Team:Washington/Celiacs/Results
From 2011.igem.org
| Line 3: | Line 3: | ||
<center><big><big><big><big>Gluten Destruction: Results Summary</big></big></big></big></center><br><br> | <center><big><big><big><big>Gluten Destruction: Results Summary</big></big></big></big></center><br><br> | ||
| + | |||
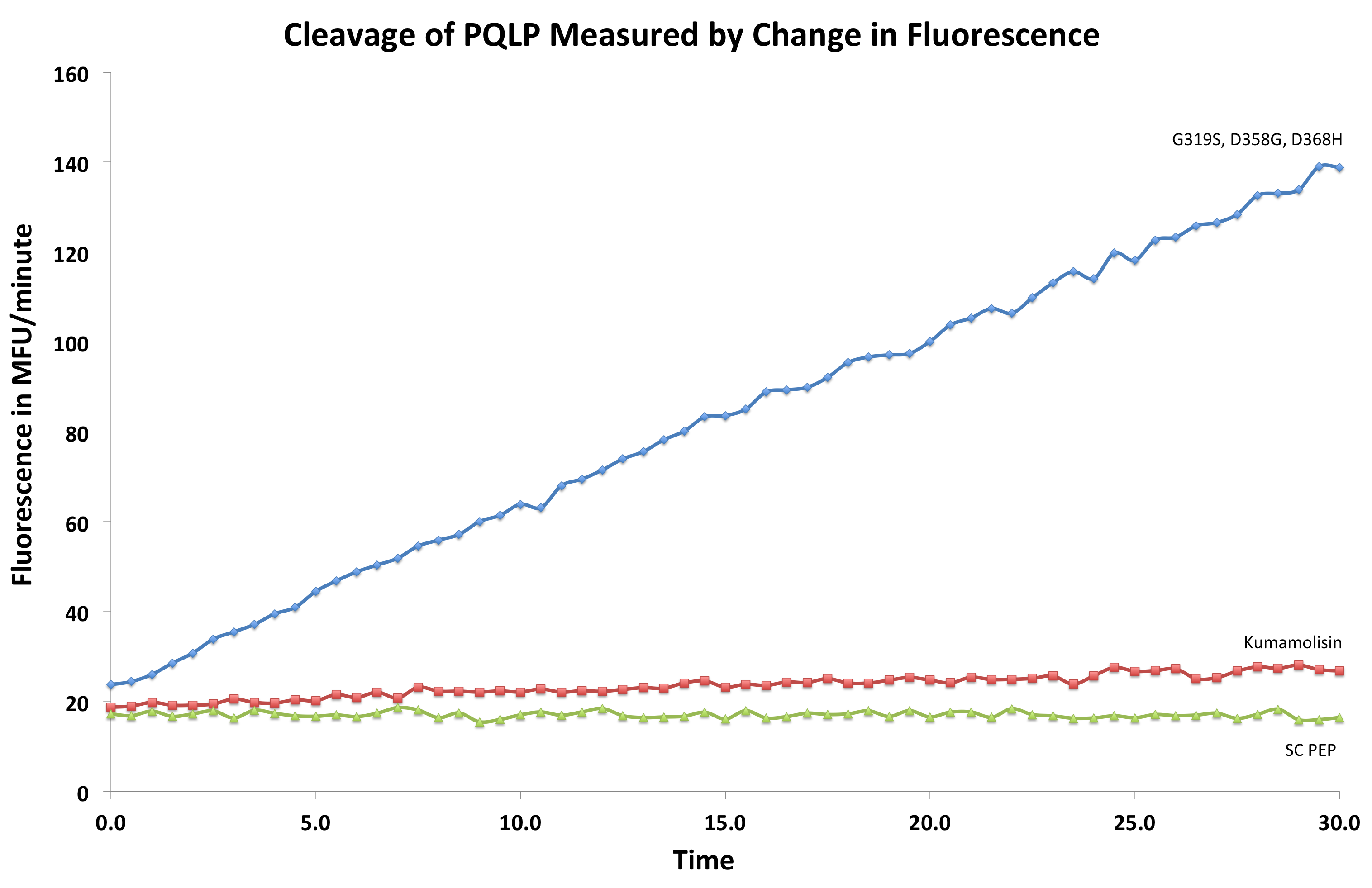
| + | [[File:Washington First Raw Data.png|center|500px|thumb|We measured fluorescence of each reaction at 30 second intervals to see the rate at which each mutant cleaved PQLP.]] | ||
