Team:Tsinghua/experiment
From 2011.igem.org
(→SDS-PAGE) |
(→SDS-PAGE) |
||
| Line 192: | Line 192: | ||
| '' 14. P 0.1mM 30℃'' | | '' 14. P 0.1mM 30℃'' | ||
|} | |} | ||
| + | |||
| + | |||
| + | Contrary to the sharp change in color, no obvious band can be seen in SDS-PAGE. Hence, further purification of protein is necessary. | ||
==Ni-column purification== | ==Ni-column purification== | ||
Revision as of 16:04, 4 October 2011

Present proteins to the outer membrane
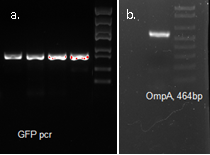
We used OmpA-His tag-GFP to test the conditions for presenting protein to the outer membrane. The design allows double check whether the C-terminal moeity can be presented outside. One is by green fluorescence, the other is by Western blotting with antibody against his tag.
Molecular cloning
OmpA, GFP His tag 1507bp confirmed by sequencing
The final plasmid is as follows.
Expression Test
Test for proper inducing conditions
| Group | 1 | 2 |
|---|---|---|
| Volume | 3ml | |
| IPTG(mM) | 0.1 | |
| 0.5 | ||
| 1.0 | ||
| Temperature | 30oC | 18oC |
| Duration | 12h | 12h |
| Result | Media remained yellowish. E. coli white | Media turned greenish. E. coli green |
Use 0.5mM IPTG, 18 centigrade, 12 hours induction in later experiments.
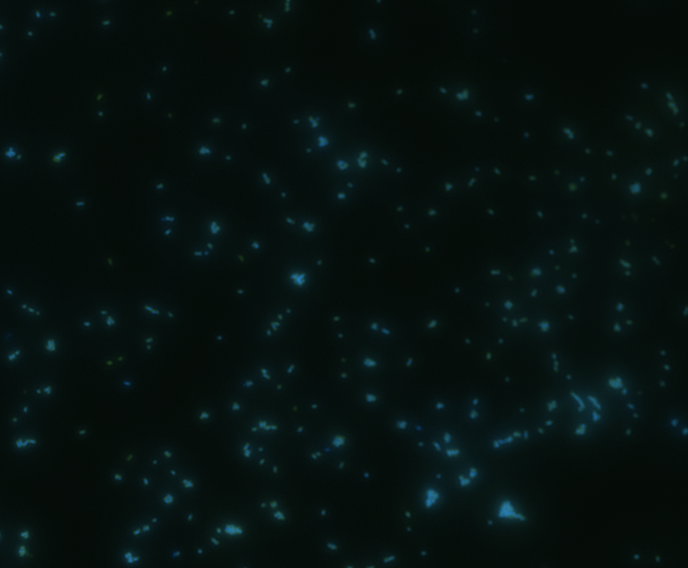
Fluorescence microscopy
Green fluorescence is bright.
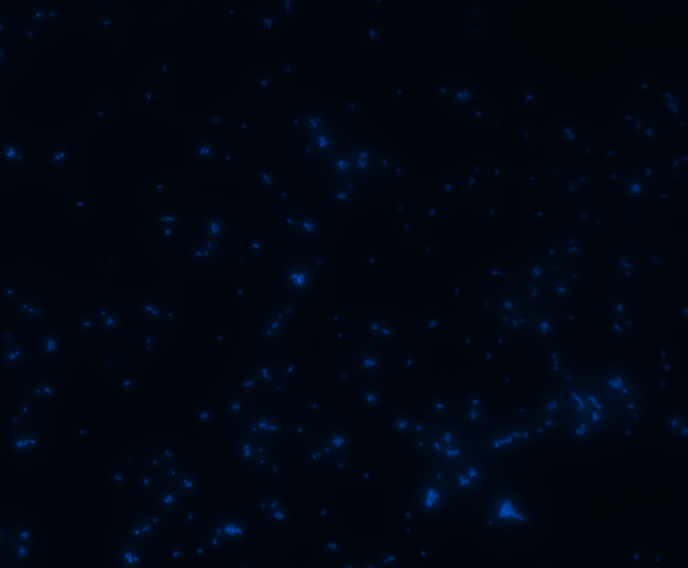
In order to determine whether GFP is presented outside the membrane, cells were digested with Protease K at 50 centigrade for 30min.
