_GibsonReacion.png|center|400px]] | <br/> [[File:Igem2011_GibsonReacion.png|center|400px]] | ||
| - | |||
<br><br> | <br><br> | ||
------------- | ------------- | ||
| Line 13: | Line 12: | ||
=== What happened last year?=== | === What happened last year?=== | ||
The Gibson Cloning method is definitely not a new method to be introduced to the iGEM community. | The Gibson Cloning method is definitely not a new method to be introduced to the iGEM community. | ||
| - | In 2010, the Cambridge iGEM team created the [http://www.cambridgeigem.org/RFC57.pdf BBF RFC 57] document which outlines a protocol for Gibson Assembly using standard BioBricks (BBF RFC 10) that would allow many fragment inserts during a single cloning step. However, while creating the Magnetosome Toolkit, we found that this BioBrick standard was incapable of producing high yields of desired Gibson products even for two-fragment assemblies. | + | In 2010, the Cambridge iGEM team created the [http://www.cambridgeigem.org/RFC57.pdf BBF RFC 57] document which outlines a protocol for Gibson Assembly using standard BioBricks ([http://dspace.mit.edu/bitstream/handle/1721.1/45138/BBFRFC10.txt?sequence=1 BBF RFC 10]) that would allow many fragment inserts during a single cloning step. However, while creating the Magnetosome Toolkit, we found that this BioBrick standard was incapable of producing high yields of desired Gibson products even for two-fragment assemblies. |
<br> | <br> | ||
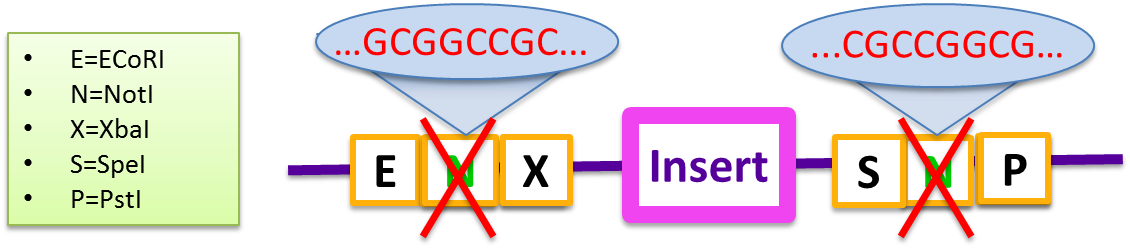
| - | The primary problem with | + | The "pSB" standard BioBrick vectors available through iGEM are not designed for efficient multiple-insert cloning beyond [http://partsregistry.org/Assembly:3A_Assembly three fragments] and are limited by ligation scars. The primary problem with a standard pSB vector is the self-complementarity of the two NotI sequences embedded in both the BioBrick prefix and suffix. These sequences prevent gene inserts from being incorporated efficiently, and do not produce a high yield of the Gibson product even in two-fragment assemblies. Generally, the backbone self-anneals and the recircularized plasmid has a combined prefix and suffix region that reads EcoRI-NotI-PstI. <br/> |
[[File:Igem2011 biobrick NotI.png|600px|center]] | [[File:Igem2011 biobrick NotI.png|600px|center]] | ||
| - | To overcome this problem, the [2010 UW iGEM] team developed new prefix and suffix regions that are based on BglBrick | + | To overcome this problem, the [https://2010.igem.org/Team:Washington/Tools_Used/Next-Gen_Cloning 2010 UW iGEM] team developed new prefix and suffix regions that are based on BglBrick (BBF [http://dspace.mit.edu/handle/1721.1/46747 RFC 21]) standard and designed to eliminate self-complementarity from the prefix and suffix of the plasmid. These vectors [https://2011.igem.org/Team:Washington/Magnetosomes/GibsonResults dramatically] increase the Gibson assembly efficiency of large-scale gene assemblies and are also compatible with iGEM standard BioBrick parts. <br/> |
[[File:Igem2011_gibsonbrick.png|600px|center]] | [[File:Igem2011_gibsonbrick.png|600px|center]] | ||
<br> | <br> | ||
| - | === What | + | === What did we do this year?=== |
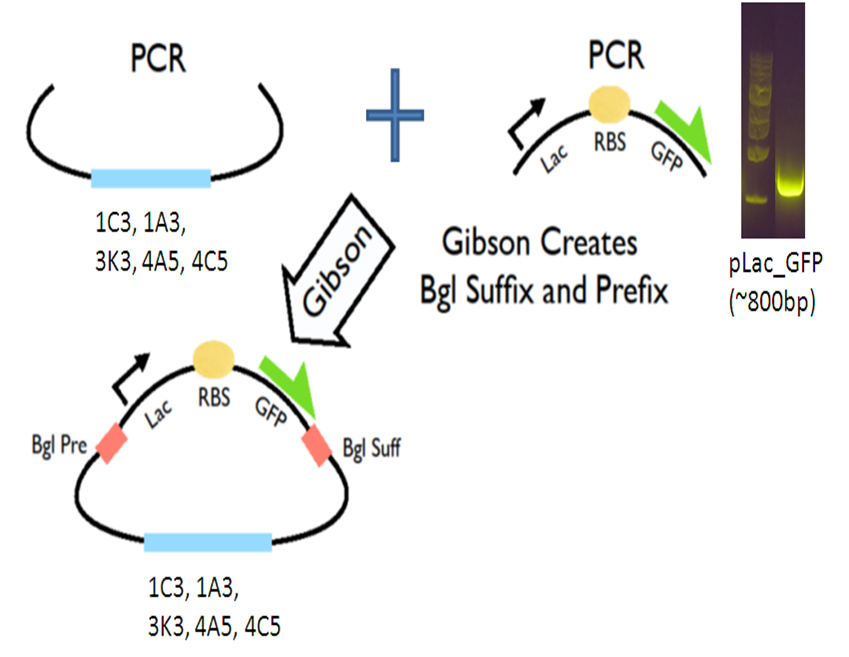
Seeing that this is a very efficient method to do cloning, we continued to make improvements to the methods and created a '''Gibson Assembly Toolkit'''! <br> <br/> | Seeing that this is a very efficient method to do cloning, we continued to make improvements to the methods and created a '''Gibson Assembly Toolkit'''! <br> <br/> | ||