=Testing Kumamolisin-As against SC-PEP= | =Testing Kumamolisin-As against SC-PEP= | ||
Revision as of 01:46, 23 September 2011
Testing Kumamolisin-As against SC-PEP
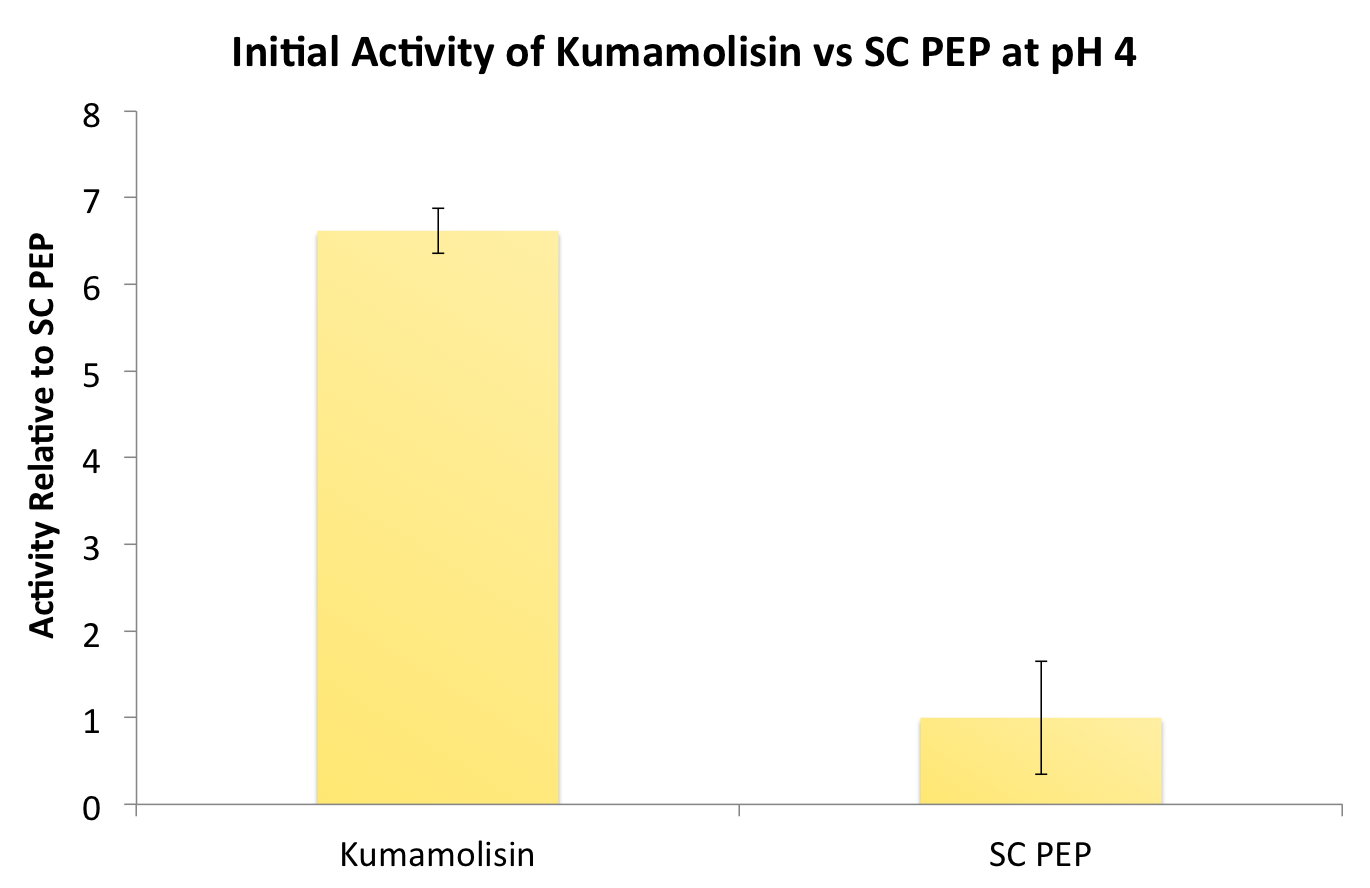
After finding Kumamolisin-As to be the ideal enzyme for our purposes, we used the assay described below to test it against SC-PEP to determine comparative activity levels, resulting in evidence that wild-type Kumamolisin-As is already over 7 fold better than SC-PEP.