| Color Channel | Post-digest | Pre-digest |
|---|---|---|
| GFP | 
| 
|
| DAPI | 
| 
|
| Merge | 
| 
|
The fluorescence disappeared completely after digestion, which seems too good to be true. Hence, we performed Western blotting to be sure of the result.
Western Blotting
The cell components were first fractionated before blotting.
1. Sonicate the bacteria culture.
2. Centrifuge 13000rpm, 30min to separate soluble protein and membrane components from the cell debris and the non-soluble protein.
3. 51000rpm 1h to separate soluble protein from membrane components.
The majority of the OmpA-GFP protein is in the pellet after the first centrifuge, indicating incomplete cell lysis or inefficient protein folding. Nonetheless, from the later result, we can see that it is exclusively in the membrane and not in the soluble proteins.
Expression of Substrate
Molecular cloning
We synthesized the multi-proline sequence and linked it to mCherry.
Test of Expression
| Group | 1 | 2 |
|---|---|---|
| Volume | 3ml | |
| IPTG(mM) | 0.1 | |
| 0.5 | ||
| 1.0 | ||
| Temperature | 30oC | 18oC |
| Duration | 12h | 12h |
| Result | Media remained yellowish. E. coli white | Media turned purple. E. coli cherry pink |
After Pro-rich mCherry expression, significant color change can be observed (bacteria cells turned cherry red as a result of mCherry expression)
From left to right, IPTG concentration: 0, 0.1mM, 0.5mM, 1mM
SDS-PAGE
The bacteria culture was sonicated and then centrifuged at 13000rpm for 15min. Both the supernatant and the pellet were loaded onto SDS-PAGE.
| 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | Marker | 9 | 10 | 11 | 12 | 13 | 14 |
| 1. Pellet, 1mM 18℃ | 2. Supernatant 1mM 18℃ | 3. P 0.5mM 18℃ |
| 4. S 0.5mM 18℃ | 5. P 0.1mM 18℃ | 6. S 0.1mM 18℃ |
| 7. P NC 18℃ | 8. S NC 18℃ | 9. S 1mM 30℃ |
| 10. P 1mM 30℃ | 11. S 0.5mM 30℃ | 12. P 0.5mM 30℃ |
| 13. S 0.1mM 30℃ | 14. P 0.1mM 30℃ |
Contrary to the sharp change in color, no obvious band can be seen in SDS-PAGE. Hence, further purification of protein is necessary.
Ni-column purification
The cell lysate was then purified by Ni-column. SDS-PAGE indicates that the protein is largely purified.
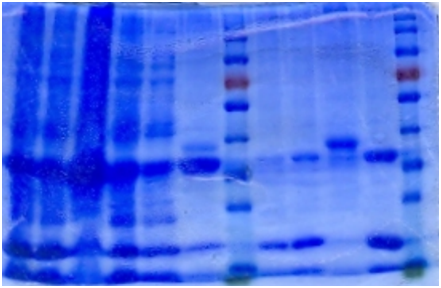 1. Sonicated
2. Supernatant
3. Pellet
4. Flowthrough
5. Buffer wash
6. Resin
7. 5mM Elution
8. 20mM Elution
9. Resin 2
10. 200mM Elution
1. Sonicated
2. Supernatant
3. Pellet
4. Flowthrough
5. Buffer wash
6. Resin
7. 5mM Elution
8. 20mM Elution
9. Resin 2
10. 200mM Elution
1 2 3 4 5 6 Marker 7 8 9 10
Western blotting
In order to be sure that the protein we got is multi-proline mCherry, Western blotting was performed.
 "
"