File:UCL-content-GBS-theory-figure-6.jpg
From 2011.igem.org
No higher resolution available.
UCL-content-GBS-theory-figure-6.jpg (750 × 275 pixels, file size: 68 KB, MIME type: image/jpeg)
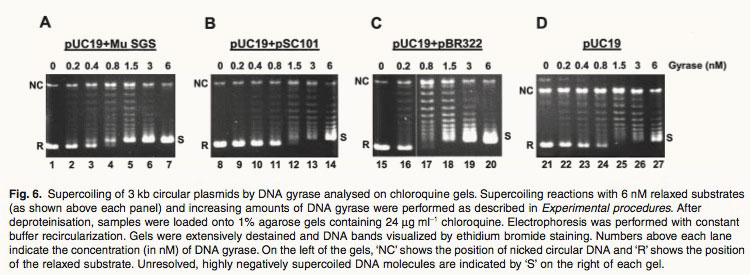
Fig. 6. Supercoiling of 3 kb circular plasmids by DNA gyrase analysed on chloroquine gels. Supercoiling reactions with 6 nM relaxed substrates (as shown above each panel) and increasing amounts of DNA gyrase were performed as described in Experimental procedures. After deproteinisation, samples were loaded onto 1% agarose gels containing 24 mg ml-1 chloroquine. Electrophoresis was performed with constant buffer recircularization. Gels were extensively destained and DNA bands visualized by ethidium bromide staining. Numbers above each lane indicate the concentration (in nM) of DNA gyrase. On the left of the gels, ‘NC’ shows the position of nicked circular DNA and ‘R’ shows the position of the relaxed substrate. Unresolved, highly negatively supercoiled DNA molecules are indicated by ‘S’ on the right of each gel. Ref: 1. Oram, M., et al., A biochemical analysis of the interaction of DNA gyrase with the bacteriophage Mu, pSC101 and pBR322 strong gyrase sites: the role of DNA sequence in modulating gyrase supercoiling and biological activity. Mol Microbiol, 2003. 50(1): p. 333-47.
File history
Click on a date/time to view the file as it appeared at that time.
| Date/Time | Thumbnail | Dimensions | User | Comment | |
|---|---|---|---|---|---|
| current | 04:45, 22 September 2011 | 750×275 (68 KB) | PhilippBoeing (Talk | contribs) | ||
| 11:09, 21 September 2011 | 750×275 (95 KB) | Eintisar (Talk | contribs) | (Fig. 6. Supercoiling of 3 kb circular plasmids by DNA gyrase analysed on chloroquine gels. Supercoiling reactions with 6 nM relaxed substrates (as shown above each panel) and increasing amounts of DNA gyrase were performed as described in Experimental pro) |
File links
The following page links to this file:
 "
"