Testing mutants for activity on breaking down PQLP
Using a whole cell lysate assay to screen a large number of mutants for good activity
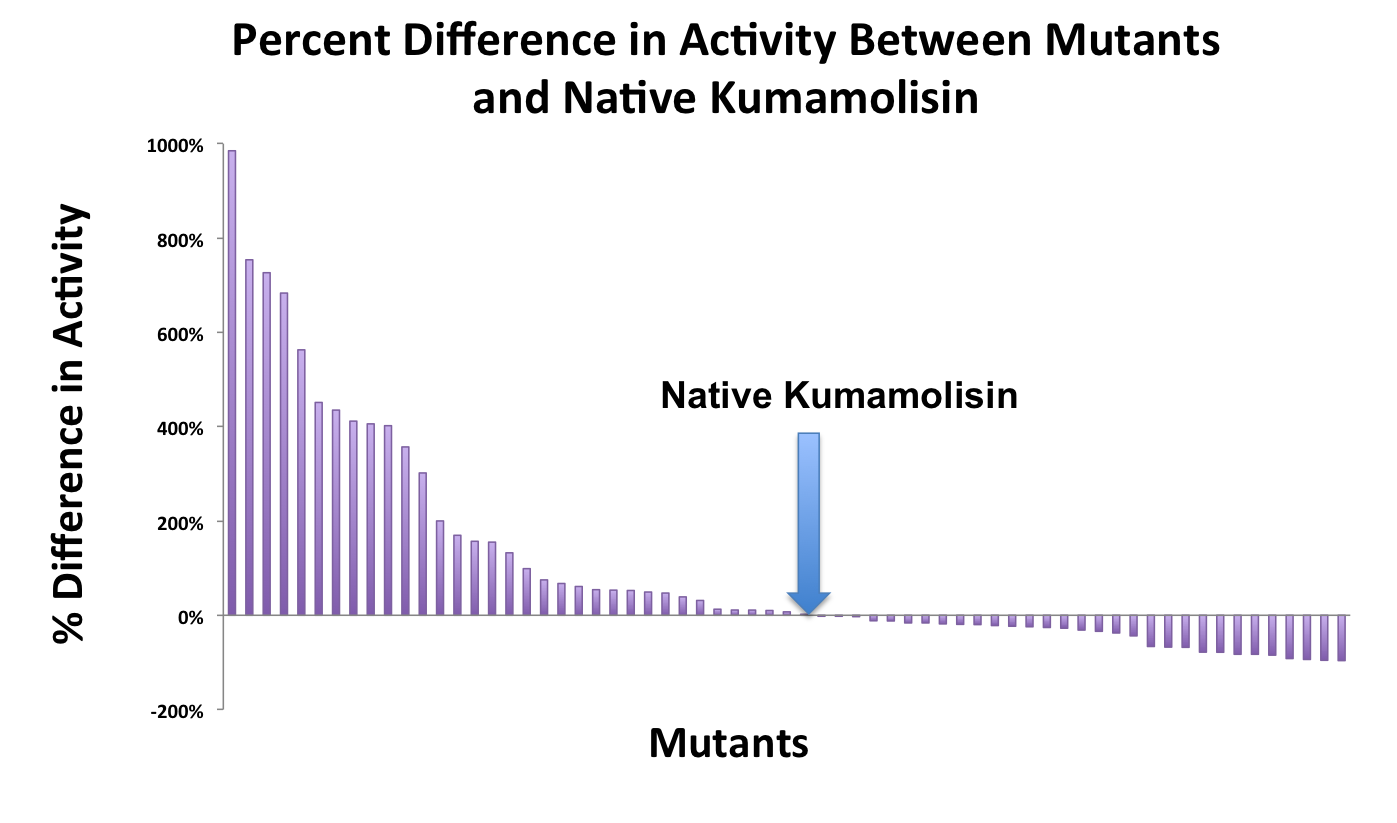
In order to determine whether our proposed mutations to the wild-type Kumamolisin improved the ability of the enzyme to break down PQLP, we tested each mutant with a whole cell lysate fluorescence assay. Cells harboring the expressed mutants were lysed and the assay was performed at pH 4, mimicking the gastric environment. The released enzymes, after being roughly separated from cell material, were added to a fluorescent PQLP that had been conjugated to a quencher. Thus, no fluorescence was achieved until the peptide had been cleaved and the fluorophore had been released from the quencher. This allowed a relative assessment of rate of enzyme activity by measuring increase in fluorescence of the system.