| - | [[File:Washington_iGEM2011_how_to_make_vector.png| | + | [[File:Washington_iGEM2011_how_to_make_vector.png|right|thumb|450px]] |
| + | |||
===Creation of 5 plasmid vectors=== | ===Creation of 5 plasmid vectors=== | ||
Our new vectors for Gibson assembly follow the naming convention of pGA. To make our pGA vectors, we first amplified the backbones and the pLac GFP insert respectively. Then we performed a Gibson reaction to combine them together to make the pGA vectors. | Our new vectors for Gibson assembly follow the naming convention of pGA. To make our pGA vectors, we first amplified the backbones and the pLac GFP insert respectively. Then we performed a Gibson reaction to combine them together to make the pGA vectors. | ||
| Line 42: | Line 42: | ||
As listed above, each of our vectors have varying copy numbers, antibiotic resistances, and purposes within the magnetosome gene assembly. However, they all appear to be very efficient will multiple gene inserts. | As listed above, each of our vectors have varying copy numbers, antibiotic resistances, and purposes within the magnetosome gene assembly. However, they all appear to be very efficient will multiple gene inserts. | ||
| - | + | <br/><br/><br/><br/><br/> | |
| - | For experimental details comparing the efficiencies of pSB1A3 and pGA1A3, see [https://2011.igem.org/Team:Washington/Magnetosomes/GibsonResults our assay results]. | + | For experimental details comparing the efficiencies of pSB1A3 and pGA1A3, see [https://2011.igem.org/Team:Washington/Magnetosomes/GibsonResults our assay results]. In addition to the characterization of assembly efficiency, we used the pGA vectors for the [https://2011.igem.org/Team:Washington/Magnetosomes/Magnet_Toolkit Magnetosome Toolkit] project. First, we efficiently extracted and BioBricked all the 18 <i>mamAB</i> magnetosome genes. We also made super-assemblies from these BioBricks to make 10 kilobase plasmids (see [http://partsregistry.org/wiki/index.php?title=Part:BBa_K590017 mamHIEJKL] and [http://partsregistry.org/wiki/index.php?title=Part:BBa_K590019 mamQRBSTUV]) from 2 kilobase pieces by assembling up to five fragments all in one cloning step! |
| Line 52: | Line 52: | ||
2. Peter A Carr and George M Church. Genome Engineering. <i>Nat Biotechnol</i> , <b>27</b>(12):1151 Dec 2009.<br> | 2. Peter A Carr and George M Church. Genome Engineering. <i>Nat Biotechnol</i> , <b>27</b>(12):1151 Dec 2009.<br> | ||
3. Daniel G Gibson, Hamilton O Smith, Clyde A Hutchison Iii, J Craig Venter, and | 3. Daniel G Gibson, Hamilton O Smith, Clyde A Hutchison Iii, J Craig Venter, and | ||
| - | Chuck Merryman. Chemical synthesis of the mouse mitochondrial genome. <i> | + | Chuck Merryman. Chemical synthesis of the mouse mitochondrial genome. <i>Nat Methods</i>, <b>7</b>(11):901, Oct 2010. |
Latest revision as of 00:24, 29 September 2011
About Gibson Assembly
Gibson assembly is a new synthetic biology tool that allows scar-free assembly of multiple gene-inserts in one isothermal reaction by using an exonuclease, polymerase, and heat-stable ligase to chew back, anneal, and repair gaps from homologous DNA fragments. This method is gaining popularity as it tends to be more efficient than standard restriction/ligation assembly, which can save a great amount of time in the cloning process. Overall, Gibson assembly allows teams to built large gene constructs with ease.
What happened last year?
The Gibson Cloning method is definitely not a new method to be introduced to the iGEM community.
In 2010, the Cambridge iGEM team created the [http://www.cambridgeigem.org/RFC57.pdf BBF RFC 57] document which outlines a protocol for Gibson Assembly using standard BioBricks ([http://dspace.mit.edu/bitstream/handle/1721.1/45138/BBFRFC10.txt?sequence=1 BBF RFC 10]) that would allow many fragment inserts during a single cloning step. However, while creating the Magnetosome Toolkit, we found that this BioBrick standard was incapable of producing high yields of desired Gibson products even for two-fragment assemblies.
The "pSB" standard BioBrick vectors available through iGEM are not designed for efficient multiple-insert cloning beyond [http://partsregistry.org/Assembly:3A_Assembly three fragments] and are limited by ligation scars. The primary problem with a standard pSB vector is the self-complementarity of the two NotI sequences embedded in both the BioBrick prefix and suffix. These sequences prevent gene inserts from being incorporated efficiently, and do not produce a high yield of the Gibson product even in two-fragment assemblies. Generally, the backbone self-anneals and the recircularized plasmid has a combined prefix and suffix region that reads EcoRI-NotI-PstI.
To overcome this problem, the 2010 UW iGEM team developed new prefix and suffix regions that are based on BglBrick (BBF [http://dspace.mit.edu/handle/1721.1/46747 RFC 21]) standard and designed to eliminate self-complementarity from the prefix and suffix of the plasmid. These vectors dramatically increase the Gibson assembly efficiency of large-scale gene assemblies and are also compatible with iGEM standard BioBrick parts.
What did we do this year?
Seeing that this is a very efficient method to do cloning, we continued to make improvements to the methods and created a Gibson Assembly Toolkit!
Creation of 5 plasmid vectors
Our new vectors for Gibson assembly follow the naming convention of pGA. To make our pGA vectors, we first amplified the backbones and the pLac GFP insert respectively. Then we performed a Gibson reaction to combine them together to make the pGA vectors.
All togeher, we created 5 pGA vectors and submitted them to the registry:
- 2 High copy extraction/cloning vectors
- pGA1A3, pGA1C3
- 1 medium copy expression vector
- pGA3K3
- 2 low copy expression vectors
- pGA4A5, pGA4C5
As listed above, each of our vectors have varying copy numbers, antibiotic resistances, and purposes within the magnetosome gene assembly. However, they all appear to be very efficient will multiple gene inserts.
For experimental details comparing the efficiencies of pSB1A3 and pGA1A3, see our assay results. In addition to the characterization of assembly efficiency, we used the pGA vectors for the Magnetosome Toolkit project. First, we efficiently extracted and BioBricked all the 18 mamAB magnetosome genes. We also made super-assemblies from these BioBricks to make 10 kilobase plasmids (see [http://partsregistry.org/wiki/index.php?title=Part:BBa_K590017 mamHIEJKL] and [http://partsregistry.org/wiki/index.php?title=Part:BBa_K590019 mamQRBSTUV]) from 2 kilobase pieces by assembling up to five fragments all in one cloning step!
References
1. Daniel G Gibson, Lei Young, Ray-Yuan Chuang, Craig J Venter, Clyde A Hutchison, and Hamilton O Smith. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat Methods, 6(5):343, Apr 2009.
2. Peter A Carr and George M Church. Genome Engineering. Nat Biotechnol , 27(12):1151 Dec 2009.
3. Daniel G Gibson, Hamilton O Smith, Clyde A Hutchison Iii, J Craig Venter, and
Chuck Merryman. Chemical synthesis of the mouse mitochondrial genome. Nat Methods, 7(11):901, Oct 2010.
 "
"