As one might expect, our first screen of mutants showed some mutants with a decrease in activity from the wild-type, some showed no change, and some actually showed great increase in activity. One single point mutant showed close to a 1000% increase in activity from wild-type Kumamolisin!
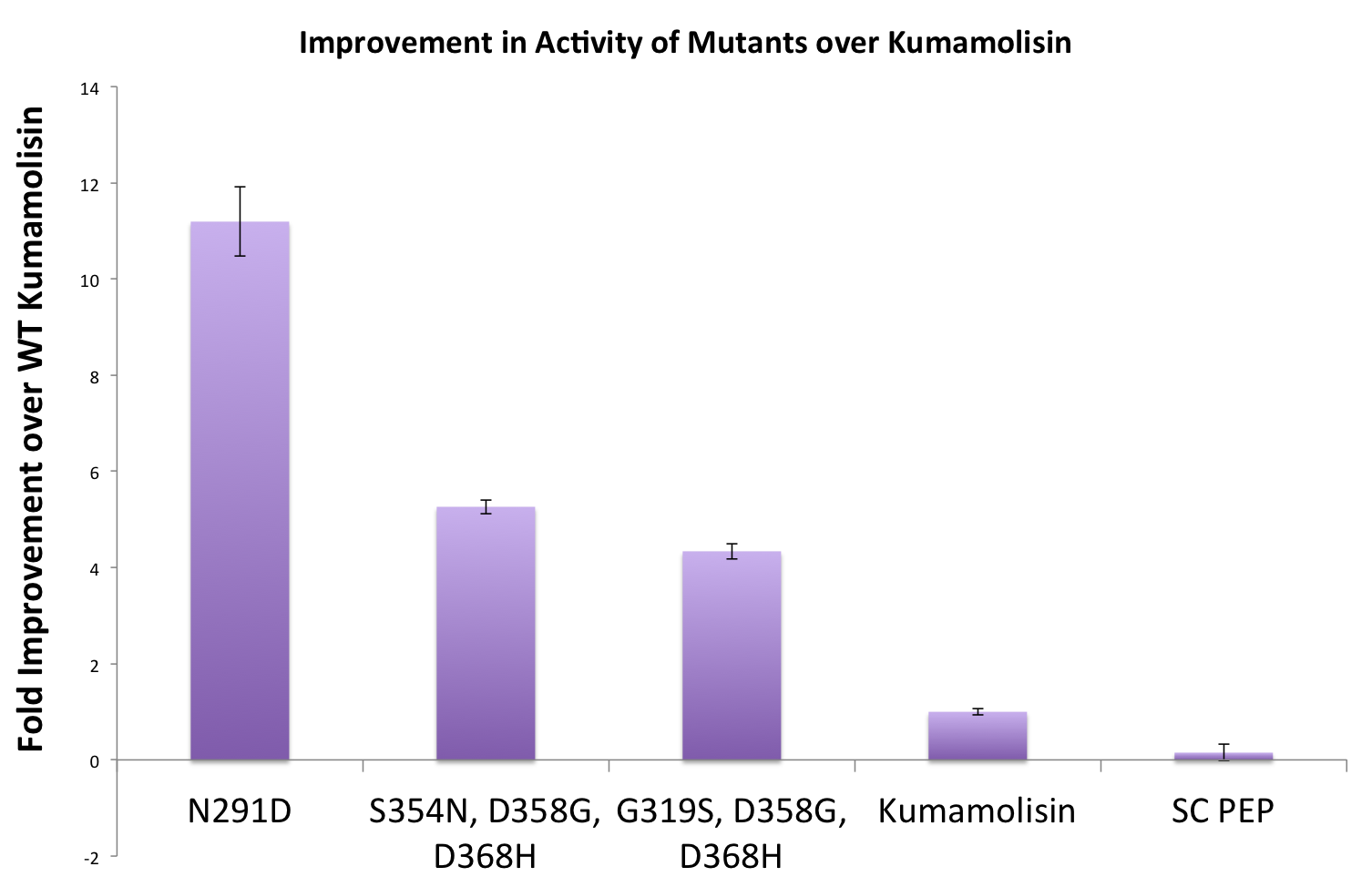
Purifying and characterizing promising mutants for accurate rate comparison
Once we had identified mutants that showed a promising increase in activity from the wild-type kumamolisin, we purified and characterized activity in concentration controlled fluorescence assays, identical to the fluorescence system used for the whole cell lysate assay. Our best mutant demonstrated an 11-fold increase in activity from the native enzyme.
Combining Mutants for the construction of a Gluten Hydrolase
A second library of based on the first round of mutagensis was constructed and tested
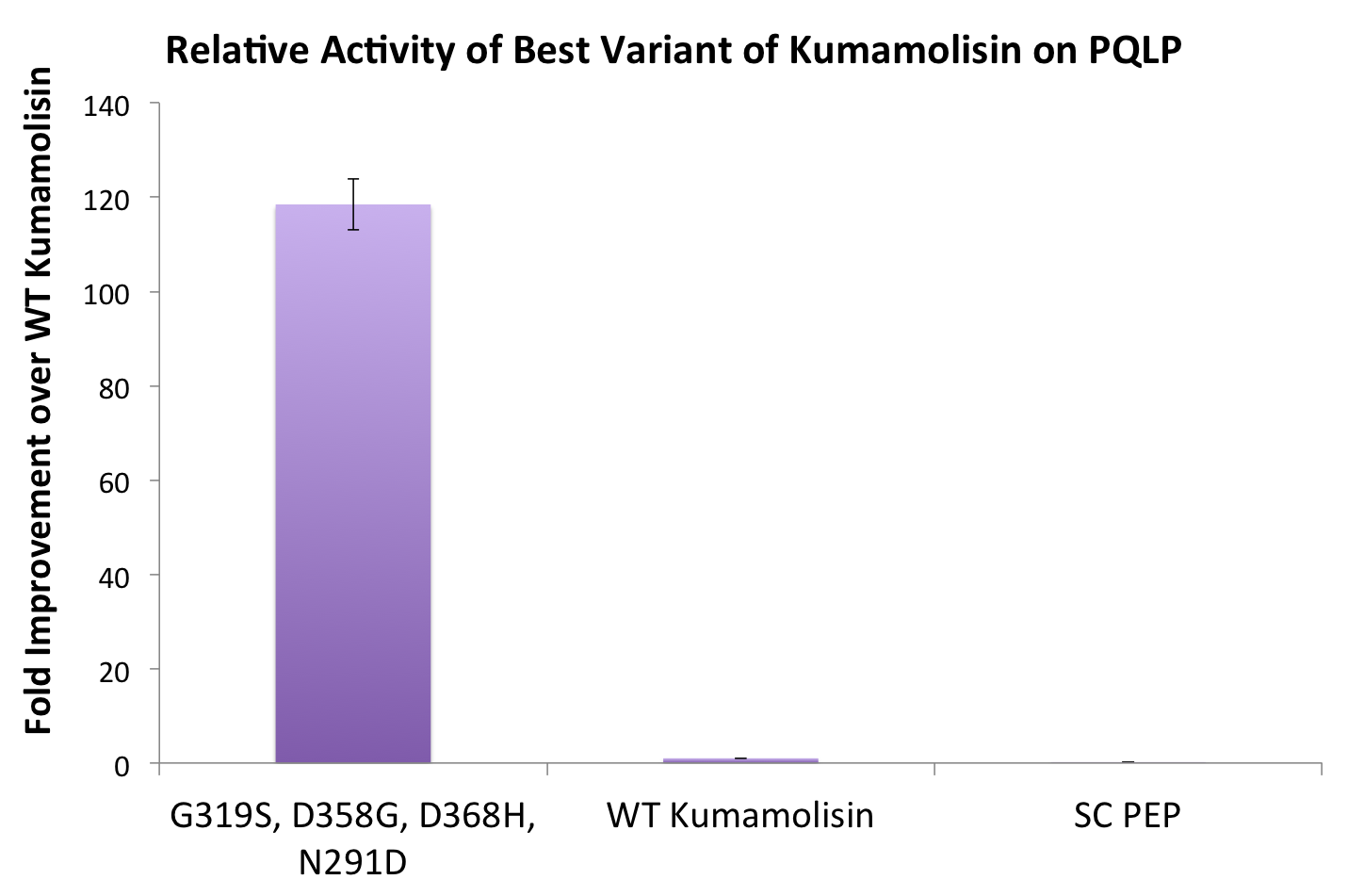
One of the combinatorial mutants resulted in over a 100-fold increase in activity
In order to achieve even more rate improvement from the native, we repeated our mutagenesis, this time taking successful mutations and adding them all to make combinatorial variants. By combining two of our top groups of mutations from the first round, we achieved an over 100-fold increase in activity on breaking down PQLP from the wild-type enzyme. This variant enzyme is ultimately 784 times better at breaking down PQLP than SC PEP, the enzyme currently in clinical trials for treating gluten intolerance!
 "
"